Infrared (IR) spectromicroscopy has been used for the past twenty years to study biological matter because of its sensitivity for detecting sub-cellular components in intact cells and tissues. As the method has improved, attention has turned to the application of this tool for the analysis of biochemical reactions within living cells.
Until now, the interpretation of IR absorption spectra was hampered by the complexity of the cell contents, as signals arising from small molecules were lost amongst those from dominant constituents such as proteins, sugars, DNA and fats. To overcome this shortfall, a novel method was developed called correlated cellular spectromicroscopy (CSM). The method involves the study of correlations between IR bands to identify small molecules based on their ‘spectral fingerprint’.
At the Multimode InfraRed Imaging and Microspectroscopy (MIRIAM) beamline at Diamond Light Source, the identification of metabolites in a human cancer cell line via CSM was followed up by IR microscopy studies with synchrotron radiation and Focal Plane Array (FPA) detector to determine their concentrations in time and space within single cells. The study of real-time metabolism has huge implications for drug discovery, since the impact of new drug candidates could be tested without damaging the cells. IR spectromicroscopy therefore has great potential for carrying out in vitro screening.
Much of our existing knowledge of biochemistry is based on the study of isolated biomolecules and simple reaction mixtures. In contrast, biochemistry in a living organism, down to the single cell or single organelle level, is described by complex and interacting reaction networks, with the simultaneous turnover of thousands of different molecules. One of the current challenges in bioanalytical chemistry is developing techniques to monitor the biochemistry in living systems in real time. Solving this challenge is a goal of both fundamental and of medical and pharmacological research. Success relies on addressing questions of sensitivity, resolution and molecular selectivity.
One approach to this end relies on the use of mid-infrared (MIR) absorption spectroscopy and microscopy. Most molecules strongly absorb mid-infrared light, including nearly all the ones of biological relevance, although only a limited subset is present in vivo at detectable concentrations1. The technique is non-destructive and can be used to monitor living cells over time without any need for labelling or staining2. The spectroscopic information can be related to structuralinformation, allowing both molecular identification and the study of structure-function relationships. One difficulty traditionally encountered in implementing MIR spectroscopy in a microscopy configuration is the degradation of signal quality when working at high spatial resolution. This difficulty has been reduced by the introduction of brighter light sources, such as synchrotrons and tuneable lasers, and of 2D multi-element imaging detectors1. In this research MIRIAM beamline at Diamond Light Source was used to perform measurements with diffraction limited spatial resolution with good signal quality via FPA detector.
An outstanding challenge in MIR spectroscopy and microscopy of living systems is the complexity of the resulting spectral information. MIR absorption spectra, even of relatively simple molecules, display complex patterns of overlapping bands. When analysing pure compounds or simple mixtures, these patterns allow the reliable identification of the molecule or molecules comprising the sample, an approach which is called spectral fingerprinting. However, the viability of fingerprinting decreases rapidly with the compositional complexity of the sample. Molecular identification in as complex a sample as a cell is a major challenge. In past years, molecular identification in MIR spectra of cells and tissue has relied in good part on our existing knowledge of the biochemistry of the cell type or tissue under study. This concern was one of the central issues at the 2016 Faraday Discussion on Advanced Vibrational Spectroscopy for Biomedical Applications, in Cambridge- UK, where the reliability of direct molecular identification in cells and tissue from MIR spectroscopy measurements was openly questioned3. Two views were discussed. According to one position, molecular assignments in cells are too unreliable and should not be attempted. Another position held that, despite existing limitations, it must be possible to develop methods that allow objective and reliable identification of molecules in living systems using MIR spectra.
