Antibodies can work together to combat malaria
Apr 25, 2022
Apr 25, 2022
It doesn't take much to contract malaria. A quick bite from an infected mosquito transfers the Plasmodium parasite to a human body. First, it heads to the liver, where it grows and multiplies. Then it moves on to infect red blood cells, hiding where the immune system can't reach it, multiplying and releasing new waves of parasites. Each one can infect a neighbouring red blood cell in under 20 seconds. The immune system doesn't stand much of a chance against such a rapid attack, so there were around 241 million malaria cases in 2020, resulting in an estimated 627,000 deaths. In the face of such a deadly parasite, the race is on to develop effective vaccines that can induce a more potent antibody response, in terms of both quantity and quality of antibodies. Researchers from the Higgins Lab at the University of Oxford have been working on the problem, using macromolecular crystallography to discover the structure of key proteins. In work recently published with the Draper lab in Nature Communications, they've found that different antibodies can work together, a synergistic effect that could be crucial to developing effective malaria vaccines.
Nearly half of the world's population is at risk from malaria, with most cases and deaths occurring in Africa, in children under five. If we can develop vaccines to prevent malaria parasites from infecting red blood cells, we can prevent the symptoms from arising and stop transmission of the disease. Whenever we are vaccinated against a disease, our immune system develops a broad range of antibodies. In the case of malaria, some of these antibodies will be effective at neutralising the parasite, but others will not. Some may even interfere with the neutralising antibodies, making the immune response less effective. Therefore, the goal is to develop good vaccines that promote the development of large quantities of high-quality antibodies that can rapidly bind to the parasite and prevent it from invading red blood cells.
Plasmodium falciparum and Plasmodium vivax are the two major parasites that cause human malaria. Plasmodium falciparum relies on a protein called cysteine-rich protective antigen (CyRPA) to invade red blood cells, making CyRPA a leading blood-stage malaria vaccine candidate. Researchers from the Higgins and Draper Labs at the University of Oxford investigated a panel of anti-CyRPA monoclonal antibodies (mAbs) that strongly inhibited parasite growth in in vitro tests.
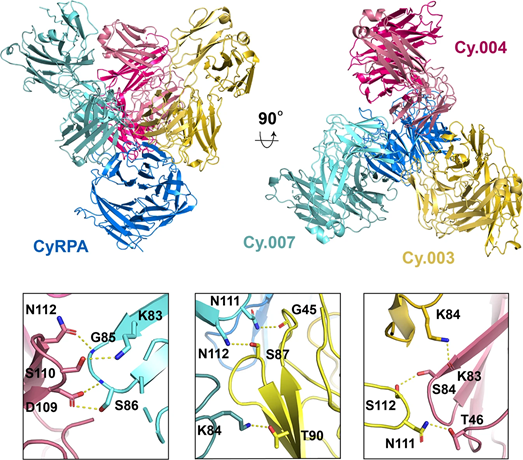
The team has BAG Access to Diamond and used its remote access protocols to send in samples for analysis using macromolecular crystallography (MX) on beamlines I03 and I04. For this work, they also conducted experiments at the Swiss Light Source and the Soleil synchrotron in France. Their structural analysis of the mAbs showed that they all bind to the same face of the CyRPA protein, which is vital information for future vaccine design.
The results also showed that these antibodies have another trick up their sleeve. Different antibodies bind to different sites on CyRPA, meaning that more than one can bind to the protein at a time. What's new is discovering that these distinct antibodies also bind to each other, helping one another bind more tightly and working together to have more neutralising power. This synergy has previously been seen in antibodies which bind to repeating structures, where multiple identical antibodies can bind to their target while also interacting with each other. Although this is the first time this mechanism of synergy has been seen for different antibodies, the team says that there's nothing special about these antibodies, and it could be a widespread effect, seen for antibodies against many different pathogens.
Prof Matt Higgins says:
We can imagine CyRPA as a Trivial Pursuit playing piece, with six wedges. All of the good antibodies we looked at bind to two neighbouring wedges. Knowing that, we can design a vaccine that just has those two wedges and use this smaller molecule to induce just the good antibodies. That may also help to create the synergy, as it's possible that less good antibodies could interfere with the antibody-antibody binding process.
CyRPA isn't the only good candidate for a blood-stage malaria vaccine targeting Plasmodium falciparum. Together with reticulocyte-binding protein homolog 5 (RH5) and RH5-interacting protein (RIPR), CyRPA forms the "RCR complex" - a trio of proteins essential for blood cell invasion. The parasite needs all three to get inside a blood cell, and all three are highly conserved and capable of inducing highly neutralising antibodies.
In previous work carried out at Diamond, the team focused on structural studies of RH5, which has led to the design of RH5-based vaccines now in clinical trials. That research featured in Diamond's 2020 Annual Review. This latest study of CyRPA should lead to the development of CyRPA-based vaccines to be tested in due course.
For more information about Diamond’s I03 MX beamline, contact MX Group Leader, Dave Hall: [email protected]. To find out more about the I04 MX beamline or discuss potential applications, contact Principal Beamline Scientist Ralf Flaig: [email protected]
Ragotte RJ et al. Heterotypic interactions drive antibody synergy against a malaria vaccine candidate. Nature communications 13.1 (2022): 1-12. DOI:10.1038/s41467-022-28601-4.
Alanine DGW et al. Human antibodies that slow erythrocyte invasion potentiate malaria-neutralizing antibodies. Cell 178.1 (2019): 216-228. DOI: 10.1016/j.cell.2019.05.025.
Campeotto I et al. One-step design of a stable variant of the malaria invasion protein RH5 for use as a vaccine immunogen. Proceedings of the National Academy of Sciences 114.5 (2017): 998-1002. DOI:10.1073/pnas.1616903114.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.