Screening Samples
There are now three methods of screening samples using Unattended Data Collection:
| Screening Method | Relative Speed | Example use case |
| None | Fastest | A crystal system with diffraction up to a known resolution. |
| Better Than | Slower | A new or inconsistently difffracting crystal system where you only want to collect full datasets with diffraction better than a given resolution |
| Collect Best N | Slower | A crystal system where you want to collect N of the best samples, such as when testing crystallisation conditions |
Any experiment kind (ligand, native, phasing) can be used with any screening method.
As the time taken to load the sample is larger than the time taken to expose the crystal to X-rays, any screening strategy will be slower than UDC without a screening strategy.
Screening Collection
Currently screening is carried out using a 15° wedge of data collected with the detector close to the sample and at a high dose.
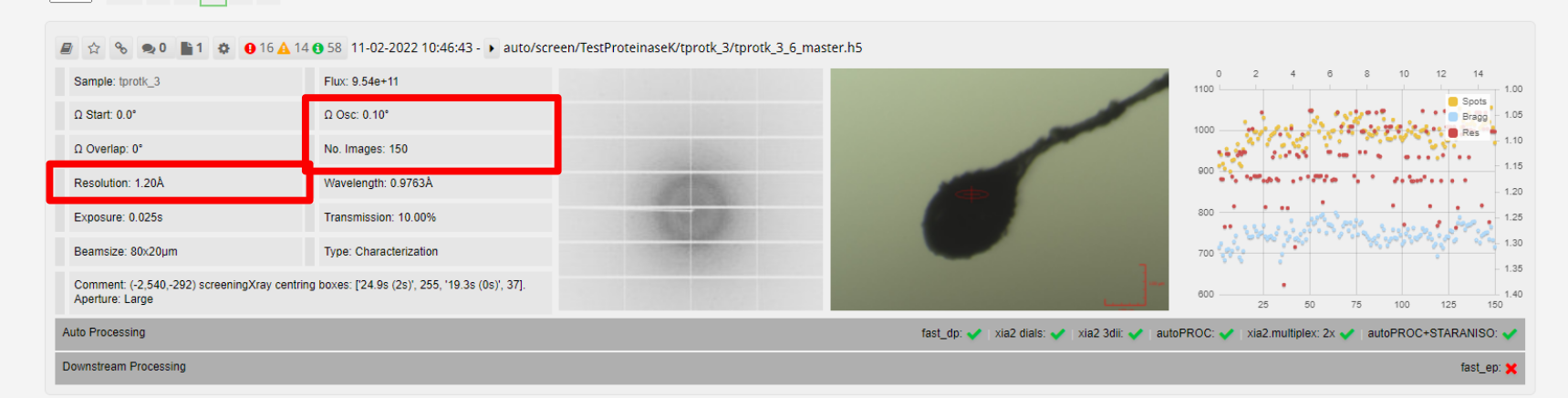
The samples that are for screening are collected first, and autoprocessing of the data begins. FastDP is used to determine a cutoff resolution. A buffer of 0.5Å is added, so if the cutoff were seen to be at 1.5Å, data would be collected at a detector position equivalent to 1Å. This is bounded within 0.9Å and 4.5Å. The buffer takes into account the larger rotation range that would be seen in a complete collection.
If all autoprocessing fails on the sample, no full collection is made. If FastDP fails, but any other processing pipeline works, then that cutoff value is used instead, and the sample is returned to, once those results are available.
This is subject to change as we develop the screening procedure to best optimise the variety of samples and improvements to the beamlines.
Sample table: Unattended
The interface to upload samples has been updated in April 2022. See Registering Samples for details on the puck creation process. See Preparation for UDC for how to specify UDC samples generally.
Sample grouping
The definition of sample groups is used for the screening strategies for Unattended Data Collection.
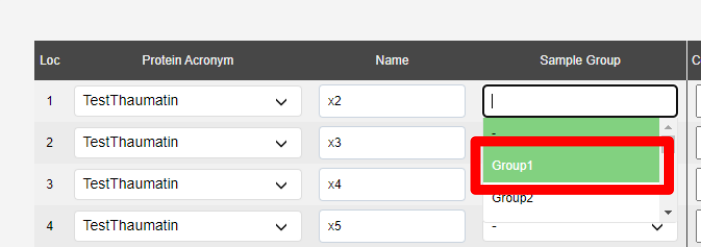
A sample group is how the screening methods knows which samples to compare. It is only needed for the Collect Best N method.
For example, consider the case where you have multiple experiments to carry out with the same protein.
