
Director of the electron Bio-Imaging Centre (eBIC)
Email: [email protected]Tel: +44 (0) 1235 77 8878

Director of the electron Bio-Imaging Centre (eBIC)
Email: [email protected]
Dr Peijun Zhang obtained her Ph.D. in molecular biophysics from University Virginia, M.S. in physics and B.S. in electrical engineering from Nanjing University. She carried out postdoctoral work at the National Cancer Institute. In 2006, she joined the faculty at the University of Pittsburgh School of Medicine as an assistant professor, and was promoted to associate professor in 2012. Her research focuses on the structural and functional studies of large molecular complexes and assemblies, viruses and cellular machineries using integrated structural, biochemical and computational approaches to understand biological complexity. Dr Zhang received many awards, including the Carnegie Science Emerging Female Scientist Award, Senior Vice Chancellor’s Award, United States Department of Health and Human Services 'On-the-Spot' Award.
Her role at Diamond, as the Director of eBIC, is establishing and leading eBIC to become a world-leading center for research, expertise and training in cryo-EM and a user facility providing access to cutting-edge cryo-EM technologies. eBIC focuses on using state-of-the-art electron microscopic techniques to determine the 3D structures of molecules, cells and tissues at high resolution, as well as developing new methods and technologies to advance 3D EM imaging.
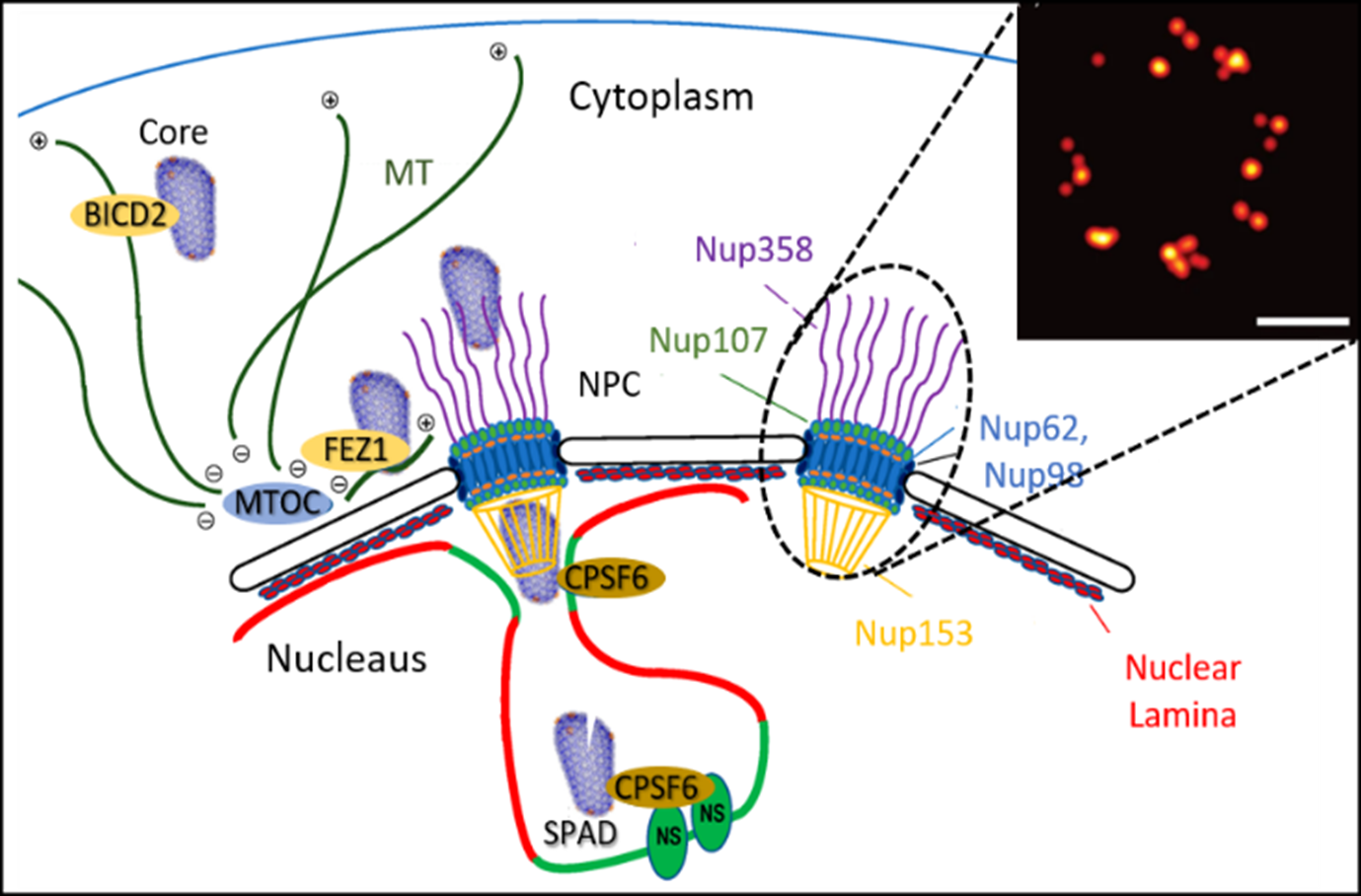
Dr Zhang's research programme is aimed to obtain an integrated, atomistic understanding of the molecular mechanisms of human virus infections. To achieve this, her team is developing novel technologies for high-resolution in situ and correlated cryo-EM/ET, combined with advanced computational methods including machine learning.
