All proteins added to UAS and approved are automatically copied across to ISPyB. To register a protein it must first be added to UAS and approved - follow this link for how to add samples in UAS.
Only proteins registered and approved in UAS can be assigned for shipping and data collection. These are summarised on the proteins page which can be acessed by selecting proteins from the proposal dropdown menu in ISPyB:
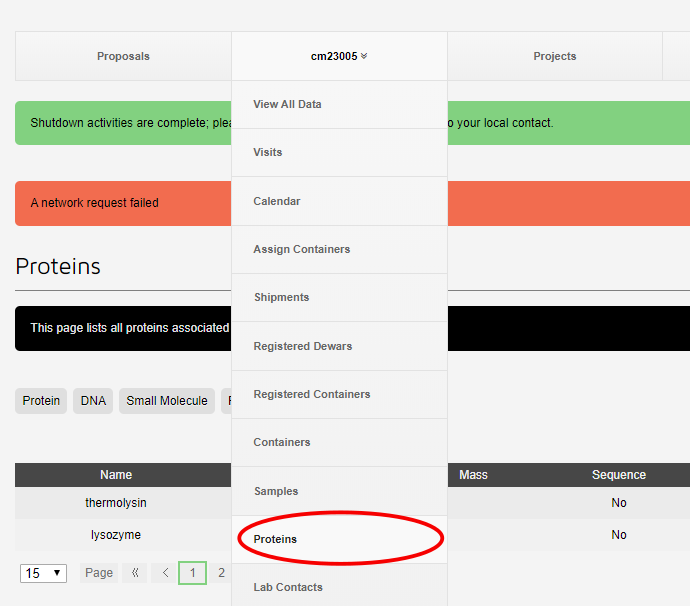
Those which have been approved are green on the proteins page. You can filter these by protein name to make finding your protein easier.
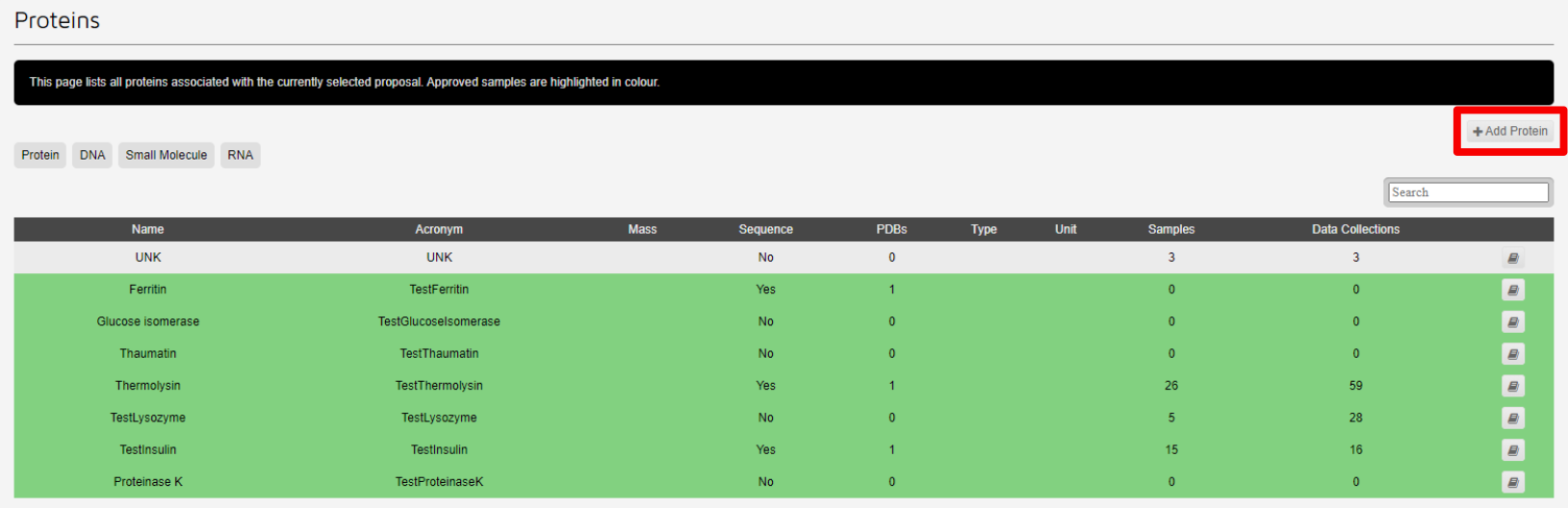
Failing to match protein names between UAS and ISPyB will result in samples not being available for the creation of a new container.
You may wish to define a new protein based on a previously approved sample, e.g. a selenomethionine derivative or point mutant. In this case you should clone the original approved sample as described below. This will allow you to define a new protein acronym but preserve other information such as sequence, pdb file etc. Only approved samples can be cloned.
