With help from a few friends
Feb 15, 2018
Feb 15, 2018
A complex made up of three proteins, Hsp90, Sgt1, and Rar1, is thought to stabilise an important immune protein known as nucleotide-binding domain and leucine-rich repeat containing protein. While the structure of the Sgt1-Hsp90-Rar1 protein is known, the stoichiometry of the complex has remained elusive. In a paper published in Frontiers in Molecular Biosciences, Dr Chrisostomos Prodromou of the University of Sussex and Dr Minghao Zhang of the University of Oxford worked with Professor Giuliano Siligardi at the Circular Dichroism beamline (B23) at Diamond Light Source to clarify the detailed make-up of the complex. Using synchrotron radiation circular dichroism, they revealed that it consists of an Hsp90 dimer, two Sgt1 molecules, and a single Rar1 molecule. The stoichiometry of the full complex potentially allows two NLR molecules to bind, a finding which may open avenues of research into how these proteins form dimers.
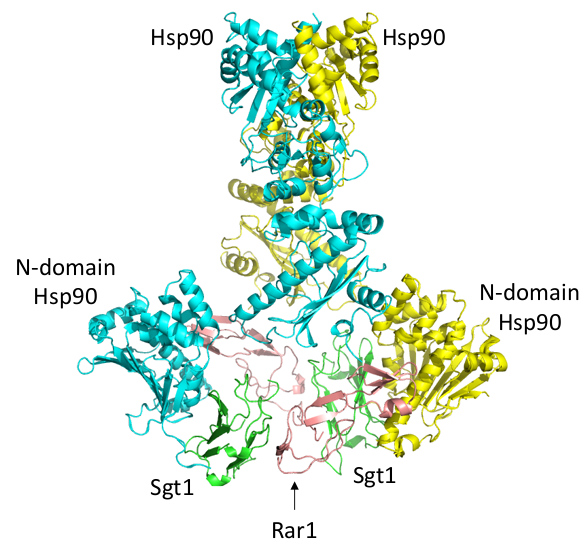
Figure 1: The structure of the Sgt1-Hsp90-Rar1 complex with an Hsp90 dimer, two Sgt1 molecules, and a single Rar1.
Subheading A complex conundrum
Heat shock protein 90 (Hsp90) is a molecular chaperone, a protein that helps other proteins attain and maintain their activity. Hsp90 is required for the correct functioning of nucleotide-binding domain and leucine-rich repeat containing (NLR) proteins, which act as intra- and extra-molecular sensors in immune response. Together with its co-chaperones Sgt1 and Rar1, Hsp90 forms a complex which interacts with NLRs and is believed to stabilise the protein and perhaps affect its dimerization.
Understanding how the Sgt1-Hsp90-Rar1 complex affects NLR calls for a clear mechanistic picture of their interactions. While the structures of several domains of the three proteins in complex were solved several years ago, “one of the outstanding issues that remained was understanding the stoichiometry of the complex,” says Dr Chrisostomos Prodromou. “Crystallographer’s often use a divide-and-conquer process where they look at the structure of some of the domains. We tend to just look at the interacting domains, which are easier to crystalise, but you lose information about what the overall stoichiometry of the complex with all the components would be.”
Two Hsp90 molecules were known to make up the core of the complex, but there was uncertainty about the number of Rar1 and Sgt1 molecules. Since Rar1 contains two domains which could interact with Hsp90, researchers were unsure whether one or two Rar1 molecules were involved. The answer to this would influence not only how many Sgt1 molecules were bound but also how many NLR receptors there would be, affecting the interactions between the complex and NLR. “We didn’t quite know how the whole thing would come together,” says Dr Prodromou.
A certain slant of light
Based on what researchers knew, there were three alternatives for the make-up of the Sg1-Hsp90-Rar1 complex. The Hsp90 dimer at the core might be connected to two Sgt1 and two Rar1 molecules, to one of each, or to two Sgt1 molecules and a single Rar1. To distinguish between these, Dr Prodromou turned to circular dichroism (CD), a spectroscopy technique based on differential absorption of left and right circularly polarised light. Since CD doesn’t require labelling or crystallisation of samples, it is an ideal approach for studying binding interactions and large multiprotein complexes.
Dr Prodromou and Dr Minghao Zhang teamed up with “probably the most experienced CD person we have in the country,” Prof Giuliano Siligardi, the principal beamline scientist on Diamond’s B23 beamline. In addition to benefitting from Prof Siligardi’s expertise, the study relied on the increased sensitivity gained by using synchrotron radiation. “In my opinion, we needed a synchrotron radiation experiment to get this done. It would be very hard to do this with desktop CD alone,” says Dr Prodromou.
The researchers began by measuring the interactions between the individual components and ADP, which is believed to help stabilise the final complex. Next, they measured the binary protein complexes. Finally, they carried out complementary titration experiments, titrating Sgt1 into a preformed complex of an Hsp90 dimer and a single Rar1, and then titrating Rar1 into a preformed complex of an Hsp90 dimer combined with two Sgt1 molecules. Analysis of the results revealed that the full complex consists of an Hsp90 dimer bound to two Sgt1 molecules and a single Rar1. “We could not have determined this with other, more commonly used techniques such as immunoprecipitation or gel filtration,” says Dr Prodromou.
Based on its stoichiometry, the Sgt1-Hsp90-Rar1 complex can accommodate two NLR proteins, though it remains unclear whether they would form a dimer or remain monomers. Dr Prodromou says “At this point, we would have liked to look at the NLRs in more detail but they’re incredibly difficult proteins to make. Currently, we can’t progress this work any further down those lines – we've reached a bit of an impasse now because we can't yet make the proteins, and we need to turn to expression systems that can do this for us.” However, he is likely to return to CD as a tool for revealing the structures of other complexes in the future. “Prof Giuliano is keen to push these types of experiments forward and push the technique to new levels.”
To find out more about the B23 beamline, or to discuss potential applications, please contact Principal Beamline Scientist Prof Giuliano Siligardi: [email protected]
Siligardi, G et al. The stoichiometric interaction of the Hsp90-Sgt1-Rar1 complex by CD and SRCD spectroscopy. Frontiers in Molecular Biosciences 4, 95 (2018). DOI: 10.3389/fmolb.2017.00095
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.