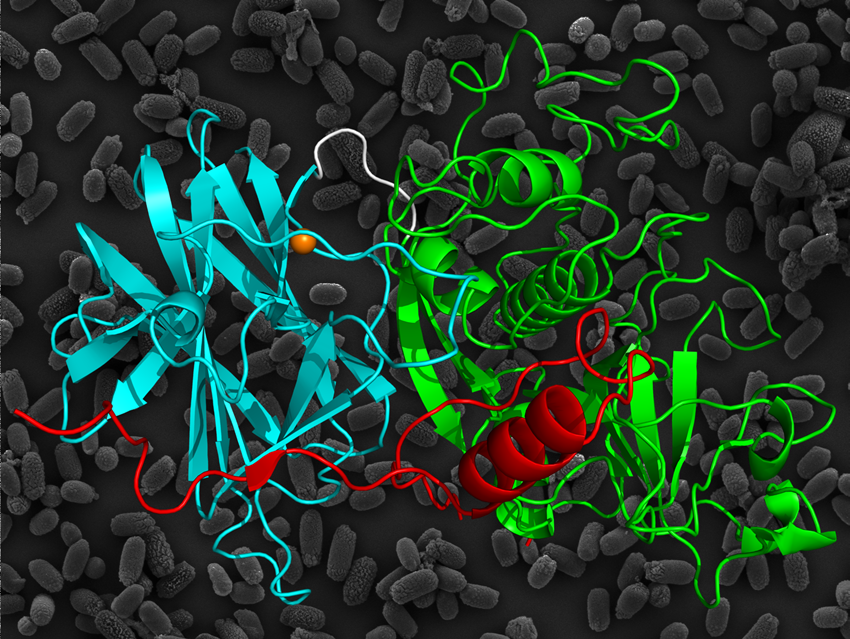
Ribbon diagram of the three-dimensional structure of the propeptide, cysteine protease and lectin-like domains of Cwp84.
(Cover image of Acta Cryst. D, vol. 70-7, 2014)
Bradshaw WJ et al. The structure of the cysteine protease and lectin-like domains of Cwp84, a surface layer-associated protein from Clostridium difficile. Acta Cryst. D70-7, 1983-1993 (2014). DOI: 10.1107/S1399004714009997
The bacterium Clostridium difficile (C. difficile) is a major health concern, particularly in the hospital environment, and counteracting it is problematic due to a poor understanding of the mechanism by which it colonises the gut. A team of structural biologists from the University of Bath and Public Health England (Porton Down) have made a breakthrough in determining the crystal structure of a protease key in the generation of the surface of the bacterium.
C. difficile infection causes diarrhoea, severe intestinal inflammation and potentially death. Disruption of the normally protective gut microbiota following a course of antibiotics, which may destroy much of the bacterial flora, can result in the colonisation and overgrowth of C. difficile bacteria. Understanding how the microbe colonises the intestine after exposure to antibiotics could lead to new ways to control infection. The growth, adherence and survival of C. difficile in the gut depends on a myriad of components particularly those on the bacterial surface. However, the mechanism of its outermost protein surface-layer protein (S-layer) biogenesis remains largely unknown. The mechanism involves the post-translational cleavage of a single peptide into low- and high-molecular weight subunits by surface protein Cwp84, a cysteine protease enzyme.
Researchers carried out high resolution crystallography at end stations I02 and I03 at Diamond to determine the structure of the C. difficile S-layer protein Cwp84. The structure was elucidated using a technique known as MAD, or multi-wavelength anomalous dispersion/diffraction, and the first crystal structure of this protein, obtained at a resolution of 1.4 Å, has been reported in Acta Cryst. Principal investigator for the project, Prof Ravi Acharya of the University of Bath, says why the team chose Diamond: “It is a world class facility, allowing cutting edge research to be performed with the some of the latest, most powerful, most accurate, and most versatile equipment available. The ability to tune the MX beamlines at Diamond was essential to this work.”
Twelve data sets were collected from four crystals containing selenomethionine and through quantification of the scattering effects of selenium, initial phase estimates were obtained. Refinement of the model with higher resolution data provided information concerning the molecular basis for the role of Cwp84 in the S-layer. Three regions were established, consisting of a cleavable propeptide, a cysteine protease domain and a newly identified linker domain, which binds calcium ions and resembles the structure of a group of proteins called lectins. Analysis of the interactions present between the domains provides information about protein folding and stabilisation, which may assist in deducing the mechanism for Cwp84. Blocking the activity of Cwp84 may prevent gut colonisation and reduce the spread of C. difficile, allowing repopulation of the intestine with “healthy” bacteria.
Future work will aim to ascertain further details about the structure and function of various proteins that make up the S-layer. Insight into surface layer growth and its relation to colonisation of the intestine could be exploited in developing drugs to treat infection. According to Prof Acharya, “Clostridium difficile is a global medical and economic problem. The development of more effective methods of preventing C. difficile infections will require a greater understanding of the bacterium.” He concludes, “These results improve the understanding of the function of the complex S-layer of C. difficile, providing insight into the role of Cwp84. The information could also potentially be used for future drug development, by targeting Cwp84.”
For more information about the capabilities of beamline I03, or to discuss potential applications, please contact Principal Beamline Scientist Katherine McAuley: Katherine.McAuley@diamond.ac.uk
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.