Dr Steven Gamblin, National Institute for Medical Research
Polycomb group proteins have an essential role in maintaining repressive epigenetic states and silencing unwanted gene expression during development. Disregulation of polycomb proteins have also been implicated in many types of cancer. Polycomb Repressive Complex 2 (PRC2) is the key enzyme complex that marks repressive chromatin domains by methylating lysine 27 of histone H3. PRC2 is composed of four ubunits, EZH2, which contains the catalytic SET domain, EED, Suz12 and RbAp48. Using beamline IO4 at Diamond, we have solved the structure of the WD40 domain from the EED subunit in complex with trimethylated lysine histone peptides known to be associated with repressive chromatin domains. We show that binding of repressive peptides to EED allosterically stimulates the PRC2 complex. From this we can suggest a model for the maintenance of repressive chromatin domains and for a transmission of histone modifications from mother to daughter cells during cell division.
The fate of a cell is specified by its gene expression profile, and an important way of determining which genes are expressed is through the control of access to DNA to enzymes such as transcription factors. In eukaryotic cells, DNA exists bound to histone molecules that package the DNA into repeating units called nucleosomes. A nucleosome consists of 146 base-pairs of DNA wrapped round an octet of histones. How adjacent nucleosomes along a DNA strand interact is key to the accessibility of the genetic information. If the nucleosomes are tightly packed then the genes contained in the region are effectively locked away and cannot be switched on, this is termed repressive chromatin. However, if the nucleosomes are loosely arranged and transcription factors can bind to the DNA then he chromatin is said to be active, and this allows gene expression. The switch between active and repressed chromatin is controlled by chemical marks, such as methylation, acetylation or phosphorylation, which are present on the tails of the histone molecules that comprise the nucleosome. These chemical marks allow gene expression to be regulated epigenetically, and the marks are often set early in development and maintained throughout the lifetime of a cell by a host of proteins that interact with histones and DNA.
The polycomb group (PcG) of proteins function by silencing inappropriate gene expression by maintaining a repressive epigenetic state [1][2]. They are essential to the correct development of eukaryotic multi cellular organisms. Indeed, PcG proteins were first identified in Drosophila through their roles in silencing homeotic (Hox) genes, though it is now known that there are many other targets of PcG proteins. The polycomb group can be divided into two main protein complexes Polycomb Repressive Complex 1 (PRC1) and PRC2. PRC2 functions by adding three methyl groups to Histone H3 Lysine 27 (H3K27) and methylation of H3K27 is associated with gene repression and silenced chromatin.
Figure 1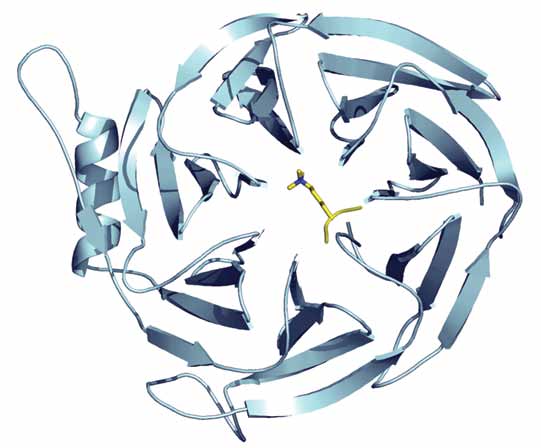 | The trimethylated state of H3K27 (H3K27me3) is the main form associated with gene silencing, though H3K27me and H3K27me2 marks are also widespread throughout the genome. Once PRC2 has methylated H3K27me3, then PRC1 recognises the trimethyl mark and binds, leading to compacting of the chromatin and gene repression. PRC2 is composed of four core protein subunits: EZH2, which contains the catalytic SET domain that is responsible for the methyltransferase activity [3]; Suz12, which is thought to bind DNA and RNA; RbAp48, which binds to histone H4; and EED, which has been shown to be essential for correct development. It is known that overexpression of components of PRC2 can enhance the pathogenicity of cancer in certain cell types [4]. |
| Figure 1. Structure of EED in complex with trimethylated histone H3 peptide. |
Using X-ray crystallography and other biochemical and biophysical methods, we have investigated the structure and the function of the EED subunit of PRC2. Initial crystallisation trials of the WD40 domain of EED resulted in small crystals that did not diffract. We found that crystals could be greatly improved by the addition of the non-detergent sulfobetaine additive NDSB-195. This allowed the structure to be solved and identification of electron density for the additive located in the cavity at one end of the WD40 barrel. We also saw that the NDSB-195 molecule had a close resemblance to a trimethylated lysine residue and hence reasoned that EED might bind trimethylated lysine peptides. A binding screen of such peptides with EED revealed that EED bound selectively to trimethylated histones associated with repressive chromatin (H3K27me3, H3K9me3, H4K20me3, H1K26me3) but not with active chromatin (H3K4me3, H3K36me3, H3K79me3).