In one (highlighted in red below) you are testing the crystallisation conditions for Thaumatin, and want to collect anything better than 2Å across all conditions.
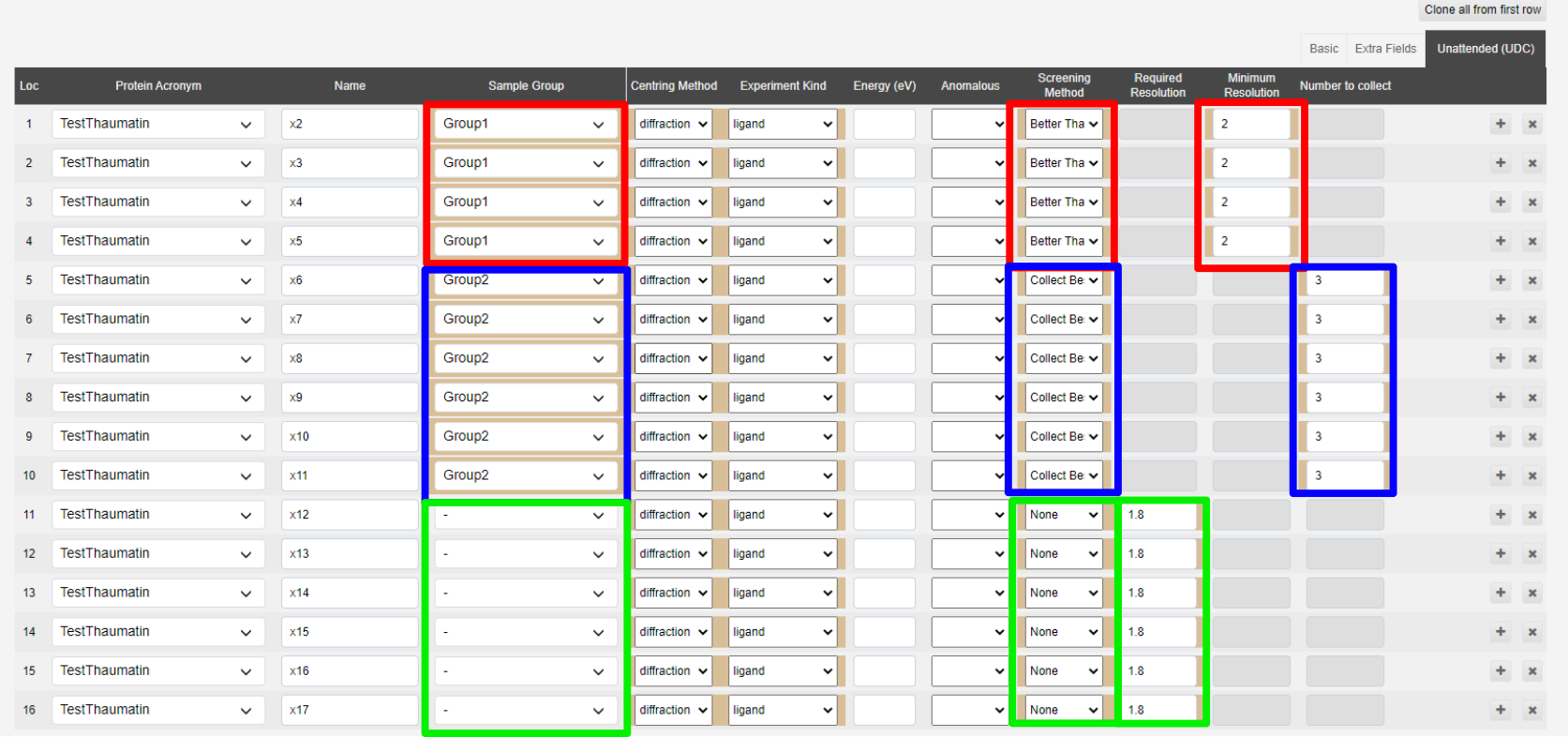
You also have a known crystallisation condition where you are testing a new co-factor binding, you want to collect the best 3 diffracting crystals, highlighted in blue.
You also have some apo collections to do in the known diffracting conditions, highlighted in green.
In each screening method, you need to provide differing information. In the Better Than screening mode, you need to provide a Minimum Resolution, in this case 2Å. In the Collect Best N you need to provide the number of datasets you wish to collect. And with screening set to None you would need to provide the resolution to set the detector at.
In Unattended Data Collection the use of sample groups does not persist between shipments, as each shipment is considered a separate experiment. The group name is available to be reused for an alternate grouping of samples on the next shipment.
CSV Upload
The screening strategies can also be defined during csv upload.