Dr Zhang's research at Diamond focuses on cryo-EM/ET technology development. Driven by the biological inquiries, Peijun's aim is to develop novel tools and methods towards 1) atomic or near-atomic resolution in situ structure determination by cryo-ET and sub-tomogram averaging; 2) single-molecule resolution 3D protein localisation in cellular tomograms of heathy and diseased cells/tissues; and 3) a robust cryoEM/ET pipeline from sample preparation, to intelligent high-throughput data acquisition, to AI-based data reduction and structure determination. These efforts will have a broad impact well beyond her own research.
Dr Zhang's research activities involve human pathogens, such as HIV-1, SARS-COV-2 and bacterial cells. In particular, her long standing interest lies in HIV-1 capsid assembly, maturation, intracellular trafficking and nuclear import, as well as its interactions with host dependency and restriction factors during the HIV-1 infection process. Another system her team have been working on is the remarkable bacterial chemotaxis sensory signalling arrays. Understanding the structural details of these challenging systems is critical for developing new antimicrobial and antiHIV/AIDS drugs.
Peer-reviewed Primary Publications (* indicates first or corresponding author)
Hou Z, Nightingale F, Zhu Y, MacGregor-Chatwin C, Zhang P* (2023) Structure of native chromatin fibres revealed by Cryo-ET in situ. Nat Commun, 10.1038/s41467-023-42072-1
Ni T, Mendonça L, Zhu Y, Howe A, Radecke J, Shah PM, Sheng Y, Krebs AS, Duyvesteyn HME, Allen E, Lambe T, Bisset C, Spencer A, Morris S, Stuart DI, Gilbert S, Zhang P* (2023) ChAdOx1 COVID vaccines express RBD open prefusion SARS-CoV-2 spikes on the cell surface. iScience 26(10):107882
Cassidy CK, Qin Z, Frosio T, Gosink K, Yang Z, Sansom MSP, Stansfeld PJ, Parkinson JS, Zhang P* (2023) Structure of the native chemotaxis core signaling unit from phage E-protein lysed E. coli cells. mBio:e0079323
Krebs AS, Liu HF, Zhou Y, Rey JS, Levintov L, Shen J, Howe A, Perilla JR, Bartesaghi A, Zhang P* (2023) Molecular architecture and conservation of an immature human endogenous retrovirus. Nat Commun 14(1):5149.
Ni T, Jiang Q, Ng PC, Shen J, Dou H, Zhu Y, Radecke J, Dykes GF, Huang F, Liu LN, Zhang P* (2023) Intrinsically disordered CsoS2 acts as a general molecular thread for α-carboxysome shell assembly. Nat Commun 14(1):5512.
Gres AT, Kirby KA, McFadden WM, Du H, Liu D, Xu C, Bryer AJ, Perilla JR, Shi J, Aiken C, Fu X, Zhang P, Francis AC, Melikyan GB, Sarafianos SG (2023) Multidisciplinary studies with mutated HIV-1 capsid proteins reveal structural mechanisms of lattice stabilization. Nat Commun 14(1):5614
Graham M, Zhang P* (2023) Cryo-electron tomography to study viral infection. Biochem Soc Trans. doi: 10.1042/BST20230103. Online ahead of print
Kleinpeter AB, Zhu Y, Mallery DL, Ablan SD, Chen L, Hardenbrook N, Saiardi A, James LC, Zhang P, Freed EO (2023) The Effect of Inositol Hexakisphosphate on HIV-1 Particle Production and Infectivity can be Modulated by Mutations that Affect the Stability of the Immature Gag Lattice. J Mol Biol. doi: 10.1016/j.jmb.2023.168037.
Waheed AA, Zhu Y, Agostino E, Naing L, Hikichi Y, Soheilian F, Yoo SW, Song Y, Zhang P, Slusher BS, Haughey NJ, Freed EO (2023) Neutral sphingomyelinase 2 is required for HIV-1 maturation. Proc Natl Acad Sci USA 120(28): e2219475120.
Pei X, Zhou L, Huang C, Boyce M, Kim JS, Liberti E, Hu Y, Sasaki T, Nellist PD, Zhang P, Stuart DI, Kirkland AI, Wang P (2023) Cryogenic electron ptychographic single particle analysis with wide bandwidth information transfer. Nat Commun 14(1):3027.
Riechmann C, Zhang P* (2023) Recent structural advances in bacterial chemotaxis signalling. Curr Opin Struct Biol. 79:102565.
Hadjidemetriou K, Kaur S, Cassidy CK, Zhang P* (2022) Mechanisms of E. coli chemotaxis signaling pathways visualized using cryoET and computational approaches. Biochem Soc Trans. 50(6):1595-1605
Yu X, Ni T, Munson G, Zhang P*, Gilbert RJC (2022) Cryo-EM structures of perforin-2 in isolation and assembled on a membrane suggest a mechanism for pore formation. EMBO J e111857.
Zhu Y, Koo CW, Cassidy CK, Spink MC, Ni T, Zanetti-Domingues LC, Bateman B, Martin-Fernandez ML, Shen J, Sheng Y, Song Y, Yang Z, Rosenzweig AC, Zhang P* (2022) Structure and activity of particulate methane monooxygenase arrays in methanotrophs. Nat Commun 13(1):5221.