 | Following the binding studies, we solved the structures of EED in complex with each of the trimethylated repressive peptides. Figure 1 shows the EED / H3K27me3 complex that was solved using data collected at beamline IO4 at Diamond Light Source. All of the peptides bind to a hydrophobic box that sits in the centre of the EED WD40 domain and is formed by residues Phe97, Tyr148, Trp364, and Tyr365 (Figure 2). EED family proteins are the only known examples of WD40 domain proteins that contain a hydrophobic box. The structure revealed why EED binds only to peptides associated with repressive chromatin: two hydrophobic cavities, which lie either side of the hydrophobic box, exclude the binding of charged residues found on peptides containing active marks. | |
| Figure 2. Hydrophobic box of EED (coloured blue) with bound H3K27me3 peptide (yellow). 2Fo-Fc electron density for the four box residues and peptide is shown. |
Using our structural data, we could design mutants to investigate further the role of EED in writing of H3K27me3 repressive chromatin marks. Using in vitro methylation assays, we showed that the binding of repressive trimethylated peptides to EED (especially H3K27me3), allosterically stimulated PRC2. If any of the hydrophobic box residues were mutated then the peptides failed to stimulate PRC2. Further studies looking at how EED mutants effected development of Drosophila revealed that the interaction of EED with repressive histone peptides through the hydrophobic box was essential for normal development. Indeed, mutant EED constructs resulted in a reduction of the global H3K27me2 and H3K27me3 levels. Taken together, our results suggest a model whereby the reading of existing H3K27me3 marks by EED allosterically stimulates PRC2 to promote the writing of new ones, therefore providing a way that repressive chromatin domains can be faithfully copied through cell division from mother to daughter cells (Figure 3).
References
[1] Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G. Cell 128, 735-45 (2007).
[2] Schwartz YB, Pirrotta V. Curr Opin Cell Biol. 20, 266-73 (2008).
[3] Müller, J., C.M. Hart, N.J. Francis, M.L. Vargas, A. Sengupta, B. Wild, E.L. Miller, M.B. O’Connor, R.E. Kingston, and J.A. Simon, Cell, 111, 197-208 (2002).
[4] Jones PA, Baylin SB. Cell, 128, 683-92 (2007). Review PubMed PMID: 17320506.
| Figure 3  | Principal Publications and Authors Margueron R, Justin N, Ohno K, Sharpe ML, Son J, Drury WJ 3rd, Voigt P, Martin SR, Taylor WR, De Marco V, Pirrotta V, Reinberg D, Gamblin SJ. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature, 461, 762-7, (2009). Funding Acknowledgement: Medical Research Council, UK. Research carried out at Diamond on I04 and at Daresbury SRS on beamline 10.1. |
| Figure 3. Schematic showing how EED might propagate H3K27me3 repressive chromatin marks. |
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.