Lentiviruses are associated with chronic disease states in a variety of mammals. However, until recently it was thought that these pathogens had no capacity for germ-line integration, and were only spread horizontally in an exogenous fashion. The discovery of the prehistoric endogenous lentiviruses in rabbits (RELIK)1 and lemurs (PSIV)2 refuted these ideas revealing lentiviruses to be present in a range of mammals, capable of germ-line integration and far more ancient than previously thought. In order to investigate the properties of ancient lentiviruses the Gag sequences of RELIK and PSIV were reconstructed from the remnants of these molecular fossils. These ancient Gags were combined with sequences from the present day lentivirus equine infectious anaemia virus (EIAV) to produce chimeric lentiviruses capable of budding, assembly and infection of non-dividing mammalian cells. Using the tuneable beamline I04 at Diamond Light Source, the crystal structures of capsid domains from PSIV and RELIK were then determined. These structures, from diverse ancient lentiviruses, were shown to be highly similar and contained many of the features found in modern day lentiviruses, including a functional cyclophilin-binding loop. Further structural, biochemical and biophysical studies of the interaction of these capsids with the host cell protein cyclophilin-A revealed the molecular details of an ancient capsid-cyclophilin interaction preserved throughout lentiviral evolution.
The discovery of the prehistoric endogenous lentiviruses RELIK1 and PSIV2 posed the notion of whether ancient sequences might encode functional proteins that are able to support the production of modern-day infectious lentiviral particles. In order to answer this question we synthesised the gag genes of the ancient lentiviruses RELIK and PSIV and constructed chimeric viruses that contained the capsid (CA) protein of RELIK or PSIV together with reverse transcriptase, protease and other proteins from EIAV. Our subsequent virological studies revealed these chimeric viruses were infectious and that ancient lentiviral capsids were able to support both budding and post-entry stages of the viral life cycle. Moreover, these viruses were capable of infecting non-dividing cells, a characteristic exclusive to lentiviruses.
In order to further investigate the capacity of RELIK and PSIV CA to support infection we determined the structures of the CA amino-terminal domains (CA-NtD) of these molecules at 1.5 and 2.0 Å resolution using beamline I04 at Diamond. The structures (Fig. 1) are representative of two diverse ancient lentiviral capsids that existed more than twelve million years ago. Remarkably, they exhibit a strong degree of similarity to each other, with an overall rmsd of only 1.3Å in the Ca positions for residues in secondary structure elements. Furthermore, the structures contain the arrangement of secondary structure elements observed in the N-terminal domains of CAs of modern retroviral genera, comprising a short ß1-ß2-hairpin and a five a-helix core made up from a1, a2, a3, a4, and a6. The presence of a highly ordered N-terminal b-hairpin in both structures is also indicative of a strong evolutionary functional conservation as formation of this feature is associated with viral maturation and is a critical requirement for the infectivity of present day lentiviruses.
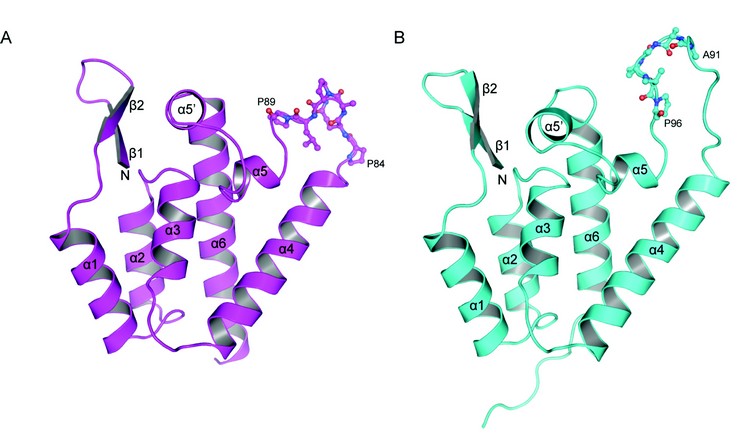
Figure 1: Crystal structures of CA amino terminal domains from (A) PSIV and (B) RELIK, cartoon representations of each molecule are shown a-helices and ß-strands are numbered sequentially from the N-terminus. Residues 84-89 (PQQPLP) in PSIV and 91-96 (AAGPVP) in RELIK that constitutes the CypA binding site are highlighted in stick representation.