Sheng Y, Morris K, Radecke J, Zhang P* (2022) Cryo-electron Tomography Remote Data Collection and Subtomogram Averaging. J Vis Exp 185. doi: 10.3791/63923.
Sheng Y, Harrison PJ, Vogirala V, Yang Z, Strain-Damerell C, Frosio T, Himes BA, Siebert CA, Zhang P, Clare DK (2022) Application of super-resolution and correlative double sampling in cryo-electron microscopy. Faraday Discuss, doi: 10.1039/d2fd00049k
Ni T, Sun Y, Burn W, Al-Hazeem MMJ, Zhu Y, Yu X, Liu LN, Zhang P* (2022) Structure and assembly of cargo Rubisco in two native α-carboxysomes. Nat Commun.13(1):4299.
Ma OX, Chong WG, Lee JKE, Cai S, Siebert CA, Howe A, Zhang P, Shi J, Surana U, Gan L (2022) Cryo-ET detects bundled triple helices but not ladders in meiotic budding yeast. PLoS One 17(4):e0266035
Krebs AS, Mendonça LM, Zhang P.* (2022) Structural Analysis of Retrovirus Assembly and Maturation. Viruses 14(1):54
Jin B, Yan F, Qi X, Cai B, Tao J, Fu X, Tan S, Zhang P, Pfaendtner J, Naser NY, Baneyx F, Zhang X, DeYoreo JJ, Chen CL. (2022) Peptoid-Directed Formation of Five-Fold Twinned Au Nanostars through Particle Attachment and Facet Stabilization. Angew Chem Int Ed Engl 61(14):e202201980
Olek M, Cowtan K, Webb D, Chaban Y, Zhang P* (2022) IceBreaker: Software for high-resolution single-particle cryo-EM with non-uniform ice. Structure 30(4):522-531
Joseph AP, Olek M, Malhotra S, Zhang P, Cowtan K, Burnley T, Winn MD (2022) Atomic model validation using the CCP-EM software suite. Acta Crystallogr D Struct Biol. 78(Pt 2):152-161
Ni T, Frosio T, Mendonça L, Sheng Y, Clare D, Himes BA, Zhang P* (2022) High-resolution in situ structure determination by cryo-electron tomography and subtomogram averaging using emClarity. Nat Protoc. 17(2):421-444
Hardenbrook N and Zhang P* (2021) A structural view of the SARS-CoV-2 virus and its assembly. Curr Opin Struct Biol. 52:123-134.
Ni T, Zhu Y, Yang Z, Xu C, Chaban Y, Nesterova T, Ning J, Böcking T, Parker MW, Monnie C, Ahn J, Perilla JR, Zhang P* (2021) Structure of native HIV-1 cores and their interactions with IP6 and CypA. Sci Adv 7(47):eabj5715.
Liu J, Alvarez FJD, Clare DK, Noel JK, Zhang P* (2021) CryoEM structure of the super-constricted two-start dynamin 1 filament. Nat Commun. 12(1):5393.
Qin Z, Zhang P* (2021) Studying bacterial chemosensory array with CryoEM. Biochem Soc Trans. 49(5):2081-2089
Mendonça L, Howe A, Gilchrist JB, Sheng Y, Sun D, Knight ML, Zanetti-Domingues LC, Bateman B, Krebs AS, Chen L, Radecke J, Li VD, Ni T, Kounatidis I, Koronfel MA, Szynkiewicz M, Harkiolaki M, Martin-Fernandez ML, James W, Zhang P* (2021) Correlative multi-scale cryo-imaging unveils SARS-CoV-2 assembly and egress. Nat Commun. 12(1):4629.
Yang Y, Liu J, Clarke BR, Seidel L, Bolla JR, Ward PN, Zhang P, Robinson CV, Whitfield C, Naismith JH. (2021) The molecular basis of regulation of bacterial capsule assembly by Wzc. Nat Commun. 12(1):4349.
Huokko T, Ni T, Dykes GF, Simpson DM, Brownridge P, Conradi FD, Beynon RJ, Nixon PJ, Mullineaux CW, Zhang P, Liu LN (2021) Probing the biogenesis pathway and dynamics of thylakoid membranes. Nat Commun. 12(1):3475.
Sarkans U, Chiu W, Collinson L, Darrow MC, Ellenberg J, Grunwald D, Hériché JK, Iudin A, Martins GG, Meehan T, Narayan K, Patwardhan A, Russell MRG, Saibil HR, Strambio-De-Castillia C, Swedlow JR, Tischer C, Uhlmann V, Verkade P, Barlow M, Bayraktar O, Birney E, Catavitello C, Cawthorne C, Wagner-Conrad S, Duke E, Paul-Gilloteaux P, Gustin E, Harkiolaki M, Kankaanpää P, Lemberger T, McEntyre J, Moore J, Nicholls AW, Onami S, Parkinson H, Parsons M, Romanchikova M, Sofroniew N, Swoger J, Utz N, Voortman LM, Wong F, Zhang P, Kleywegt GJ, Brazma A. (2021) REMBI: Recommended Metadata for Biological Images-enabling reuse of microscopy data in biology. Nat Methods. doi: 10.1038/s41592-021-01166-8.
