Data collected on I04 at Diamond Light Source, for haem-containing (native) EfeB, confirms it is a structural member of the DyP-peroxidase family and despite the presence of a catalytic aspartate, EfeB acts via a typical haem-peroxidase catalytic cycle. We have shown, using UV-VIS spectroscopy, that EfeB is able to accept electrons from an artificial peroxide substrate (guaiacol) suggesting that a complex between EfeB and reducing substrate is not required and that EfeB can accept electrons at a distance (manuscript in preparation). This supports the notion that EfeB is able to accept electrons from EfeO (generated by the oxidation of Fe2+).
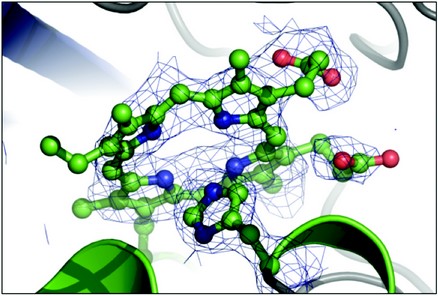
Until recently, it was believed that haem is internalised before degradation of the tetrapyrrol ring in the cytoplasm and extraction of iron, through the action of haem oxygenases and haem degrading monooxygenases. Our structural studies of EfeB, performed at the ESRF (ID14-EH2, ID23-EH1) and Diamond Light Source (I04), have demonstrated that soaking haem into preformed apo-EfeB crystals results in the presence of protoporphyrin IX (Fig. 1a), rather than haem in the EfeB active site, as noted for the haem purified form (Fig. 1b). Our observations to date support a deferrochelatase activity for EfeB and suggest an additional role for EfeB, retrieval of iron from exogenous haem (manuscript in preparation). Further work remains to determine the physiological significance of this novel enzyme activity, which may be of interest to industrial processes (such as food processing).
Additional interest in EfeB is driven by two factors; i) homologs of EfeB are widespread amongst Gram-negative and Gram-positive bacteria but absent in higher eukaryotic organisms and ii) iron uptake systems are important determinants of bacterial virulence, since the host specifically withdraws iron availability as part of its innate defence against infectious disease (such as tuberculosis). These factors support EfeB as a potential new target in the search for species-selective antimicrobial drugs involving iron transport-mediated uptake.
Our bioinformatic analysis of EfeO showed that the N-terminal region consists of an approximately 100 residue cupredoxin-like domain (Cup) with preference for binding copper and possibly iron, whereas the C-terminal region forms a ‘peptidase M75’ (M75) domain of approximately 250 residues, containing a highly conserved ‘HXXE’ motif with preference for zinc. 4 Using SRCD at B23 at Diamond Light Source, we were able to experimentally verify the secondary structural composition of EfeO from E.coli and an M75 domain protein, EfeM (an EfeO M75 domain homolog) from P. syringae. 10 SRCD showed EfeO secondary structure as, α = 51%, b = 12%, other = 38%, while the Cup domain of EfeO showed high β-sheet and low α-helix content (α = 8%, b = 34%, other = 38%), consistent with its anticipated Greek key topology. The M75 domain protein, EfeM, exhibited high α-helix content (α = 52%, b = 13%, other = 35%), consistent with the recently solved structure of an imelysin peptidase (insulin cleaving extracellular peptidase, PDB id 3N8U, JCSG unpublished) from P. aeruginosa: both belong to the metallopeptidase M75 family.
Like the Cup domain, the M75 domain was found to be a consistent feature of EfeUOB(M) transporters. Interestingly, there is very little structural data available for proteins belonging to the M75 metallopeptidase family, thus, we embarked on structure determination of EfeM and, very recently, collected 1.6Å resolution data for EfeM on beamline I04 at Diamond Light Source. Structure refinement and analysis are underway, but generally, the structure consists of two α-helical bundle domains hinged at one end and open at the other, resembling that of a periplasmic binding protein. The ‘HXXE’ motif is located at the open end of the cleft, poised for metal binding. Whether the two domains are able to move with respect to each other, allowing an opening/closing of the cleft, is unknown as yet but clearly a possibility.
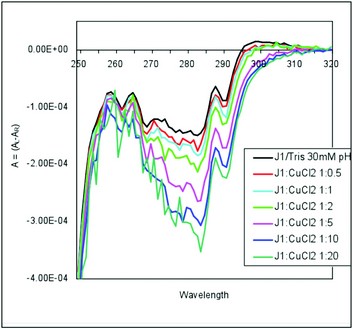
Figure 2a: Copper binding to the Cup domain protein from E.coli. Metal titration experiments, in the near-UV CD region using the Chirascan instrument provided in the B23 peripheral lab at Diamond, show localised changes upon copper binding in the 280 nm region. This is consistent with a copper binding site located near a hydrophobic (aromatic) residue. Our structural model shows a conserved Tyr residue located at the base of the predicted copper binding site pocket, as found in other cupredoxins. refined in the presence of haem.
