Towards better understanding of retroviral DNA integration and more effective antiviral drugs
Sep 28, 2009
Sep 28, 2009
Understanding how viruses such as HIV function is vital in the search for more effective antiviral medicines, as well as opening up novel possibilities in gene therapy. A collaborative study involving researchers from Imperial College London, Yale University, and Dana-Farber Cancer Institute, led by Dr. Peter Cherepanov from the Division of Medicine at Imperial took advantage of Diamond’s I02 and I04 macromolecular crystallography beamlines to reveal important mechanistic aspects of retroviral replication.
Retroviruses are a family of viruses that carry their genetic material as RNA (ribonucleic acid), rather than DNA. This is converted into DNA by enzymes they bring with them as they enter a host cell. For infection to be effective, the viral DNA has to be inserted into the genome of the cell, thus making it a permanent carrier of the viral genetic material. This process requires integrase, the essential viral enzyme that cuts a host cell chromosome and pastes in the viral DNA. HIV integrase is an important drug target, and disrupting its function has already produced potent antivirals. In addition, a greater understanding of retroviral integration will aid the development of gene therapy applications, which utilize modified retroviruses as vehicles for delivery of genetic material.
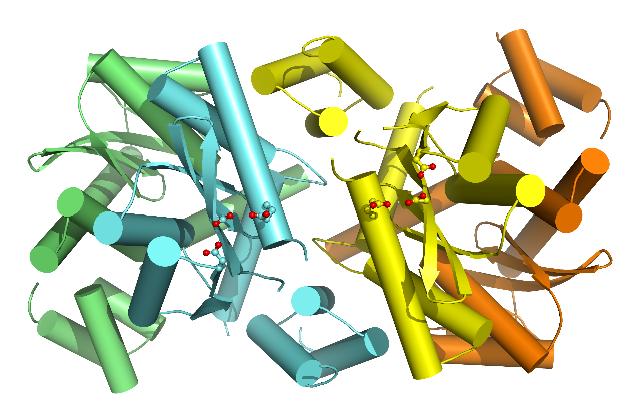 |
| The closed tetrameric structure of MVV integrase. One integrase dimer is coloured cyan and green and the other yellow and orange. The amino acids of the central active sites, correctly arranged for joining both ends of the viral DNA to the host DNA, are shown in "ball and stick" representation, with their acidic groups coloured red. |
HIV belongs to Lentivirus, a retroviral genus whose members have the unique ability to efficiently infect terminally differentiated non-dividing cells, such as neurons. Lentiviruses are therefore of great interest for gene therapy as gene delivery vectors, whereby genetic material can be delivered into cells as part of the treatment process. During integration lentiviruses highjack a cellular chromosomal protein called LEDGF. By virtue of binding to LEDGF, lentiviral preintegration complex is directed towards the most suitable regions of the host cell genome for integration.
In the first of two recent studies published in PLoS Pathogens, the researchers used X-ray crystallography to reveal a structure of a two-domain fragment of HIV integrase in complex with LEDGF. This work elucidated the entire integrase-LEDGF molecular contact surface and suggested how gene therapy vectors could be retargeted using engineered proteins derived from LEDGF.
A long-standing question in the field has been about the architecture of the active integrase multimer. Despite considerable efforts in many laboratories, researchers have yet been unable to obtain the complete three-dimensional structure of the viral protein. Because both ends of a viral DNA molecule must be joined to a host cell chromosome, the enzyme must provide a pair of active sites, where the joining reactions occur, in a suitable geometry. The catalytic core domain of integrase is known to form a dimer – two molecules paired together, however the pair of active sites in the dimer are too far away from each other. Such structural insights and functional data suggested that integrase must minimally form a tetramer, or more precisely a dimer-of-dimers, to accomplish its function.
“In our second study, we looked at integrase from maedi-visna virus (MVV), a retrovirus of sheep, functionally similar to but genetically divergent from HIV. By producing a two-domain fragment of this integrase with LEDGF we obtained a protein-protein complex, which we were able to crystallize in two forms. The resulting crystal structures revealed a common overall tetrameric arrangement of integrase with considerable flexibility at the dimer-dimer interface. We were very excited when we observed, for the first time, a tetramer in the closed conformation in one of our structures. Importantly, the closed tetramer features the correct arrangement of the active sites and appears compatible with the integration process.’’Dr Stephen Hare, Imperial College London
"This new structural information, allowed us to speculate how the active integrase tetramer forms and how it binds cellular DNA. However, although the partial structures tell us a lot about the mechanism of integration, some very important nuances will only be revealed when the structure of a full-length retroviral integrase protein in complex with its cognate viral DNA will be solved. Determination of such structure is the next big challenge.”Dr Peter Cherepanov
Hare S, Shun MC, Gupta SS, Valkov E, Engelman A and Cherepanov P. (2009) A novel co-crystal structure affords the design of gain-of-function lentiviral integrase mutants in the presence of modified PSIP1/LEDGF/p75. PLoS Pathogens, 5(1):e1000259. Data collected at the ESRF.
Hare S, Di Nunzio F, Labeja A, Wang J, Engelman A and Cherepanov P. (2009) Structural basis for functional tetramerization of lentiviral integrase. PLoS Pathogens, 5(7):e1000515. Data collected at Diamond Light Source.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.