Fragment-based screening is now well-established as a powerful approach to early drug ("lead") discovery.
Email: [email protected]
Tel: +44 (0) 1235 778926
XChem: X-ray structure-accelerated, synthesis-aligned lead discovery
Fragment-based screening is now well-established as a powerful approach to early drug ("lead") discovery. Of the many suitable biophysical techniques, X-ray crystallography was one of the first to be used, and is the most directly informative (reference); however, the experimental overheads have historically been too high for it to be widely used for primary screening.
At Diamond in Lab34 and beamline I04-1, the full X-ray screening experiment has now been implemented as a highly streamlined process, allowing up to 1000 compounds to be screened individually in less than a week (including 36 hours' unattended beamtime). The process covers soaking, harvesting, automatic data collection, and data analysis; fragment libraries are available, though users can bring their own.
Since April 2015, the XChem facility has been available to users with dedicated weekly beamtime, and is now part of the MX User Programme (details on how to apply here); as a world-wide first facility, we welcome international proposals. The future focus is on exploring how best to harness the technique to proceed rapidly to potent compounds.
There's a longer narrative of the facility in The Translational Scientist (free registration required) and also in this podcast.
The user community is growing and you can follow current discussions in the XChem bulletin board.
The facility is based in Lab34 and the adjacent beamline I04-1.
Access is structured as Tier 1, for establishing crystal systems and Tier 2 for full screens. Currently under discussion is Tier 3 access, to support hit follow-up chemistry. More information on the Tier system can be found here.
In practice, the experiment will span a few days and even multiple visits to establish crystals' suitability. Users must generate the crystals in their home lab or provide materials and protocols for the XChem team, and are required to come and perform soaking and harvesting themselves. Multi-day Lab Visits will be scheduled separate from your normal beamtime allocation. In contrast, users do not need to be present for the X-ray data collection, which will also not be booked to your usual BAG or other allocation. A local contact will be assigned, in the same way as for beamtime.
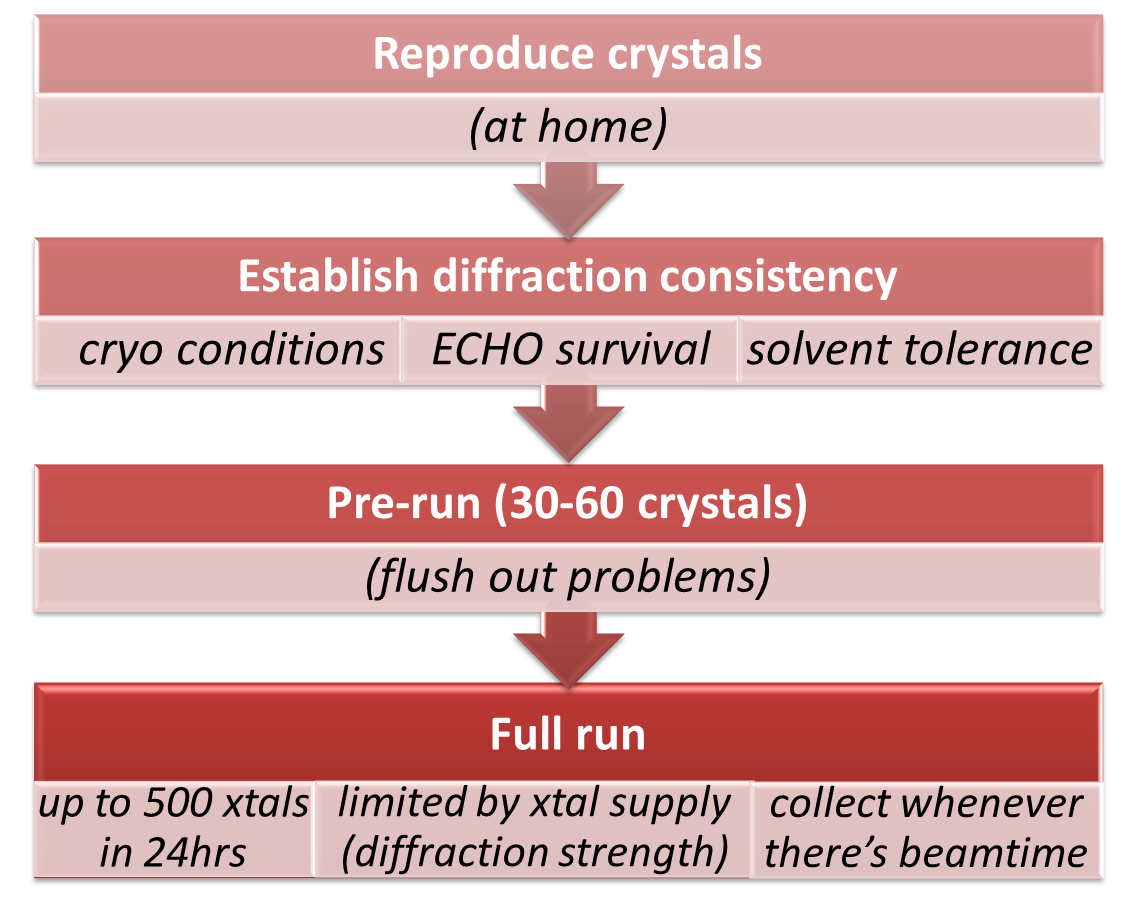
The workflow of an ideal experiment is outlined in the adjacent schema. In practice, the first three steps are iterative and require a few dozen crystals, and in difficult cases even several Lab Visits but associated diffraction testing will be fitted in during the Lab Visit where possible. The final "Full run" soaking and harvesting will be scheduled once the soaking protocol is confirmed (in well-behaving cases, during the same Lab Visit).
Data analysis builds on the existing automatic data processing, and we have developed tools to streamline density interpretation and refinement (PanDDA and XChemExplorer). Use of these tools at Diamond is optional but highly recommended: they are taylored very specifically to the many-dataset problem, and have been extensively tested. We also provide some support in analysis and presentation of hit results, and depositing hit structures.
Full details are provided here.
Academic access is governed by the same usage policies as the rest of Diamond's user programme. In particular, users are required to publish their results and deposit their hit structures in the PDB, as for all MX beamtime. In addition, we may occasionally request brief reports on how hits are progressing to potent compounds, to help us evaluate the effectiveness of the facility.
Academic access is not limited to academic groups; industry-based users can also apply by the same peer-review route, but will then be expected to publish and deposit just like academic groups. (This is in contrast to commercial access via the Industrial Liason Office, which is incidentally also available to academic groups provided they are prepared to pay commercial rates.)
To make enquiries, please contact us here.
However, please first carefully read the contents of these pages, especially the 'Accessing XChem' section, which ought to answer most questions.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.