Watanabe Y, Mendonça L, Allen E, Howe A, Lee M, Allen J, Chawla H, Pulido D, Donnellan F, Davies H, Ulaszewska,M, Belij-Rammerstorfer S, Morris S, Krebs A, Dejnirattisai W, Mongkolsapaya J, Supasa P, Screaton G, Green C, Lambe T, Zhang P*, Gilbert S, Crispin M (2021) Native-like SARS-CoV-2 spike glycoprotein expressed by ChAdOx1 nCoV-19 vaccine. ACS Cent. Sci. 7(4):594-602
Zhong Z, Ning J, Boggs E, Jang S, Wallace C, Telmer C, Bruchez M, Ahn J, Engelman A, Zhang P, Watkins S, and Ambrose Z (2021) Cytoplasmic CPSF6 regulates HIV-1 capsid trafficking and infection in a cyclophilin A-dependent manner. mBio12(2): e03142-20.
Mendonça L, Sun D, Ning J, Liu J, Kotecha A, Olek M, Frosio T, Fu X, Himes BA, Kleinpeter AB, Freed EO, Zhou J, Aiken C, Zhang P* (2021) CryoET structures of immature HIV Gag reveal a complete six-helix bundle. Commun Biol. 4(1):481.
Zhu Y, Sun D, Schertel A, Ning J, Fu X, Gwo PP, Watson AM, Zanetti-Domingues LC, Martin-Fernandez ML, Freyberg Z, Zhang P* (2021) Serial cryoFIB/SEM Reveals Cytoarchitectural Disruptions in Leigh Syndrome Patient Cells. Structure 29(1):82-87
Wilbourne M and Zhang P* (2021) Visualizing HIV-1 Capsid and Its Interactions with Antivirals and Host Factors. Viruses 13(2):246.
Bárcena M, Barnes CO, Beck M, Bjorkman PJ, Canard B, Gao GF, Gao Y, Hilgenfeld R, Hummer G, Patwardhan A, Santoni G, Saphire EO, Schaffitzel C, Schendel SL, Smith JL, Thorn A, Veesler D, Zhang P, Zhou Q. (2021) Structural biology in the fight against COVID-19. Nat Struct Mol Biol 28(1):2-7
Mendonça L, Zhang P* (2021) Analysis of Viruses in the Cellular Context by Electron Tomography, Encyclopedia of Virology (Fourth Edition) 1:242-247
Liu C., Mendonça L., Yang Y., Gao Y., Shen C., Liu J., Ni T., Ju B., Liu C., Tang X., Wei J., Ma X., Zhu Y., Liu W., Xu S., Liu Y., Yuan J., Wu J., Liu Z., Zhang Z., Liu L., Wang P., Zhang P.* (2020) The Architecture of Inactivated SARS-CoV-2 with Postfusion Spikes Revealed by CryoEM and CryoET. Structure 28(11):1218-1224
Sutton G., Sun D., Fu X., Kotecha A., Hecksel C.W., Clare D.K., Zhang P.*, Stuart D.I., Boyce M. (2020) Assembly intermediates of orthoreovirus captured in the cell. Nat Commun 11, 4445.
Ni T., Gerard S., Zhao G., Dent K., Ning J., Zhou J., Shi J., Anderson-Daniels J., Li W., Jang S., Engelman AN., Aiken C., Zhang P.* (2020) Intrinsic curvature of HIV-1 CA hexamer underlies capsid topology and interaction with cyclophilin A. Nat Struct Mol Biol 27(9):855-862
Beale E.V., Waterman D.G., Hecksel C., van Rooyen J., Gilchrist J.B., Parkhurst J.M., de Haas F., Buijsse B., Evans G., Zhang P.* (2020) A Workflow for Protein Structure Determination from Thin Crystal Lamella by Micro-Electron Diffraction. Front Mol Biosci 7:179.
Zhou L., Song J., Kim J., Pei X., Huang C., Boyce M., Mendonca L., Clare D., Siebert A., Allen C., Liberti E., Stuart D.I., Pan X., Nellist P., Zhang P., Kirkland A., and Wang P. (2020) Low-Dose Phase Retrieval of Biological Specimens using Cryo-Electron Ptychography. Nat Commun 11 (1):2773.
Sun D., Varlakhanova N.V., Tornabene B.A., Ramachandran R., Zhang P.*, Ford M.G.J. (2020) The cryo-EM structure of the SNX-BAR Mvp1 tetramer. Nat Commun 11(1):1506.
Ni T., Jiao F., Yu X., Aden S., Ginger L., Williams S.I., Bai F., Pražák V., Karia D., Stansfeld P., Zhang P., Munson G., Anderluh G., Scheuring S., Gilbert R.J.C. (2020) Structure and mechanism of bactericidal mammalian perforin-2, an ancient agent of innate immunity. Sci Adv 6(5):eaax8286.
Cassidy C.K., Himes B.A., Sun D., Ma J., Zhao G., Parkinson J.S., Stansfeld P.J., Luthey-Schulten Z. and Zhang P.* (2020) Structure and dynamics of the E. coli chemotaxis core signaling complex by cryo-electron tomography and molecular simulations. Commun Biol. 3(1):24. doi:10.1038/s42003-019-0748-0
Alvarez F.J.D., Zhang P.* (2020) Purification and Characterization of MxB. Methods Mol Biol. 2159:55-65.