Metal titration experiments, using CD spectroscopy on beamline B23, demonstrated a preference for copper binding to the Cup domain and zinc binding to the EfeM protein (Fig. 2a and 2b, respectively). Further results from recent SRCD metal titrations, together with metal binding studies using mass spectrometry, ITC and metal saturation experiments have been used to elucidate metal binding preferences and affinities for copper, iron and zinc for the individual domains (Cup and M75) of EfeO, and of the full length EfeO protein (manuscript in preparation). As predicted, the combined analyses demonstrate a preference for up to two copper (or one copper and one iron) by the Cup domain and a single zinc by the M75 domain. Comparison of the ferric-iron binding capacity of full length EfeO with EfeM (lacking the Cup domain) showed far higher iron levels for the former protein than the latter, suggesting that the Cup domain contributes to ferric-binding capacity.
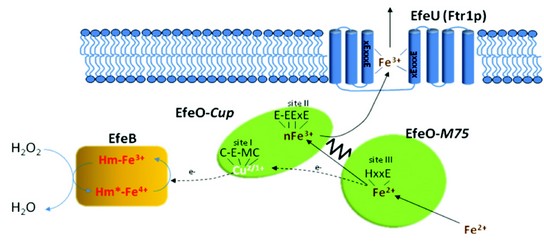
Figure 3: Schematic model of EfeUOB iron transport in E.coli. The oxidant employed is speculatively shown as hydrogen peroxide. It is anticipated that conversion of the excited form of EfeB to the resting state involves transfer of two electrons, sequentially, from EfeO.
Our work to date on this novel transporter has led to a proposed mechanism of iron transport in bacteria (Fig. 3)4. We propose that EfeO acts as a periplasmic Fe2+-binding protein and ferroxidase, and that EfeB accepts electrons from the ferroxidase reaction and disposes of these by combining them with an oxidant (possibly a peroxide). Once oxidised, the iron is then passed from EfeO to EfeU for translocation across the membrane. Although many questions still remain, the work to date supports the proposed model and accounts for the three potential metal binding sites in EfeO and the need of a haem peroxidase-like protein, EfeB.
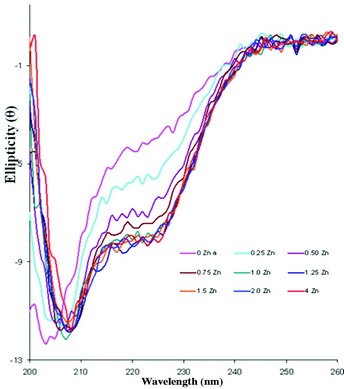
Figure 2b: Zinc binding to the M75 domain protein from P. syringae. Metal titration experiments, in the far-UV CD region using module B on B23 at Diamond, show secondary structural changes upon zinc binding with no significant change beyond 1:1 stoichiometry consistent with a single metal binding site, as predicted.
Rajasekaran, M.B., Mitchell, S.A., Gibson,T.M., Hussain,R., Siligardi, G., Andrews, S.C., Watson, K.A. Isolation and characterisation of EfeM, a periplasmic component of the putative EfeUOBM iron transporter of Pseudomonas syringae pv. Syringae Biochem. Biophys. Res. Commun. 398, 366-71. (2010)
References
- Cao, J., Woodhall, M.R., Alvarez, J. et al. Mol. Microbiol. 65, 857-75 (2007).
- Grosse, C., Scherer, J., Koch, D. et al. Mol. Microbiol. 62, 120-31 (2006).
- Ollinger, J., Song, K.B., Antelmann, H. et al. J. Bacteriol. 188, 3664-73 (2006).
- Rajasekaran, M.B., Nilapwar, S., Andrews, S.C. et al. Biometals. 23, 1-17 (2010).
- Askwith, C. and Kaplan, J. J. Biol. Chem. 272, 401-5 (1997).
- De Freitas, J., Wintz, H., Kim, J.H. et al. Biometals. 16, 185-97 (2003).
- Paronetto, M.P., Miele, R., Maugliani, A. et al. Arch. Biochem. Biophys. 392, 162-7 (2001).
- Cartron, M.L., Mitchell, S.A., Woodall, M.R. et al. Acta. Crystallogr. F63, 37-41 (2007).
- Sturm, A., Schierhorn, A., Lindenstrauss, U. et al. J. Biol. Chem. 281, 13972-8 (2006).
- Rajasekaran, M.B., Mitchell, S.A., Gibson, T.M. et al. Biochem. Biophys. Res. Commun. 398, 366-71 (2010).
Acknowledgements
A Felix Trust Scholarship to Mohan Babu Rajasekaran and the Research Endowment Trust Fund, University of Reading, to Vicki Bamford, funded this research. This work is in collaboration with Professor Simon Andrews, University of Reading.