The major differences between the RELIK and PSIV structures and that of modern-day lentiviruses are in the region that joins a4 to a6, the equivalent of the HIV-1 cyclophilin-A (CypA) binding loop3. The capacity for lentiviruses to bind to cyclophilin domains is important for infectivity and likely key in the mechanism they employ to integrate into non-dividing cells. In RELIK, this region contains a large fifteen-residue extended loop containing a cis Gly93-Pro94 dipeptide linkage and is similar in conformation but not in composition to that in HIV-1. However, in PSIV the loop is much shorter, only nine residues in length, has an entirely different conformation and contains Glu-Pro and Leu-Pro rather than a single Gly-Pro dipeptide. Nevertheless, the presence of a cis Gly93-Pro94 dipeptide linkage and HIV-1-like loop conformation in the a4-a6 region of the RELIK CA-NtD prompted us to examine if a similar interaction took place between the host-cell peptidyl proline isomerase CypA and the CA-NtD from ancient lentiviruses as it does with the CA-NtD of HIV-1. Analysis of the incorporation of CypA into viral particles in vivo revealed that HIV-1, together with chimeric EIAV-RELIK and EIAV-PSIV viruses contained significant quantities of CypA whilst control EIAV and SIVmac did not. Further characterisation of the interaction in vitro using isothermal titration calorimetry (ITC) and Biacore surface plasmon resonance (SPR) revealed the affinity of ancient CA-CypA interactions. Both RELIK and PSIV CA-NtDs bound to CypA with a 1:1 stoichiometry and with equilibrium dissociation constants in the range of 10-100µM, comparable to that observed with HIV-1. These data also revealed that CypA appears to be able to accommodate a4-a6 loops with linkages other than Gly-Pro, as in PSIV only Glu-Pro and Leu-Pro dipeptides are present.
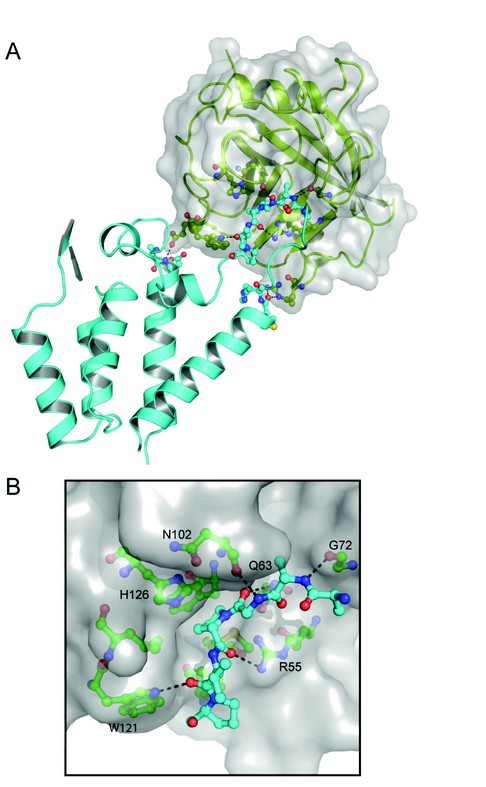
Figure 2: Structure of the RELIK-CypA complex. (A) Crystal structure of the RELIK CA-NtD-CypA, RELIK is shown in cyan and CypA in green cartoon representation. The molecular surface of CypA is also displayed. (B) Details of the RELIK-CypA molecular interface, residues 91-96 of RELIK CA-NtD that are located in the CypA binding groove are shown as sticks (blue). CypA residues that form the binding cleft are labelled and shown in green, hydrogen-bonding interactions are indicated with dashed lines.
