Each circle on the figure below represents 180o sweep on the same crystal at three different energies. For example GDA will first collect sweep 1 (0 to 45o) at Energy 1, then sweep 2 (0 to 45o) at Energy 2, followed by sweep 3 (0 to 45o) at Energy 3 and so on...

The advantage using wedged MAD collection compared to regular MAD data collection is that the same reflection at three (or more) different energies will be collected with equivalent dose. It will result in a better assesment of the anomalous signal less polluted by radiation damage.
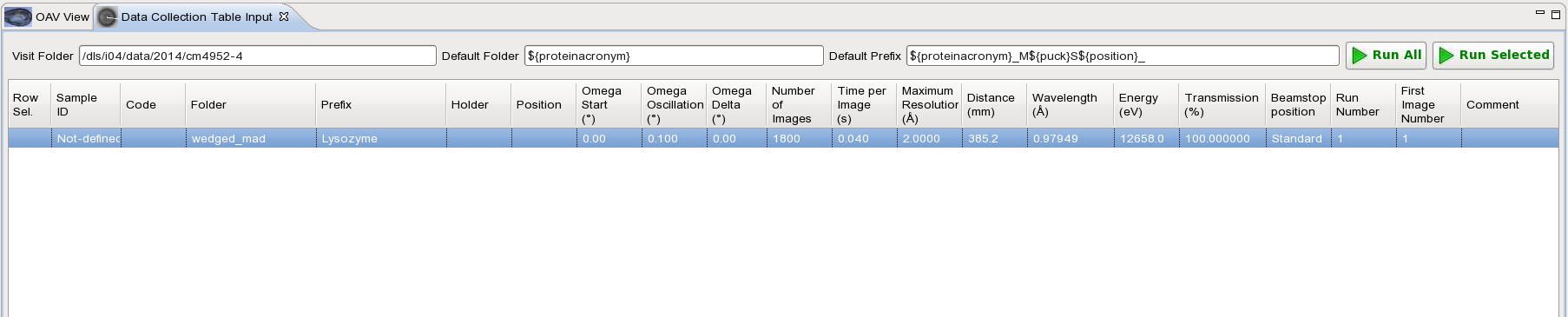
In GDA, go to "Data Collection" then click on the tab "Data Collection Table Input":
![]()
Enter the Folder where you want to collect your data, Prefix and the specs for the data collection. For example for each energy we will collect 1800 images, 0.1o oscillation, 0.040s exposure and 100% transmission:
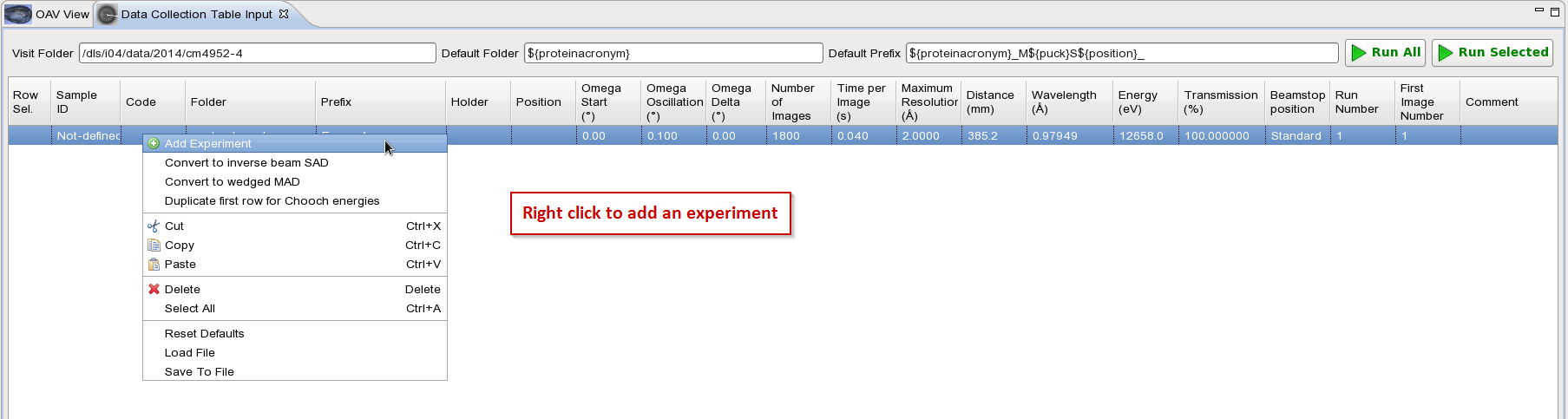
Add experiments until you have one for each energy you want to collect for your MAD data. In this example we will do 3 energies.