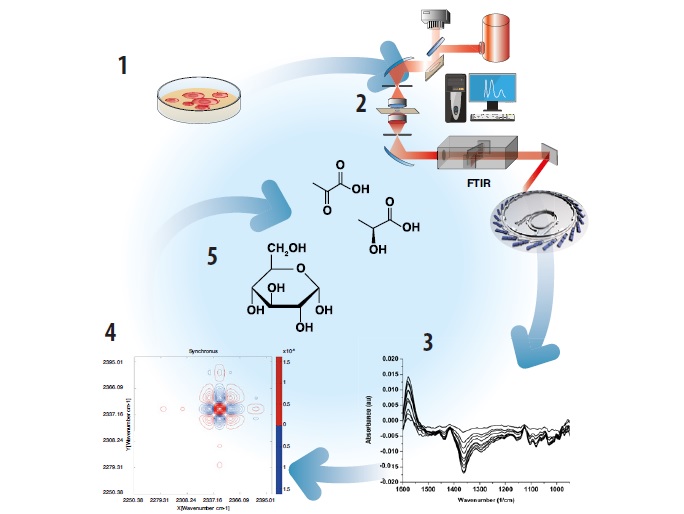
Figure 1: Experimental and data processing steps for identifying molecular features in live cells by IR microscopy. 1 – The cells are cultured on IR optical windows. 2 – The cells are transferred to an IR microscope. 3 - IR absorption measurements are performed and spectral information is plotted as a function of time. 4- Spectra are analysed using correlation analysis. 5 -The analysis is used to identify individual molecules and to assign sequences of reaction events. The result is a conversion of the complex spectroscopic properties of the cellular system into a description of reactant and product evolution.
This project follows the second view and aims to improve molecular identification by using a protocol called Correlated Cellular SpectroMicroscopy (CSM)4. The protocol relies on the analysis of the correlation properties of spectral bands to isolate the ones that arise from the same molecule, allowing their assignment by fingerprinting (Fig. 1). CSM constitutes a major improvement in the reliability of MIR spectroscopy to selectively monitor molecules in live cells. The analysis relies only on spectroscopic information. While a priori knowledge of the sample and additional information from other techniques can help in datainterpretation, it is neither necessary nor can it be performed on a sample which is unknown. Our approach has also been integrated with isotopic substitution, which is one of the most accurate strategies for band assignment in complex systems3. This creates an opportunity to make MIR absorption microscopy a real discovery method in biology, since it allows the identification of unknown or unexpected molecules in the sample.
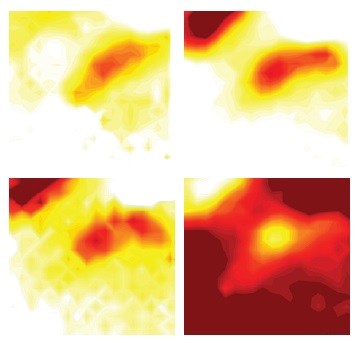
In our recent work, CSM has been applied to the study of metabolic turnover in the A549 lung cancer cell line and succeeded in identifying several metabolites involved in glycolysis. The analysis were followed up with IR imaging experiments on MIRIAM beamline, which allowed us to obtain images of metabolite distribution around individual living cells5. The results provide a description of local metabolic turnover of small molecules, including the formation of the end product of fermentation, lactate and the consumption of glucose (Fig. 2). This work is a proof of concept that IR microscopy allows imaging cellular metabolic production in a non-destructive way and at single cell level. In addition to the glycolytic metabolites detected in these experiments, several other small molecules could potentially provide a measurable contribution to cellular IR absorption, based on their reported cytoplasmic concentrations. Only a small fraction of known human metabolites is detectable5. Nonetheless this corresponds to at least 30 molecular species. This opens the way to the use of IR imaging to study the biochemistry of live cells, individually, in clusters or in tissue. The role of fluctuations of local concentrations in metabolic networks and the interaction between metabolic activities in adjoining cells can all be measured and hypothesis can be tested. It has also been proposed that the integration of CSM with other multivariate analysis methods, such as Partial Least Square Regression analysis, will potentially lead to the automated evaluation of metabolic kinetics in living cells3.
A clear target application is the study of cancer metabolism and tumor environment. The subject is receiving considerable attention because of the possibility to control or suppress tumor cells by selectively targeting their glycolytic metabolism. MIR microscopy and CSM can provide a wealth of information to this end, including identification of metabolites and quantification of rates of production, which is key in assessing the activity of drug candidates.
References:
Funding acknowledgement:
LQ and TZ acknowledge Diamond Light Source for beam time allocation at the MIRIAM beamline B22, (proposal SM12221) and support which allowed the results presented here.
Corresponding author:
Dr Luca Quaroni, Faculty of Chemistry, Jagiellonian University, [email protected]
Related publication:
Quaroni L, Zlateva T, Wehbe K, Cinque G. Infrared imaging of small molecules in living cells: from in vitro metabolic analysis to cytopathology. Faraday Discussion 187, 259-271, doi:10.1039/C5FD00156K (2016).
Publication keywords:
Single cell; Infrared spectroscopy; Infrared microscopy; Band assignment; Metabolism, Synchrotron infrared; FTIR; FPA
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.