Fu X., Ning J., Zhong Z., Ambrose Z., Watkins S.C. and Zhang P.* (2019) AutoCLEM: An Automated Workflow for Correlative Live-Cell Fluorescence Microscopy and Cryo-Electron Tomography. Sci Rep 9, 19207 doi:10.1038/s41598-019-55766-8
Kim J.Y., Yeom J., Zhao G., Calcaterra H., Munn J., Zhang P., Kotov N. (2019) Assembly of Gold Nanoparticles into Chiral Superstructures Driven by Circularly Polarized Light. J Am Chem Soc. 141(30):11739-11744
Xiao J., Liu M., Qi Y., Chaban Y., Gao C., Pan B., Tian Y., Yu Z., Li J., Zhang P., Xu Y. (2019) Structural insights into the activation of ATM kinase. Cell Res. 29(8):683-685
Zhang P.* (2019) Advances in cryo-electron tomography and subtomogram averaging and classification. Curr Opin Struct Biol. doi:10.1016/j.sbi.2019.05.021.
Himes B.A. and Zhang P.* (2018) emClarity: Software for High Resolution Cryo-electron Tomography and Sub-tomogram Averaging. Nat Methods 15(11):955-961
Siegmund S.E., Grassucci R., Carter S.D., Barca E., Farino Z.J., Juanola-Falgarona M., Zhang P., Tanji K., Hirano M., Schon E.A., Frank J., Freyberg Z. (2018) Three-Dimensional Analysis of Mitochondrial Crista Ultrastructure in a Patient with Leigh Syndrome by In Situ Cryoelectron Tomography. iScience 6:83-91.
Duyvesteyn H.M.E., Kotecha A., Ginn H.M., Hecksel C.W., Beale E.V., de Haas F., Evans G., Zhang P., Chiu W., Stuart D.I. (2018) Machining protein microcrystals for structure determination by electron diffraction. Proc Natl Acad Sci USA 115(38):9569-9573
Varlakhanova N.V., Alvarez F.J.D., Brady T.M., Tornabene B.A., Hosford C.J., Chappie J.S., Zhang P., Ford M.G.J. (2018) Structures of the fungal dynamin-related protein Vps1 reveal a unique, open helical architecture. J Cell Biol 217(10):3608-3624
Ning J., Zhong Z., Fischer D.K., Harris G., Watkins S.C., Ambrose Z., Zhang P.* (2018) Truncated CPSF6 forms higher order complexes that bind and disrupt HIV-1 capsid. J Virol 92(13). pii: e00368-18
Tao C., Liu Y., Sun R., Zhang B., Qi L., Shivakoti S., Tian C., Zhang P., Lau P., Zhou Z.H., Bi G. (2018) Differentiation and characterization of excitatory and inhibitory synapses by cryo-electron tomography and correlative microscopy. J Neurosci 38(6):1335-1350, Featured on the cover of J Neurosci.
Wang M., Quinn C.M., Perilla J.R., Zhang H., Shirra Jr. R., Hou G., Byeon I.J., Suiter C.L., Ablan S., Urano E., Nitz T.J., Aiken C., Freed E.O., Zhang P., Schulten K., Gronenborn A.M., Polenova T. (2017) Quenching protein dynamics interferes with HIV capsid maturation. Nat Commun 8(1):1779
Alvarez F.J.D., He S., Perilla J.R., Jang S., Schulten K., Engelman A.N., Scheres S.H.W., Zhang P.* (2017) CryoEM structure of MxB reveals a novel oligomerization interface critical for HIV restriction. Sci Adv 3(9):e1701264
Cassidy C.K., Himes B.A., Luthey-Schulten Z., Zhang P.* (2017) CryoEM-based Hybrid Modeling Approaches for Structure Determination. Curr Opin Micro Biol 43:14-23
Perilla J.R., Zhao G., Lu M., Ning J., Hou G., Byeon I.L, Gronenborn A.M., Polenova T., Zhang P.* (2017) CryoEM Structure Refinement by Integrating NMR Chemical Shifts with Molecular Dynamics Simulations. J Phys Chem B 121(15):3853-3863
Yang M., Chan H., Zhao G., Bahng J.H., Zhang P.*, Petr Král P., Kotov N.A. (2017) Self-assembly of Nanoparticles into Biomimetic Capsid-like Nanoshells. Nat Chem 9(3):287-294.
Clare D.K., Siebert C.A., Hecksel C., Hagen C., Mordhorst V., Grange M., Ashton A.W., Walsh M.A., Grünewald K., Saibil H.R., Stuart D.I., Zhang P. (2017) Electron Bio-Imaging Centre (eBIC): the UK national research facility for biological electron microscopy. Acta Crystallogr D Struct Biol 9(1):65
Ning J., Erdemci-Tandogan G., Yufenyuy E.L., Wagner J., Himes B.A., Zhao G., Aiken C., Zandi R., Zhang P.* (2016) In vitro Protease Cleavage and Computer Simulations Reveal the HIV-1 Capsid Maturation Pathway. Nat Commun 7:13689
Merg A.D., Boatz J.C., Mandal A., Zhao G., Mokashi-Punekar S., Liu C., Wang X., Zhang P., van der Wel P.C.A., Rosi N.L. (2016) Peptide-Directed Assembly of Single-Helical Gold Nanoparticle Superstructures Exhibiting Intense Chiroptical Activity. J Am Chem Soc 138 (41) 13655-13663
Liu C., Perilla J. R., Ning J., Lu M., Hou G., Ramalho R., Bedwell G., Byeon I., Ahn J., Gronenborn A. M., Prevelige P., Rousso I., Aiken, C., Polenova T., Schulten K., Zhang P.* (2016) Cyclophilin A Stabilizes the HIV-1 Capsid through a Novel Non-canonical Binding Site. Nat Commun 7: 10714.
