The proteins that bind
May 8, 2018
May 8, 2018
Serine-rich repeat proteins (SRRPs), which help bacteria attach to surfaces, have been structurally characterised in pathogenic bacteria but not in beneficial bacteria such as those present in the gut. Dr Nathalie Juge’s team at the Quadram Institute Bioscience has previously identified SRRP as a main adhesin in Lactobacillus reuteri strains from pigs and mice. Now, together with colleagues at the University of East Anglia, they have described the structure and activity of the binding region of L. reuteri SRRPs in a paper published in PNAS. Using the Macromolecular Crystallography beamlines (I03 and I04) at Diamond Light Source, they discovered that the structure of these proteins is unique among characterised SRRPs and is surprisingly similar to pectin degrading enzymes. Molecular simulations and binding experiments revealed a pH-dependent binding to pectin and to proteins from the epithelium known as mucins. Altogether, these findings shed light on the activity of a key protein in these bacteria and may help guide the development of more targeted probiotic interventions.
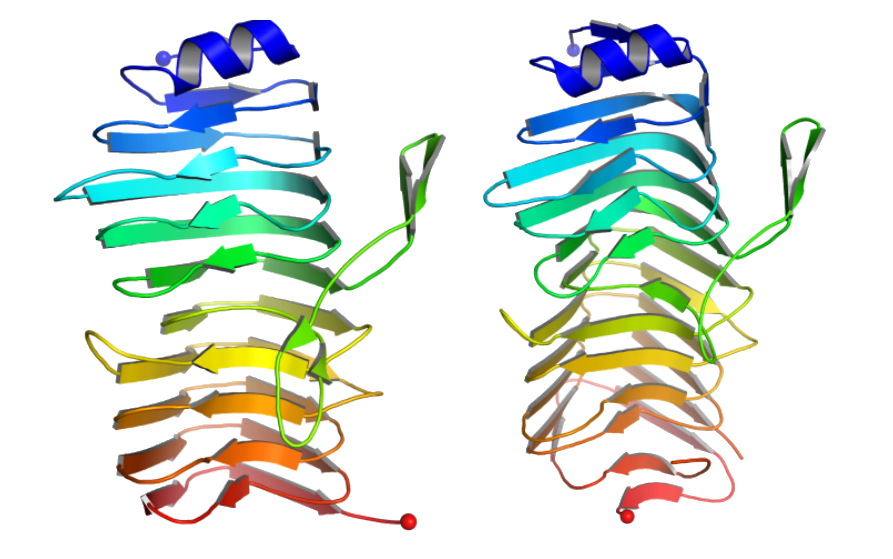 Figure 1: (Left) Cartoon representation of crystal structures of the binding region of SRRP53608. (Right) Cartoon representation of crystal structures of the binding region of SRRP100-23. The N-terminus is shown with blue balls and the C-terminus is shown with red balls.
Figure 1: (Left) Cartoon representation of crystal structures of the binding region of SRRP53608. (Right) Cartoon representation of crystal structures of the binding region of SRRP100-23. The N-terminus is shown with blue balls and the C-terminus is shown with red balls.Working with friends
To address this, Dr Nathalie Juge led an investigation of the structure and binding properties of SRRPs from the beneficial gut bacteria Lactobacillus reuteri. “In addition to its reported health properties, L. reuteri is used as a model in academia, to study the mechanism of host adaptation of gut symbionts,” says Dr Juge. In the current study, the team began by using bioinformatics to identify SRRPs in 58 strains of L. reuteri. Next, they analysed the sequence of the binding region of these SRRPs and grouped them accordingly. This analysis grouped most of the SRRPs into host-specific clusters, including a cluster with SRRP53608 from a pig strain and SRRP100-23 from a mouse strain.
Earlier collaborative research by Dr Juge had shown that SRRP100-23 is crucial for the mouse strain to form biofilms in the mouse forestomach, leading the researchers to focus on how these proteins bind. They concentrated on the binding region of SRRP53608 and SRRP100-23, which they crystallised following in situ limited proteolysis. Next, they used the I03 and I04 beamlines at Diamond to solve the binding regions’ structure through X-ray crystallography. Overall, both structures were quite similar, with the binding regions of both consisting primarily of a right-handed coil made of beta-strands.
A surprising twist
When the team compared these coils against other protein structures, they discovered an unexpected similarity. “We were really surprised when we saw the fold in the protein, because it was different from all of the ones that had been reported so far for SRRPs from pathogenic bacteria,” says Dr Juge. The search revealed that the SRRP binding region structure was strikingly similar to that of pectinases, a class of proteins that break down the plant molecule pectin. “That was not what we expected.”
When the team overlaid the structure of the binding region on the structure of pectinase bound to its ligand, they found that the binding site of the pectinase corresponded with a highly conserved region of charged, aromatic amino acids in the SRRP, leading them to hypothesise that this might be its binding site. “The literature reported that pectinases show a pH-dependent conformational change, so we decided to test it,” says Dr Juge. Molecular dynamics simulations confirmed the hypothesis. The simulations predicted pH-dependent conformational changes near the putative binding site, suggesting that SRRP binding activity might depend on the pH gradient in the gut.
Finally, the researchers screened potential ligands to determine what these SRRPs bind. They began the test with arrays of sialic acid compounds, since many SRRPs from pathogenic bacteria bind to sialic acids. “We couldn’t see any significant interaction, and we found ourselves having to screen more widely,” recalls Dr Juge. Eventually, they found that SRRP53608 and SRRP100-23 bound to some pectin components at low pH and to mucin glycoproteins and DNA from the epithelial lining at higher pH. In other words, these proteins bind to plant components in food or to the epithelium, depending on the pH, which may help them to persist in the gut.
By characterising key adhesive proteins of Lactobacillus reuteri, this study helps explain how these beneficial gut bacteria interact with their environment and how they form biofilms, which may guide us in developing more effective probiotic interventions in animal husbandry. “To date, biofilm formation is associated with detrimental effects of pathogenic bacteria and resistance to antibiotics, but using biofilm-forming Lactobacillus could offer a novel strategy to combat pathogens and antimicrobial resistance,” says Dr Juge. Although the strains used in this study are specific to mice and pigs, the findings may also eventually have implications for humans. “It might be possible to use what we’re learning to modify strains so they can persist longer and target the right site for the right application,” says Dr Juge.
To find out more about the I03 or I04 beamlines, or to discuss potential applications, please contact Principal Beamline Scientist Dr Katherine McAuley ([email protected]) or Dr Dave Hall ([email protected]), respectively.
Sequeira S et al. Structural basis for the role of serine-rich repeat proteins from Lactobacillus reuteri in gut microbe-host interactions. PNAS 115, E2706-E2715 (2018).
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.