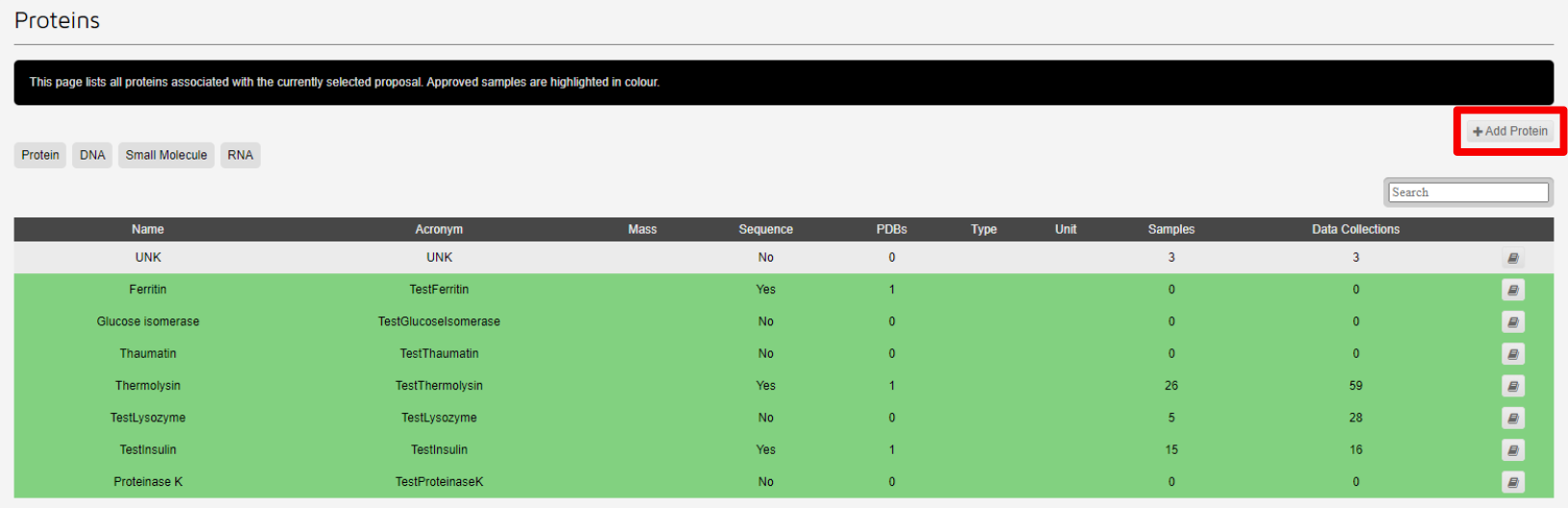
Samples highlighted in green have matching details supplied in UAS. These samples can be cloned.
To clone a protein, choose the original approved protein you wish to clone and click on it to bring up the detail page.
This brings up a page summarising the protein. If the sample is approved, then the 'Clone Protein' button will be displayed at the top right of this page.
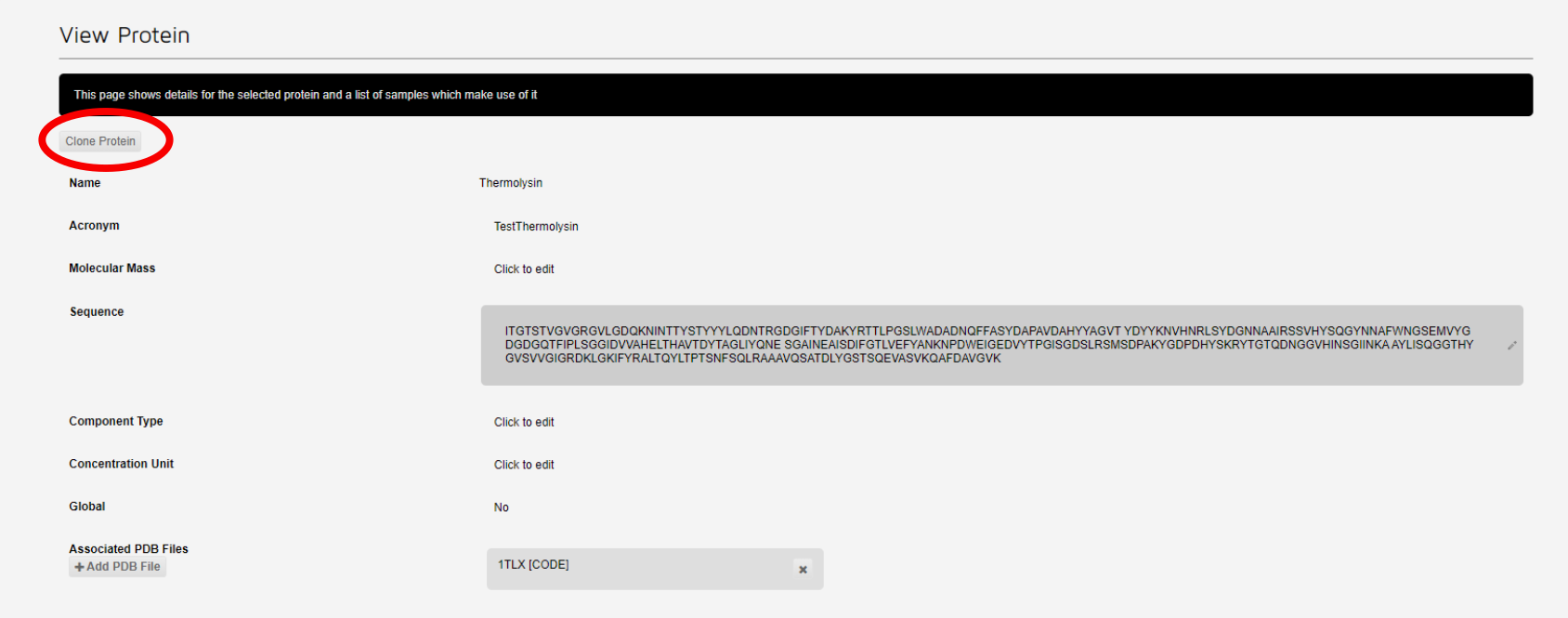
Click 'Clone Protein' to copy the protein to a new entry. This will display a form prefilled with the original protein information. You may edit the protein acronym, pdb file, sequence etc here. When you are finished, click the 'Add Protein' button to create your new protein.
