A human protein is essential for spreading Flu
Nov 27, 2020
Nov 27, 2020
A team of researchers in the UK have uncovered how a human protein allows the influenza virus to replicate inside human cells. The protein called ANP32 is essential for genome replication of our ownvcells, but when they become infected with influenza C virus it functions quite differently. ANP32 protein helps provide the stability that the viral enzymes need to both produce functional proteins as well as replicate the influenza genome. This leads to more virus particles being produced resulting in more cells being infected and ultimately the transmission of the virus. This study highlights the intricacy of genome replication in influenza C and the complex interplay with the host cells. It also highlights very specific targets for the development of inhibitors which could become new treatments for flu.
Viruses that cannot replicate their genome cannot cause disease. To infect a cell, a virus is essentially destroyed, but the ability to copy genetic material inside a cell means that many new viruses will be produced that will go on to cause infection. Influenza viruses have RNA genomes and they produce an RNA polymerase which is an enzyme that can make copies of the viral genetic material. Research over the years has shown that this process is rather complex, requiring many different interactions of protein complexes with different RNA molecules, one of those proteins being the human ANP32 which is essential for viral replication. The research team knew that the ANP32 protein varies between humans and birds, which turns out is very important for influenza virus replication. RNA polymerases from avian influenza strains cannot replicate in the presence of ANP32 from humans and vice versa, so what was the cause?
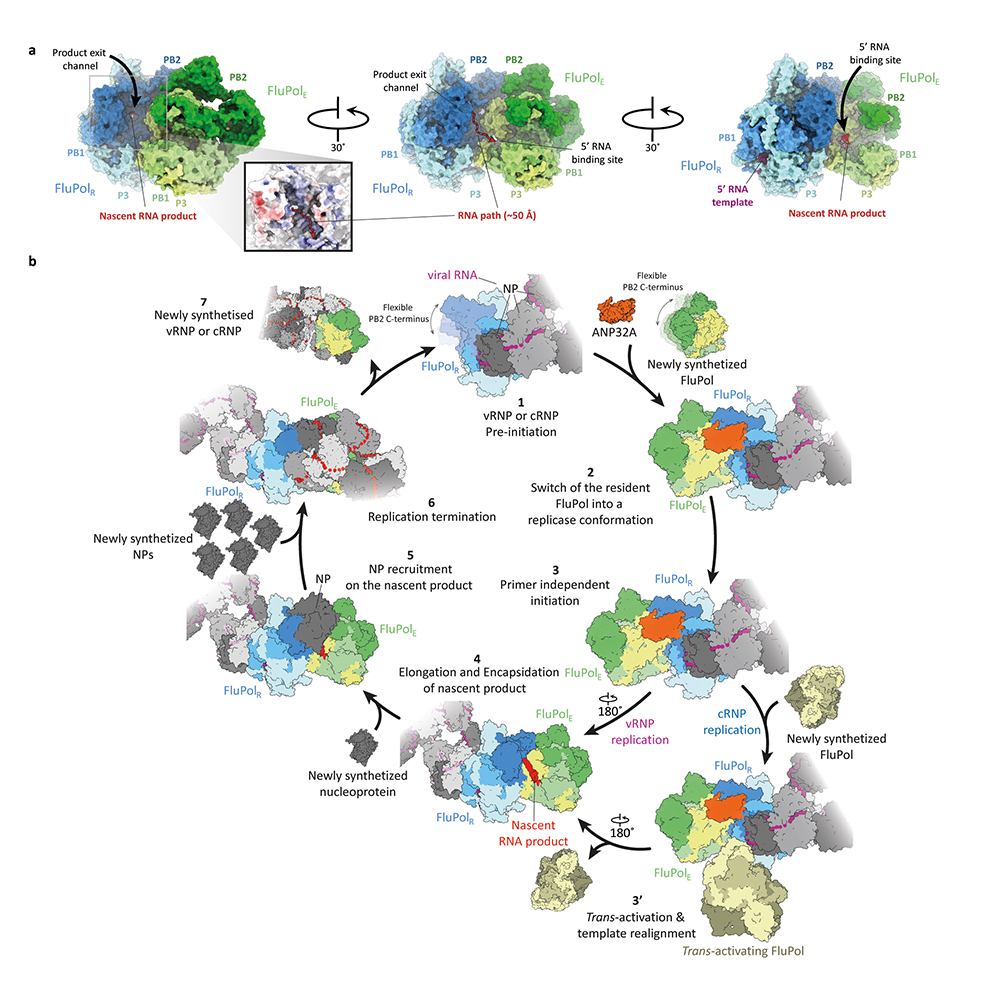
The research team used Cryo-Electron microscopy to understand the interactions of the viral RNA polymerase with ANP32. They found that the human protein helped to form a stable bridge between two molecules of the viral polymerase. Crucially, this stability was only seen between human ANP32 and influenza virus C polymerases from viruses that could infect humans. The question was how did this stability help the virus to replicate? By probing deeper into the Cryo-EM data, the research team found an answer. When the virus first infects a cell, there are fewer copies of the polymerase enzyme, and this means that instead of replicating the genome, the enzymes produce a molecule called mRNA which is used to create more proteins, including more polymerase enzymes. As the concentration increases, polymerase molecules come closer together when the ANP32 can also interact with them and form a stable bridge. When the two enzymes fastened together, they can then replicate the influenza virus genome. This intricate mechanism ensures that genome replication only occurs later in the infection process.
This new insight into a critical step in viral replication opens the door for novel inhibitors. The complex of two polymerase enzymes with ANP32 is not only essential for genome replication, but it also protects the viral RNA from being recognised and destroyed by the cell. If the complex could not form, then enzymes inside the cell may be able to destroy the viral genome before it is replicated, putting a halt to viral transmission. Given that animals present a large reservoir for influenza viruses, this new finding provides an avenue for producing bird flocks with a modified ANP32 protein that cannot form a stable bridge, making the animals resistant to the virus.
Carrique, L., Fan, H., Walker, A.P. et al. Host ANP32A mediates the assembly of the influenza virus replicase. Nature 587, 638–643 (2020). DOI: 10.1038/s41586-020-2927-z
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.