Sun J., Chen Y., Li K., Huang Y., Fu X., Zhang X., Zhao W., Wei Y., Xu L., Zhang P., Venkataramanan R., Li S. (2016) A Prodrug Micellar Carrier Assembled from Polymers with Pendant Farnesyl Thiosalicylic Acid Moieties for Improved Delivery of Paclitaxel. Acta Biomater. 43:282-91.
Tillman T.S., Alvarez F.J., Reinert N.J., Liu C., Wang D., Xu Y., Xiao K., Zhang P., Tang P. (2016) Functional Human α7 Nicotinic Acetylcholine Receptor (nAChR) Generated from E. coli. J Biol Chem. 291(35):18276-82.
Lu M., Hou G., Zhang H., Suiter C.L., Ahn J., Byeon I.L., Perilla J.R., Langmead C., Hung I., Gorkov P.L., Gan Z., Brey W., Aiken C., Zhang P., Schulten K., Gronenborn A.M., Polenova T. (2015) Dynamic Allostery Governs Cyclophylin A - HIV Capsid Interplay. Proc Natl Acad Sci USA 112(47):14617-22.
Cassidy C. K, Himes B. A., Alvarez F. J., Ma J., Zhao G., Perilla J. R., Schulten K., Zhang P.* (2015) CryoEM and Computer Simulations Reveal a Novel Kinase Conformational Switch in Bacterial Chemotaxis Signaling. Elife. 2015 Nov 19;4. pii: e08419. doi: 10.7554/eLife.08419.
Lu J., Liu C., Wang P., Ghazwani M., Xu J., Huang Y., Ma X., Zhang P., Li S. (2015) The Self-assembling Camptothecin-tocopherol Prodrug: An Effective Approach for Formulating Camptothecin. Biomaterials 62:176-87.
Yeom J., Yeom B., Chan H., Smith K.W., Dominguez-Medina S., Bahng J.H., Zhao G., Chang W., Chang S.J., Chuvilin A., Melnikau D., Rogach A.L., Zhang P., Link S., Král P., Kotov N.A. (2015) Chiral Templating of Self-Assembling Nanostructures by Circularly Polarized Light. Nat Mater 14(1):66-72.
Fu X., Himes B., Ke D., Rice W.J., Ning J., Zhang P.* (2014) Controlled Bacterial Lysis for Electron Tomography of Native Cell Membranes. Structure 22(12):1875-82.
Fribourgh J.L., Nguyen H.C., Matreyek K.A., Alvarez F.J., Summers B.J., Dewdney T.G., Aiken C., Zhang P., Engelman A., Xiong Y. (2014). Structural Insight into HIV-1 Restriction by MxB. Cell Host & Microbe 16(5):627-38.
Saini S.G., Liu C., Zhang P., Lee T.H. (2014). Membrane Tethering by the Atlastin GTPase Depends on GTP Hydrolysis But Not on Forming the Crossover Configuration. Mol Biol Cell 25(24):3942-53.
Park J., Nguyen T.D., Silveira G.Q., Bahng J.H., Srivastava S., Zhao G., Sun K., Zhang P., Sharon C. Glotzer S.C., Kotov N.A. (2014). Terminal Supraparticle Assemblies from Similarly Charged Protein Molecules and Nanoparticles. Nat Commun 5:3593.
Schirra R.T. Jr and Zhang P.* (2014) Correlative Fluorescence and Electron Microscopy. Curr Protoc Cytom. 70:12.36.1–12.36.10.
Hickey R.J., Koski J., Meng X., Riggleman R.A., Zhang P., Park S.J. (2014) Size-controlled Self-assembly of Superparamagnetic Polymersomes. ACS Nano 8(1):495-502.
Zhao G. and Zhang P.* (2014) CryoEM Analysis of Capsid Assembly and Structural Changes upon Interactions with a Host Restriction Factor, TRIM5α. Methods Mol Biol 1087:13-28.
Zhao G., Perilla J.R., Yufenyuy E.L., Meng X., Chen B., Ning J., Ahn J., Gronenborn A.M., Schulten K., Aiken C., Zhang P.* (2013) Mature HIV-1 Capsid Structure by Cryo-electron Microscopy and All-atom Molecular Dynamics. Nature 497(7451):643-6. Featured on the cover of Nature.
Zhang P.* (2013) Correlative Cryo-electron Tomography and Optical Microscopy of Cells. Curr Opin Struct Biol. 23(5):763-70.
Zhang P., Huang Y., Makhov A.M., Gao X., Zhang P., Li S. (2013). Characterization of Spherulites as a Lipidic Carrier for Low and High Molecular Weight Agents. Pharm Res. 30(6):1525-35.
Mowrey D., Cui T., Jia Y., Ma D., Makhov A.M., Zhang P., Tang P., Xu Y. (2013). Open-Channel Structures of the Human Glycine Receptor α1 Full-Length Transmembrane Domain. Structure 21(10):1897-904.
Song C., Blaber M.G., Zhao G., Zhang P., Fry H.C., Schatz G.C., Rosi N.L. (2013) Tailorable Plasmonic Circular Dichroism Properties of Helical Nanoparticle Superstructures. Nano Letter 13(7):3256-61.
