Lassa fever virus kills more than five thousand people in West Africa every year, with many survivors having hearing and neurological problems.1 As no vaccine or effective drugs are available, and the infection can transmit from person to person, the Lassa fever virus is classed as a hazard group four virus and is regarded as a potential biological weapon. The nucleoprotein of Lassa fever is the most abundant protein in the virus and plays essential roles in virus translation, transcription, encapsidation and host immune suppression. The nucleoprotein crystal structure of the Lassa fever virus was determined to a resolution of 1.8Å, showing that the nucleoprotein consists of an N-terminal domain and a C-terminal domain. The C-terminal domain is a 3’-5’ nuclease, essential for immune evasion, whereas the N-terminal domain is a cap-binding polypeptide, which is essential for viral transcription. In addition, the structure also shows a zinc-binding motif at the C-terminal domain, which is a part of the nuclease active site. The structure indicates a cleft between the N-terminal and the C-terminal domains in the viral RNA binding site and both C-terminal and the N-terminal domains are involved in the nucleoprotein oligomerization. These findings reveal a novel mechanism for virus evasion from the human immune system, where we propose that the C-terminal nuclease specifically removes the pathogen-associated pattern ligand RNAs of the viruses, so that human cellular receptors do not detect the PAMP RNA ligands, resulting in no type I interferon (IFN) production. This could be important for the development of a vaccine and effective anti-viral drugs.
Arenaviruses currently have more than 22 members, and new members are emerging. Arenaviruses are not only important clinical pathogens, but also important experimental models. At least six arenaviruses can cause severe viral hemorrhagic fevers, and LCMV is one of the most popular experimental model viruses.2
The whole nucleoprotein of the arenavirus Lassa fever virus was overexpressed, purified and crystallized. The crystal structure of the Lassa fever virus nucleoprotein was determined to 1.8 Å resolution, which shows that the nucleoprotein contains N-terminal and C-terminal domains (Fig.1), with a structure not similar to any reported viral nucleoproteins.
The C-terminal domain consists of eight a-helices and five ß-strands, forming a typical a/ß/a sandwich structure. The structure of the C-terminal domain unexpectedly resembles 3’-5’ exonucleases, such as human TREX1, Ecoli DNA polymerase III epsilon chain, and Pseudomonas aeruginosa RNase T. Particularly, the five catalytic residues (D389,E391,D466,H528 and D533) are located at identical positions to the 3’-5’ exonucleases, which strongly suggests that the nucleoprotein of the Lassa fever virus is a 3’-5’ exonuclease. In vitro assays using different RNA substrates have confirmed that the nucleoprotein is a 3’-5’ nuclease. We propose that the nucleoprotein mimics the human TREX1 in vivo to degrade the pathogen-associated pattern ligand RNAs, such as the dsRNA and triphosphate dsRNA as shown in the in vitro assays, to prevent PAMP RNA ligands activating interferon production pathways.
Indeed, the cell base assays confirmed that substitution of the catalytic residues with alanine results in the loss of the nucleoprotein’s immune suppression. The results provide a totally new model for viral evasion of human immune response by degrading viral PAMP RNA ligands. By fluorescent scanning, a zinc ion was identified in the nucleoprotein, coordinated by the side chains of residues E399, C506, H509 and C529. This zinc-binding motif is part of the active site of the C-terminal nuclease.
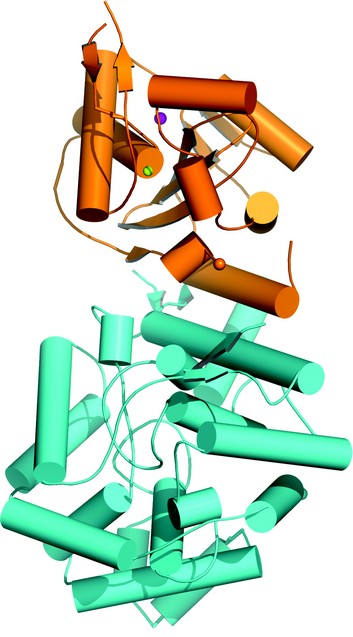
Figure 1: Protomer structure of Lassa fever virus nucleoprotein. The C-domain is in orange and N-terminal domain is in cyan. Mn2+ is shown in the pink sphere and Zn2+ is shown in the yellow sphere.
The N-terminal domain has a novel fold, mainly formed by helices and coils. As the arenaviruses cannot synthesise the cap for the viral mRNA, which is similar to other negative viruses such as influenza and bunyaviruses, the arenaviruses have to snatch the caps from human cellular mRNA. Our structural studies have, for the first time, provided evidence to show that the Lassa fever virus nucleoprotein is a cap-binding protein, which may be involved in cap snatching. Cap analogues, UTP and dTDP were located on the cap-binding site, which revealed a deep cavity at the N-terminal domain for the cap binding. Although the structure of the N-terminal domain is not similar to other cap-binding proteins, the cap-binding model of the N-terminal domain is similar. The cap (m7G) is located between two hydrophobic residues F176 and W164 (or L120), forming a sandwich structure. However, the deep cavity for the cap binding has the unique feature that the positive charged residues K253, K309 and R323 are in the middle of the cavity for binding the triphosphate of the m7Gppp and the entrance of the cavity consists of hydrophobic residues Y319, Y209, Y213 and L265 for binding another base. This indicates that the whole m7GpppN can be hidden in the cap-binding cavity, which can keep the m7GpppN from the exonucleases. To confirm the N-terminal domain’s function in cap binding, we performed a cap-dependent transcription assay based on the Lassa fever virus mini-genome replicon system, which proved that the proposed residues for the cap binding are truly crucial. This assay strongly supports our structural finding that the N-terminal domain forms the cap-binding polypeptides. The arenaviruses are reported to use a unique mechanism for cap snatching, as the non-template is only 1 to 4 nucleotides, which is much shorter than those used by influenza and bunyaviruses. The cap-binding feature of the N-terminal domain and the 3’,5’-nuclease of the C-terminal domain may help us to understand the unique cap-snatching mechanism.
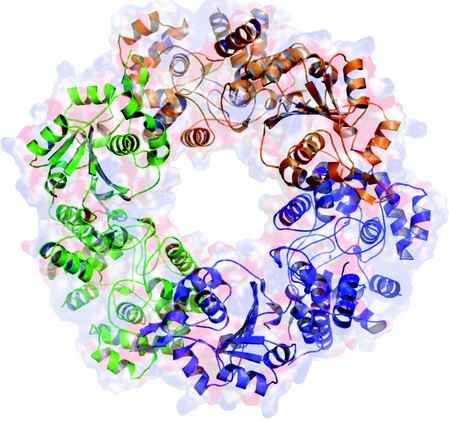
Figure 2: Trimeric structure of Lassa fever virus. The different subunits are highlighted in different colors.
In summary, the structure of the Lassa fever virus nucleoprotein is the first nucleoprotein structure of arenaviruses, which reveals a viral nucleoprotein that is not only key to viral RNA encapsidation, but also plays unexpected roles in immune suppression and viral transcription using novel mechanisms. These findings have great potential for the development of new effective vaccines and drugs.
Qi, X et al. Cap binding and immune evasion revealed by Lassa nucleoprotein structure. Nature 468(7325):779-83 (2010)
References
Acknowledgements
CD is a Wellcome trust career development fellow. CD thanks collaborators Dr. Yuying Liang and Dr. Hinh Ly from Emory University and Prof. Wenjian Wang from Sun Yat-Sen University. We thank the staff at I02 and I03 beamlines of Diamond Light Source for their assistance in data collection.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.