Beamline phone numbers:
+44 (0) 1235 567594 (EC1 - main)
+44 (0) 1235 567499 (EC1 - DECT)
+44 (0) 1235 567499 (EC1 - DECT)
+44 (0) 1235 777596 (CC3)
Tel: +44 (0) 1235 778757
E-mail: [email protected]
Email: [email protected]
Tel: +44 (0)1235 778518
B24 is a correlative cryo-imaging beamline offering 3D imaging with soft X-ray tomography (cryoSXT) complemented by super resolution fluorescence structured illumination microscopy (cryoSIM). All associated sample preparation and evaluation as well as post-data collection processing are also available at the beamline (see sections below).
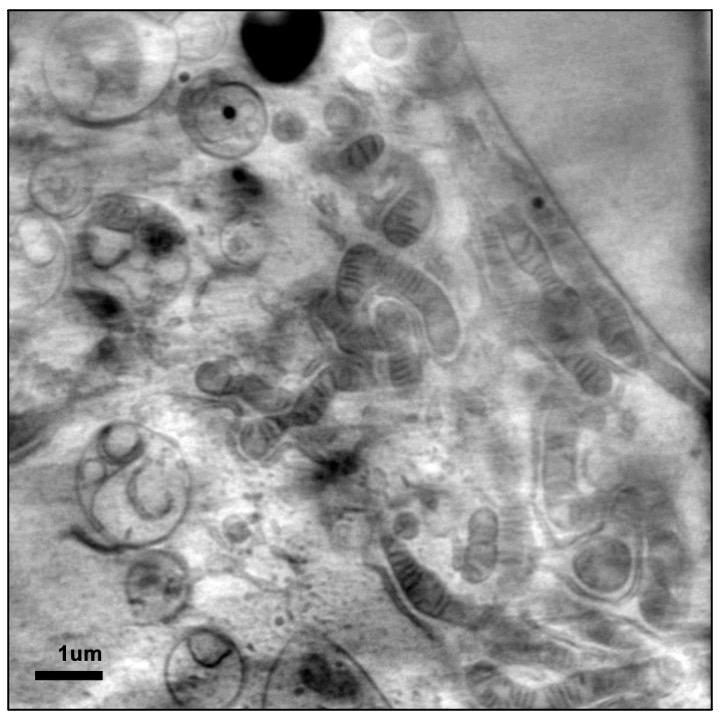
CryoSXT offers us the opportunity to image cells and cell populations in 3D to a resolution of 25 nm. To put this in context, ribosomes are only 20-30 nm across while mitochondria can be anywhere between 500 to 3000 nm. This places SXT well within capacity to deliver clear cellular imaging and, given the sizeable fields of view achieved (anywhere between 10 to 20 μm), document ultrastructure through swathes of intracellular and pericellular space in 3D. SXT imaging is done through absorption contrast in the water window of X-ray light where carbon-rich biological structures obstruct light as it passes through them as opposed to the oxygen-rich surrounding medium that does not. As a result, the impression left on a detector when an image is taken is a negative projection of the cell structure exactly like in the case of medical X-ray imaging. In fact, in concept and optical implementation this method is the cellular equivalent of a full body CT scan in a medical setting only adapted for the microworld of cells. Because imaging depends on cellular content alone, there is no requirement for contrast enhancing chemicals to be added and therefore the information captured has not been altered in any way by foreign material. An additional benefit comes from the fact that to help cells endure sustained exposure to X-ray light during imaging we snap freeze them before use, thereby perfectly preserving the exact instant of their lives that is of interest without altering any of the structures that we are hoping to see.

Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.