Hickey R.J., Meng X., Zhang P., Park S.J. (2013) Low-dimensional Nanoparticle Clustering in Polymer Micelles and Their Transverse Relaxivity Rates. ACS Nano 7(7):5824-5833.
Aiken C. and Zhang P. (2013) HIV-1 Maturation. Advances in HIV-1 Assembly and Release:153-166.
Jun S., Zhao G., Ning J., Gibson G., Watkins S., Zhang P.* (2013) Correlative Microscopy for 3D Structural Analysis of Dynamic Interactions. J Vis Exp. 76: e50386.
Gao X., Huang Y., Makhov A.M., Epperly M., Lu J., Grab S., Zhang P., Rohan L., Xie X., Wipf P., Greenberger J., Li S. (2013) Nano-assembly of Surfactants with Interfacial Drug-Interactive Motifs as Tailor-Designed Drug Carriers. Mol. Pharmaceut.10(1):187-198.
Yang Y., Bhatti A., Ke D., Gonzalez-Juarrero M., Lenaerts A., Kremer L., Guerardel Y., Zhang P. and Ojhaa A.K.(2013) Exposure to a Cutinase-like Serine Esterase Triggers Rapid Lysis of Multiple Mycobacterial Species. J. Biol. Chem. 288(1):382-392.
Zhang P.*, Meng X., Zhao G. (2013) Tubular Crystals and Helical Arrays: Structural Determination of HIV-1 Capsid Assemblies Using Iterative Helical Real-Space Reconstruction. Methods Mol Biol 955: 381-399.
Yang H., Ji X., Zhao G., Ning J., Zhao Q., Aiken C., Gronenborn A.M., Zhang P., Xiong Y. (2012) Structural Insight into HIV-1 Capsid Recognition by Rhesus TRIM5α. Proc Natl Acad Sci USA 109(45): 18372-18377.
Wang K., Strunk K., Zhao G., Gray J.L., Zhang P.* (2012) 3D Structure Determination of Native Mammalian Cells using Cryo-FIB and Cryo-electron Tomography. J Struct Biol 180(2): 318-326.
Meng X., Zhao G., Yufenyuy E., Ke D., Ning J., DeLucia M., Ahn J., Gronenborn A.M., Aiken C.*, Zhang P.* (2012) Protease Cleavage Leads to Formation of Mature Trimer Interface in HIV-1 Capsid. PLoS Pathog 8(8): e1002886.
Strunk K., Wang K., Ke D., Gray J.L., Zhang P.* (2012) Thinning of Large Mammalian Cells for Cryo-TEM Characterization by Cryo-FIB Milling. J. Microscopy 247(3):220-227.
Jun S., Ke D., Debiec K., Zhao G., Meng X., Ambrose Z., Gibson G.A., Watkins S.C., Zhang P.* (2011) Direct Visualization of HIV-1 with Correlative Live-Cell Microscopy and Cryo-Electron Tomography. Structure 19(11):1573-1581.
Zhao G., Ke D., Vu T., Ahn J., Shah V.B., Yang R., Aiken C., Charlton L.M., Gronenborn A.M., Zhang P.* (2011) Rhesus TRIM5alpha Disrupts the HIV-1 Capsid at the InterHexamer Interfaces. PLoS Pathog 7(3):e1002009.
Meng X., Zhao G., Zhang P.* (2011) Structure of HIV-1 Capsid Assemblies by Cryo-electron Microscopy and Iterative Helical Real-space Reconstruction. J Vis Exp (54), e3041.
Ahn J., Novince Z., Concel J., Byeon C., Makhov A.M., Byeon I.L., Zhang P., Gronenborn A.M. (2011) The Cullin-RING E3 Ubiquitin Ligase CRL4-DCAF1 Complex Dimerizes via a Short Helical Region in DCAF1. Biochemistry 50(8): 1359-1367.
Morin-Leisk J., Saini S.G., Makhov A.M., Meng X., Zhang P., Lee T.H. (2011) An Intra-molecular Salt Bridge Drives the Soluble Domain of GTP-bound Atlastin into the 'Post-fusion' Conformation. J. Cell Biol. 195(4):605-615.
Du S., Betts L., Yang R., Shi H., Concel J., Ahn J., Aiken C., Zhang P., Yeh J.I. (2011) Structure of the HIV-1 Full-Length Capsid in a Conformationally-Trapped Unassembled State Induced by Small-molecule Binding. J. Mol. Biol. 406(3):371-386.
Hwang L., Zhao G., Zhang P., Rosi N.L. (2011) Size-controlled Peptide-directed Synthesis of Hollow Spherical Gold Nanoparticle Superstructures. Small 7(14):1939-1942.
Song C., Zhao G., Zhang P., Rosi N.L. (2010) Expeditious Synthesis and Assembly of Spherical Gold Nanoparticle Superstructures. J. Am. Chem. Soc. 132(40):14033-14035.
Byeon I.L., Meng X, Jung J., Zhao G., Yang R., Shi J., Ahn J., Concel J., Aiken C., Zhang P.*, Gronenborn A.M. (2009) Structural Convergence between CryoEM and NMR Reveals Novel Intersubunit Interactions Critical for HIV-1 Capsid Assembly and Function. Cell 139: 780–790.
Chen C., Zhang P., Rosi N.L. (2008) A New Peptide-Based Method for the Design and Synthesis of Nanoparticle Superstructures: Construction of Highly-Ordered Gold Nanoparticle Double-Helices. J. Am. Chem. Soc. 130(41):13555-13557.
