A collaboration of scientists from Diamond Light Source and the
University of Reading has revealed the binding mechanism of a so-called ‘light-switch’ effect complex, a type of chemical compound that fluoresces on binding to DNA. There are two possible applications for these compounds – in sensitive diagnostic tests and as sensitizers for photodynamic therapy. The team used the Diamond synchrotron to determine exactly how the photoreactive metal complex binds to DNA, revealing that the binding is sequence specific. Their findings, which were published online in
Nature Chemistry 24 June 2012, may pave the way for improved cancer diagnostics.
Since the 1980s, researchers have been interested in the interaction between ruthenium complexes and DNA, with a view to activation of the complexes by means of a light source. This is the basis of photodynamic therapy, a highly selective technique for destroying cancerous cells whilst ensuring most healthy cells go unharmed. Although there are currently a small number of clinical drugs that work in this way, the applications are relatively few. This new information opens up the potential to develop this technique further to enable wider application and more specific targeting.
The compounds involved in photodynamic therapy are designed to kill cells, which is currently done by forming ultra-reactive oxygen within the cell which then destroys them. However, compounds which attack and destroy DNA are currently in development and as such the key to making them work successfully is understanding their mode of action.
Until now, previous research was only able to determine which chemical compounds would bind with DNA when irradiated; the details of the interaction were highly debated. The structural data collected on two of the
Macromolecular Crystallography (MX) beamlines at Diamond, I02 and I04, has revealed what is happening at the molecular level.
James Hall, a PhD student with the University of Reading and Diamond, explains, “Solving the crystal structures of compounds bound to DNA showed us for the first time that they bind in two different ways and that the binding is specific to the DNA sequence. We now know that the complex will only bind with DNA with TA base pairs but not AT. This means we can look towards targeting an individual DNA sequence, potentially allowing future development of highly specific binding agents.”
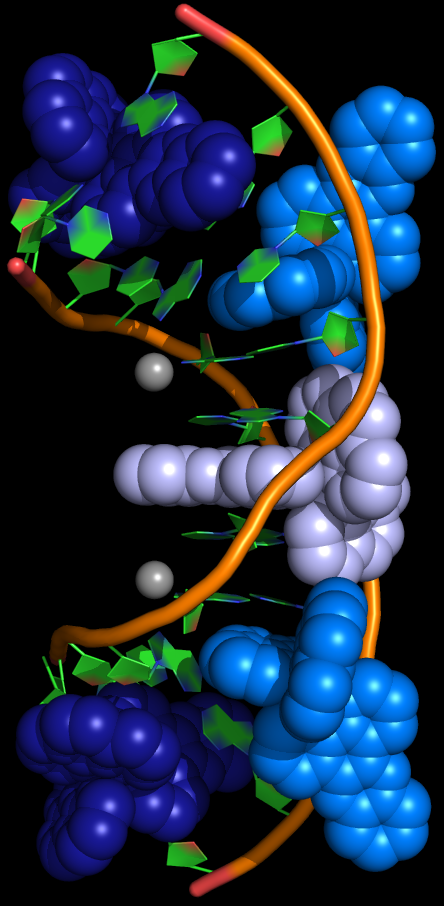
The image to the right shows the ruthenium complex bound with DNA with TA base pairs. Two of the ruthenium complexes are shown in dark blue, a third complex is shown in grey, semi-intercalated complexes are shown in light blue, the DNA backbone is shown in orange; carbon, green; oxygen, red; nitrogen, blue.
James is one of over 40 joint PhD students in post at Diamond, with each being affiliated to and part funded by a UK university. Although the projects are extremely varied, they are all pushing the boundaries of what is possible on the beamlines at the Diamond synchrotron.
James was responsible for growing some of the crystals for this experiment, and the subsequent data analysis. One part of his PhD project with Reading and Diamond is to develop reliable crystals that can be used to test and commission the experimental equipment on the MX beamlines at Diamond.
- ENDS -
Crystal structures of L-[Ru(phen)2dppz]2+ with oligonucleotides containing TA/TA and AT/AT steps show two intercalation modes
Niyazi, H, Hall, J.P., O'Sullivan, K., Sorensen, T, Winter, G, Kelly, J.M. and Cardin, C.J
Published in Nature Chemistry online, 24th June 2012
DOI: 10.1038/NCHEM.1397
 Since the 1980s, researchers have been interested in the interaction between ruthenium complexes and DNA, with a view to activation of the complexes by means of a light source. This is the basis of photodynamic therapy, a highly selective technique for destroying cancerous cells whilst ensuring most healthy cells go unharmed. Although there are currently a small number of clinical drugs that work in this way, the applications are relatively few. This new information opens up the potential to develop this technique further to enable wider application and more specific targeting.
Since the 1980s, researchers have been interested in the interaction between ruthenium complexes and DNA, with a view to activation of the complexes by means of a light source. This is the basis of photodynamic therapy, a highly selective technique for destroying cancerous cells whilst ensuring most healthy cells go unharmed. Although there are currently a small number of clinical drugs that work in this way, the applications are relatively few. This new information opens up the potential to develop this technique further to enable wider application and more specific targeting.