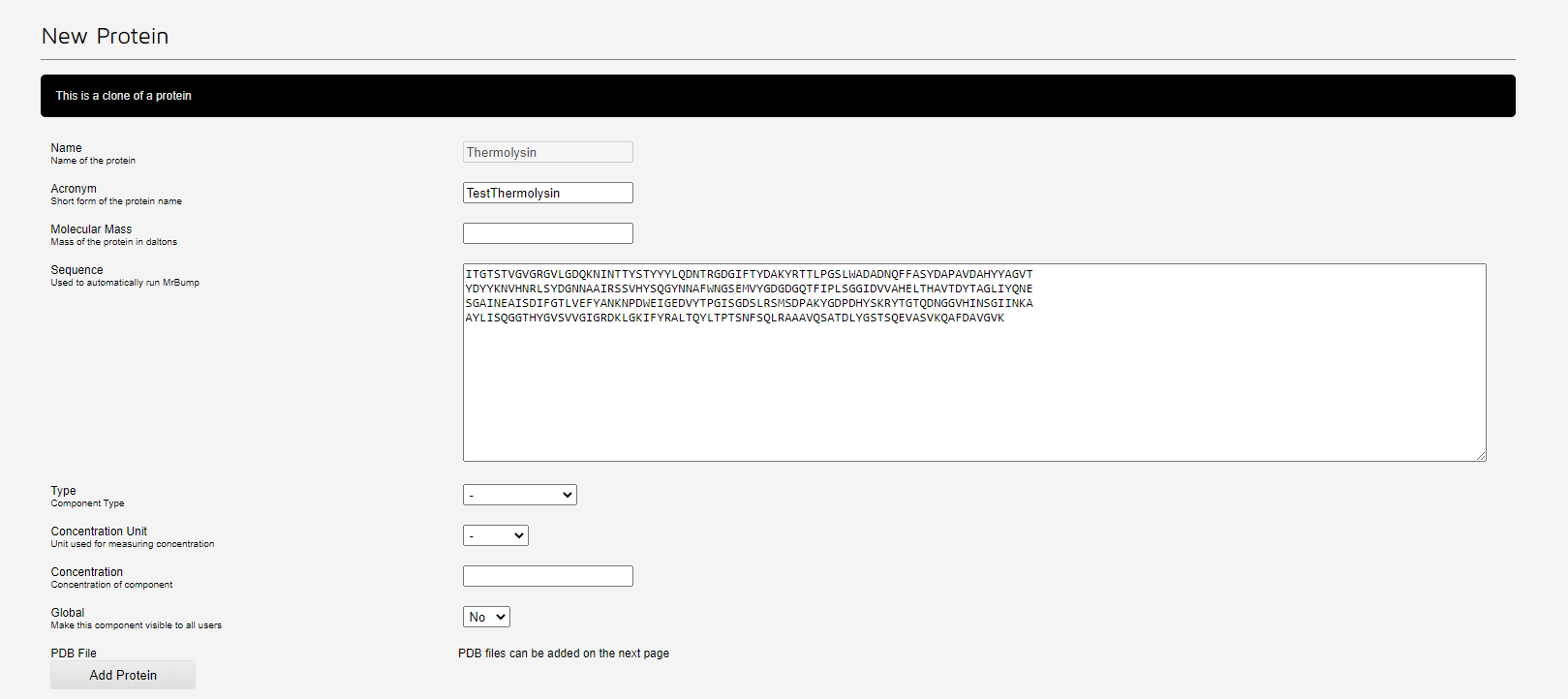
Supplying the sequence will allow automated molecular replacement (MrBump) to be run. Supplying a PDB file, which can be done once the protein is added, will allow automated refinement (DIMPLE) to be run. The result of these can be obtained after data collection on ISPyB as part of the automatic software pipelines.
Sample specific conditions such as anomalous scatterers can be defined within sample registration.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.