Khursigara C.M., Wu X., Zhang P.*, Lefman J., Subramaniam S. (2008) Role of HAMP Domains in Chemotaxis Signaling by Bacterial Chemoreceptors. Proc Natl Acad Sci USA 105(43):16555-16560.
Zhang P.*, Khursigara C.M., Hartnell L.M., Subramaniam S. (2007) Direct Visualization of E.coli Chemotaxis Receptor Arrays using Cryo-electron Microscopy. Proc Natl Acad Sci USA 104(10):3777-3781.
Zhang P.*, Weis R.M., Peters P.J., Subramaniam S. (2007) Electron Tomography of Bacterial Chemotaxis Receptor Assemblie. Methods Cell Biol. 79:373-384.
Zhang P.*, Juliani J., Lefman J., Land W., Smith S., Lee S., Leapman R., Kessel M., Rouault T., Subramaniam S. (2005) Electron Tomography of Degenerating Neurons in Mice with Abnormal Regulation of Iron Metabolism. J. Struct. Biol. 150(2):144-153.
Lefman J, Zhang P.*, Hirai T., Weis R.M., Juliani J., Bliss D., Kessel M., Bos E., Peters P.J., Subramaniam S. (2004) Three-Dimensional Electron Microscopic Imaging of Membrane Invaginations in Escherichia coli Overproducing the Chemotaxis Receptor Tsr. J. Bacteriol. 186(15):5052-5061.
Zhang P.*, Bos E., Heymann J., Gnaegi H, Kessel M, Peters P.J., Subramaniam S. (2004) Direct Visualization of Receptor Arrays in Frozen-hydrated Cells and Sections from Bacteria Overproducing the Chemotaxis Receptor Tsr. J. Microscopy 216(1):76-83.
Chen Y., Zhang P., Egelman E.H., Hinshaw J.E. (2004) The Stalk Region of Dynamin Drives the Constriction of Dynamin Tubes. Nature Struct. Mol. Biol. 11(6):574-575.
Borgnia M.J., Shi D., Zhang P., Subramaniam S., Milne J.L. (2004) Visualization of Alpha-helical Features in a Density Map Constructed using 9 Molecular Images of the 1.8 MDa Icosahedral Core of Pyruvate Dehydrogenase. J. Struct. Biol. 147(2):136-145.
Shen W., Zhang P., German D., Rouault T., Subramaniam S. (2004) A Template-propagation Method for Segmentation of Filamentous Structures in Electron Tomograms. IEEE Int. Symp. Biomed. Imaging 1:1-4.
Zhang P.*, Borgnia M.J., Mooney P., Shi D., Pan M., O'Herron P., Mao A., Brogan D., Milne J.L., Subramaniam S. (2003) Automated Image Acquisition and Processing using a New Generation of 4K x 4K CCD Cameras for Cryo Electron Microscopic Studies of Macromolecular Assemblies. J. Struct. Biol. 143(2):135-144.
Zhang P.* and Hinshaw J.E. (2001) Three-dimensional Reconstruction of Dynamin in the Constricted State. Nat Cell Biol. 3:922-926.
Zhang P.*, Beatty A., Milne J.L., Subramaniam S. (2001). Automated Data Collection with a Tecnai 12 Electron Microscope: Applications for Molecular Imaging by Cryo-microscopy. J. Struct. Biol. 135:251-261.
Young H.S., Xu C., Zhang P., Stokes D.L. (2001) Locating the Thapsigargin Binding Site on Ca(2+)-ATPase by Cryoelectron Microscopy. J. Mol. Biol. 308(2): 231-240.
Stokes D.L., Auer M., Zhang P., Kuhlbrandt W. (1999) Comparison of H+-ATPase and Ca2+-ATPase Suggests that a Large Conformational Change Initiates P-type Ion Pump Reaction Cycles. Curr. Biol. 9(13):672-679.
Zhang P.*, Toyoshima C., Yonekura K., Green M.N., Stokes D.L. (1998) Structure of the Calcium Pump from Sarcoplasmic Reticulum at 8Å Resolution. Nature 392:835-839.
Stokes D.L., Zhang P., Toyoshima C., Yonekura K., Ogawa H., Lewis M.R., Shi D. (1998) Cryoelectron Microscopy of the Calcium Pump from Sarcoplasmic Reticulum: Two Crystal Forms Reveal Two Different Conformations. Acta Physiol Scand Suppl 643:35-43.
Luthin D.R., Lee K.S., Okonkwo D., Zhang P., Linden J. (1995) Photoaffinity Labeling with 2(-)[2-(4-azido-3(-)[125I]- iodophenyl)ethylamino]adenosine and Autoradiography with 2(-)[2-(4-amino-3(-)[125I]iodophenyl)ethylamino]adenosine of A2a Adenosine Receptors in Rat Brain. J. Neurochemistry 65:2072-2079.
Zheng J. G., Li Q., Zhang P., Feng D. (1992) A TEM Study of One-dimensional Incommensurate Modulation on (001) Plane of Single Crystal Bi2Sr2CaCu2Oy. Supercon. Sci. Technol. 5:472-475.
Zheng J. G., Li Q., Zhang P., Wang L., Wei M., Feng D. (1991) HREM Studies on Stacking Faults in YBa2Cu3O7-y. Progress in Natural Science 1(4):323-327.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.