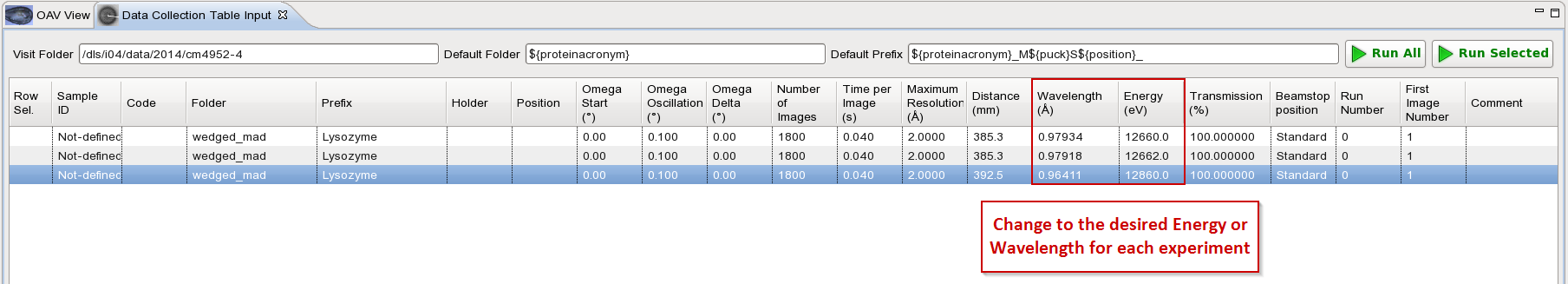
Change the wavelength or energy in the table to the desired value for each experiment:
Then convert entry into wedged MAD as shown below:
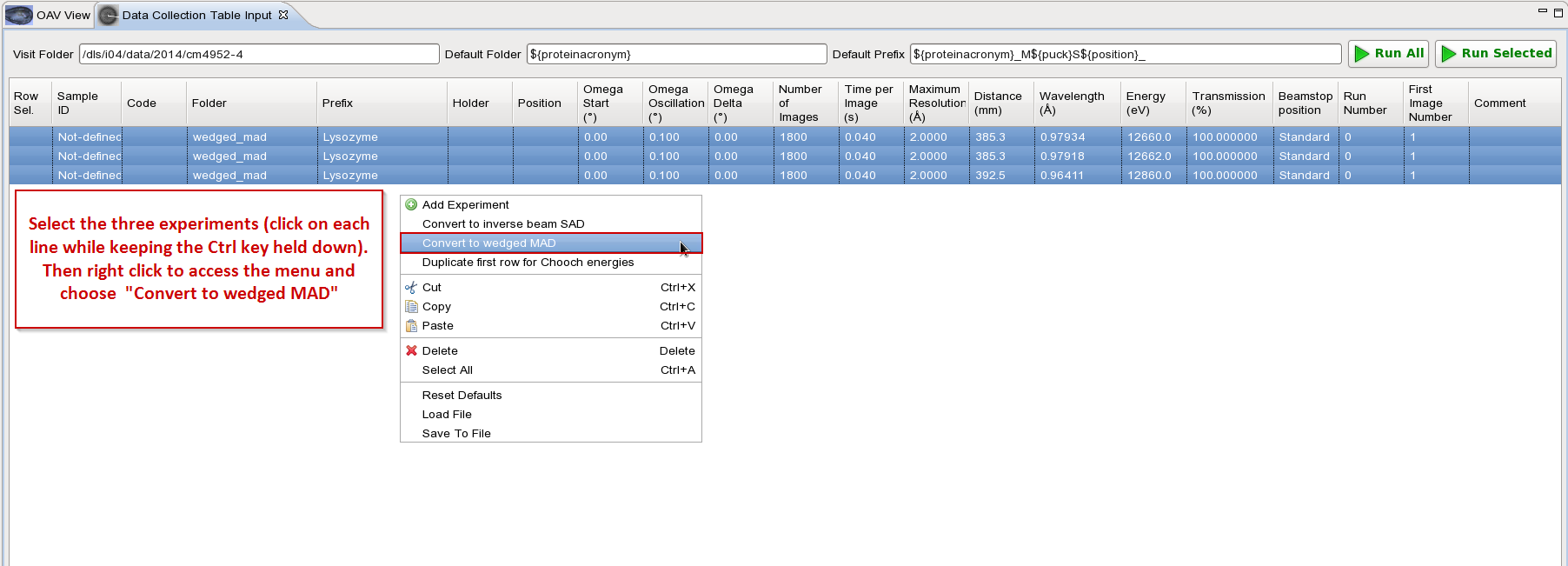
In this example we choose to do wedge of 25o(250 images with 0.1o oscillation). Note that for wedged MAD data collection we recommend to use 5o wedge or lower.
The data collection has been broken up into 25o wedges and reorganized so that each wedge is collected at 3 different energies before moving on to the next wedge. The prefix name has also been updated to indicate which energy is collected: E1, E2 or E3. To run it just click on the "Run All" button.

In this case the results will appear as 3 data sets: Lysoszyne_E1, Lysozyme_E2 and Lysozyme_E3.
Note that the wedged MAD experiment is not restricted to 3 energies.
VMXm is a micro/nanofocus MX beamline aimed at atomic structure determination in cases where the production of significant quantities of protein material and crystals is problematic.
More information
High throughput and highly automated beamline for optimised MAD and SAD experiments. Capable of accepting CL3 type experiments on crystals of pathogens.
More information
A unique facility for solving the crystallographic phase problem, using the small anomalous signals from sulphur or phosphorous which are present in native protein or RNA/DNA crystals. Additionally, anomalous difference Fourier maps can be used to locate sulphur and phosphorous positions to assist model building at low resolution and/or identify lighter atoms such as chlorine, potassium and calcium.
More information
Variable focus from 5 to 100 microns, high throughput and highly automated beamline for optimised MAD and SAD experiments.
More information
High throughput variable microfocus beamline for optimised MAD and SAD experiments on crystals down a few microns in size.
More information
The Versatile Macromolecular X-tallography in-situ (VMXi) beamline will be an entirely automated facility for characterisation of, and data collection directly from, crystallisation experiments in situ.
More informationDiamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.