In order to determine the solution shape of human tropoelastin, small angle X-ray and neutron scattering measurements were made on solutions of tropoelastin and overlapping fragments. These included full-length tropoelastin (ID02 ESRF), exons 2-25 (I22 Diamond Light Source) and exons 2-18 (BioCAT). In addition, neutron scattering data was collected on full-length tropoelastin at ISIS. These data were processed using the ATSAS 2.4 suite of biosaxs software http://www.embl-hamburg.de/biosaxs/software.html. To determine the 3D solution shape of tropoelastin and overlapping fragments, the ab initio programs GASBOR and DAMMIN were used.2 Average shapes were created from multiple runs using the DAMAVER suite of programs, in order to identify common structural features. The ab initio shape of full-length tropoelastin from both SAXS and SANS data consistently revealed an elongated asymmetric molecule (Fig. 1). The dimensions were 16 nm end-to-end and 3 nm wide at the narrowest point which opens to 7.5 nm at the widest point. The molecule has clearly distinct regions starting at one terminus with a long narrow region that branches to give a larger more open appearance at the opposite terminus. The narrow elongated shape persists for ~11 nm before branching to give a “foot” at the end of the molecule. Therefore the molecule is not compact but is relatively extended.
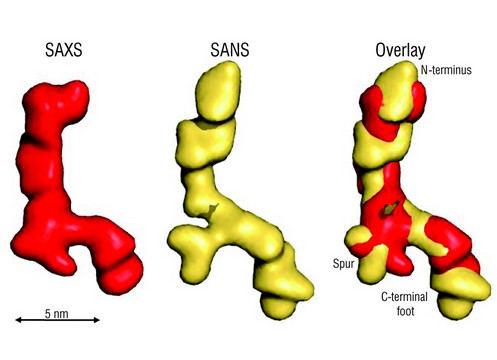
Figure 1: Ab initio shapes of full-length tropoelastin calculated from solution SAXS or SANS data. The filtered average shapes of 20 individual SAXS (red) and SANS (yellow) simulations are shown as a surface representation. An overlay of the models from the two scattering methods is also shown. The proposed locations of the N-terminus, the spur region containing exons 20-24 and the C-terminus are indicated. Scale bar is 5 nm.
In order to determine which ends of the tropoelastin shape correspond to the N- and C-termini, further SAXS analysis was performed on defined fragments that extended from a fixed N-terminus. X-ray scattering was conducted on the tropoelastin fragments exons 2 – 18 and exons 2 – 25. Data were processed using ab initio simulations as described above. The shape of exons 2-18 was a uniform elongated 14.5 nm long rod-shape that was 2 - 3 nm wide. It had a slight bend at approximately 60 % along its length to give an internal angle of 133o. The shape of exons 2-25 overall was remarkably similar to that of exons 2-18; however exons 2-25 had an additional protrusion at one end. The ab initio models of exons 2-18, 2-25 and full length tropoelastin were superimposed (Fig. 2). Exons 2-18 and 2-25 overlay closely with the linear region of the full length molecule, and since exons 2-18 are common between all three constructs, this suggests that the linear end corresponds to the N-terminus of tropoelastin. The protrusion seen in exons 2-25 corresponds exactly to the branching region in the full-length molecule and points to branching occurring at around exon 25 and the more open region therefore corresponding to the C-terminus of tropoelastin.
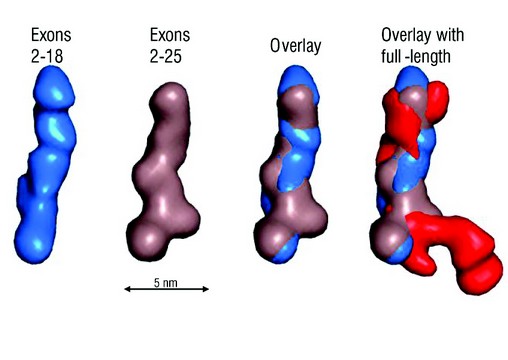
Figure 2: SAXS analysis of overlapping fragments of human tropoelastin. Ab initio models were calculated from SAXS data for tropoelastin constructs 2-18 (blue), 2-25 (brown) and full-length (red). An overlay of the two N-terminal fragments shows a conserved linear region. Scale bar is 5 nm.
We have found that tropoelastin is an asymmetric molecule with a gradual coil along the long, spring-like axis of the molecule. This coil region accounts for most of the elasticity of tropoelastin.3 The spur region protruding from the side of the molecule corresponds to a predicted hinge region containing exons 20-24.4 Beyond the spur there is a bridge to the C-terminal region. The tropoelastin molecule terminates in a more compact “foot-like” region. This part of the molecule includes the cell-interactive C-terminus of tropoelastin. 5 In summary, structural analysis reveals two dominant, functionally relevant parts of the molecule: the coil which contributes to elasticity and the foot that encompasses the C-terminal cell contact region. Our findings shed light on how biology uses this single protein to build durable elastic structures that allow for cell attachment to an appended foot. We present a new model for head-to-tail assembly which allows for the propagation of the molecule’s asymmetric coil through a stacked spring design (Fig. 3).
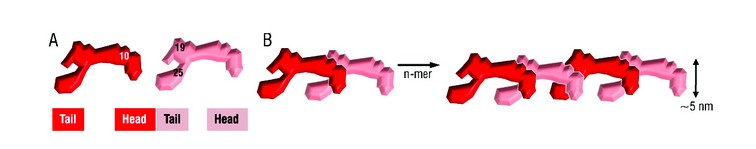
Figure 3: Head-to-tail model for elastin assembly. A) Juxtaposed domains 19 and 25 on one tropoelastin molecule and domain 10 on an adjacent monomer would allow the formation of a three-way desmosine cross-link found in vivo. B) Tandem assembly of tropoelastin monomers displaying n-mer propagation as an outcome of covalently bonded molecules.
Baldock, C., Oberhauser, A. F., Ma, L., Lammie, D., Siegler, V., Mithieux, S. M., Tu, Y., Chow, J. Y., Suleman, F., Malfois, M., Rogers, S., Guo, L., Irving, T. C., Wess, T. J., and Weiss, A. S. Shape of tropoelastin, the highly extensible protein that controls human tissue elasticity. Proc. Natl. Acad. Sci. USA. 108 (11), 4322 (2011)
References
- Wise, S. G. and Weiss, A. S. Tropoelastin. Int. J. Biochem. Cell. Biol. 41 (3), 494 (2009).
- Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76 (6), 2879 (1999).
- Holst, J. et al. Substrate elasticity provides mechanical signals for the expansion of hemopoietic stem and progenitor cells. Nat. Biotechnol. 28 (10), 1123 (2010).
- Dyksterhuis, L. B. et al. Domains 17-27 of tropoelastin contain key regions of contact for coacervation and contain an unusual turn-containing crosslinking domain. Matrix. Biol. 26 (2), 125 (2007).
- Bax, D. V. et al. Cell adhesion to tropoelastin is mediated via the C-terminal GRKRK motif and integrin alphaVbeta3. J. Biol. Chem. 284 (42), 28616 (2009).