These structural, biochemical, and virological studies provide compelling evidence for an ancient association between the lentiviral capsid and CypA. However, in order to understand the molecular details of the CypA interaction with an ancient capsid, we also determined the X-ray structure of a RELIK CA-NtD-CypA complex (Fig. 2A). The overall conformation of RELIK is essentially the same as in the free structure, except for a 10-12Å displacement of the a4-a6 loop. CypA adopts the same conformation that has been observed in other X-ray structures4 comprising an eight-stranded ß-barrel with a concave surface containing the hydrophobic peptide-binding groove and the proline-isomerase active site of the molecule. In the structure, residues Ala91-Pro96 of the RELIK are located in the binding groove of CypA and make extensive hydrophobic interactions together with main chain-mediated polar contacts at the interface. Within the groove, Pro94 adopts a cis configuration and is positioned adjacent to histidine-126 of CypA (cypHis126) in the hydrophobic pocket that constitutes the CypA active site. The key catalytic arginine, cypArg55, is located on the opposing side of the pocket from cypHis126 sandwiching Pro94 and hydrogen bonding to its main-chain carbonyl. This active site configuration is further stabilised through hydrogen bonding of the carbonyl group of the Gly93-Pro94 cis-peptide with the amide side chain of cypGln63 and by hydrogen bonds between the amide of Gly93 and the main-chain carbonyl of cypAsn102, and by the carbonyl of Val95 and the Ne1 of the cypTrp121 ring (Fig. 2B).
Comparison of the RELIK-CypA structure with that of HIV-1-CypA complexes5 shows that despite sequence and isomeric configuration differences, the backbone trajectory of the cyclophilin-binding loop together with side-chain orientation and hydrogen bonding contributions from cypTrp121, cypHis126, cypGly72 and cypAsn102 are structurally conserved. However, although the detailed organisation of the active site is conserved the complexes differ significantly in respect to the orientation of CypA relative to the RELIK and HIV-1 CA-NtDs. In the RELIK complex, CypA is rotated ~90° about the cyclophilin binding loop compared to the orientation observed in the HIV-1 structure. Furthermore, the bound Gly-Pro dipeptide in RELIK is permuted five residues C-terminal to the Gly-Pro dipeptide in HIV-1. This rotation combined with the shift in site position brings CypA closer to the core of the RELIK NtD than in the CypA-HIV complex. Consequently, the RELIK-CypA interface comprises a larger buried surface and includes specific interactions between cypArg148, cypGlu120 and cypTrp121 with secondary sites located at the termini of the RELIK a4 and a6.
These structures of molecular fossils demonstrate the extent of the similarities between ancient lentiviruses that colonised mammalian genomes 4-12 million years ago and their modern-day exogenous descendents. Moreover, they reveal that ancient capsids underwent the same maturation process required for infectivity and had the capacity to bind CypA through a diverse a4-a6 loop still present in many modern-day lentiviruses.
Goldstone, D.C., Yap, M.W., Robertson, L.E., Haire, L.F., Taylor, W.R., Katzourakis, A., Stoye, J.P., Taylor, I. A.
Structural and functional analysis of prehistoric lentiviruses uncovers an ancient molecular interface. Cell host & microbe. 8(3), 248-259, (2010)
References
- Katzourakis, A. et al. Discovery and analysis of the first endogenous lentivirus. Proc. Natl. Acad. Sci. USA. 104 (15), 6261 (2007).
- Gifford, R. J. et al. A transitional endogenous lentivirus from the genome of a basal primate and implications for lentivirus evolution. Proc. Natl. Acad. Sci. USA. 105 (51), 20362 (2008).
- Gamble, T. R. et al., Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid. Cell. 87 (7), 1285 (1996).
- Ke, H., Mayrose, D., and Cao, W. Crystal structure of cyclophilin A complexed with substrate Ala-Pro suggests a solvent-assisted mechanism of cis-trans isomerization. Proc. Natl. Acad. Sci. USA. 90 (8), 3324 (1993).
- Howard, B. R. et al., Structural insights into the catalytic mechanism of cyclophilin A. Nat. Struct. Biol. 10 (6), 475 (2003).
Acknowledgements
Medical Research Council, UK
Research carried out at Diamond Light Source on I04.

