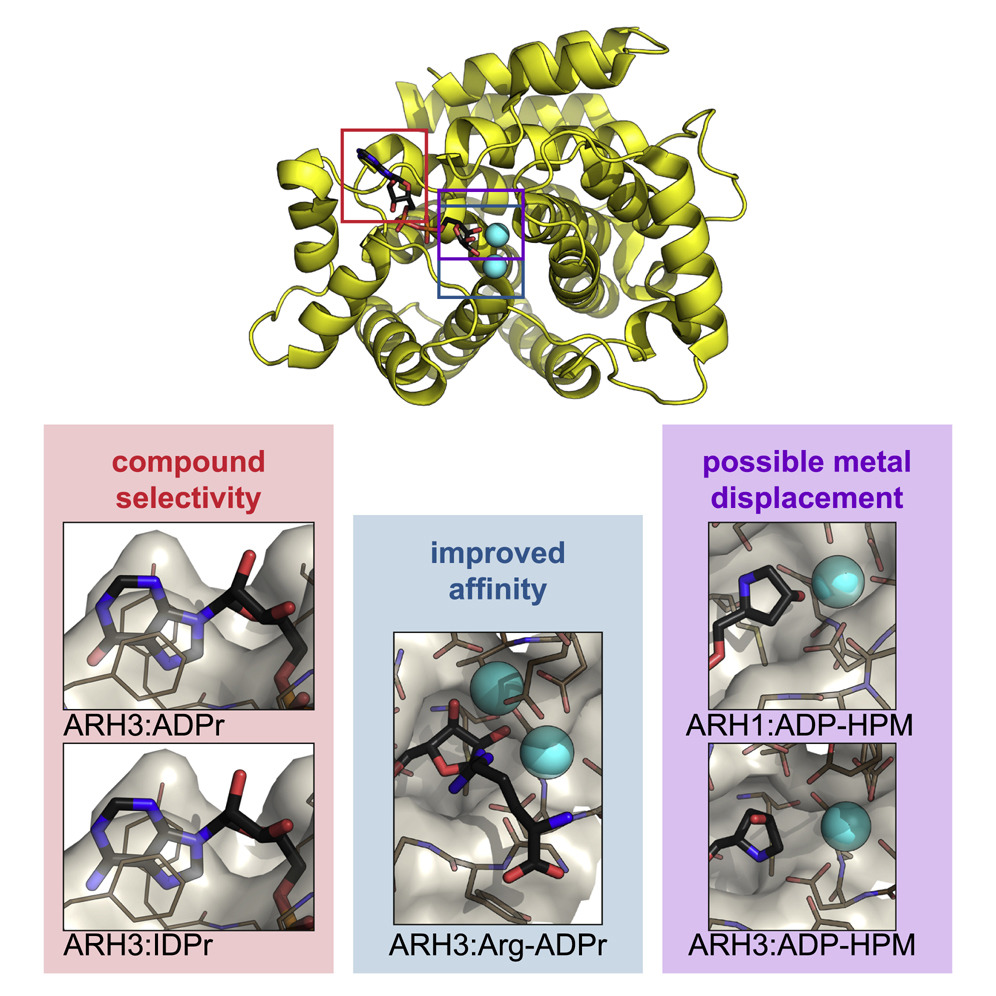
When cells suffer DNA damage, they send out an SOS signal. When the repair crew arrives, the emergency signal is cancelled as it is no longer needed. This two-stage process is an important one in many different diseases, but the underlying mechanisms are not well understood. ADP-ribosylation is one of the DNA damage response mechanisms used by cells to send out an SOS signal; and the ARH family of enzymes is involved in cancelling the signal. In work recently published in Cell Chemical Biology, researchers from the University of Oxford's DNA Damage Response Lab investigated the structure of two ADP-(ribosyl)hydrolases (ARHs). Their results provide important insights into the differences between these two proteins and offer useful tools for targeted drug design.

Experiments for this research were carried out on beamlines I03, I04, I04-1, and I24. The ARH proteins are magnesium-dependent, and their structure had to be determined both with and without magnesium present. The paper, therefore, contains ten protein structures, collected at atomic resolution. This kind of intensive research was possible because the University of Oxford has BAG access to Diamond, which allows groups of users to fill a beamtime shift with a series of short experiments.
Dr Antonio Ariza explains:
Before Diamond opened, I had to fly to Grenoble to conduct MX experiments, which was a considerable time investment. Experiments are so quick now, and Diamond makes continuous improvements. I visit Diamond several times every month now. Many of our experiments could be automated, but I like coming in person to get my hands dirty, and to keep up with the changes.
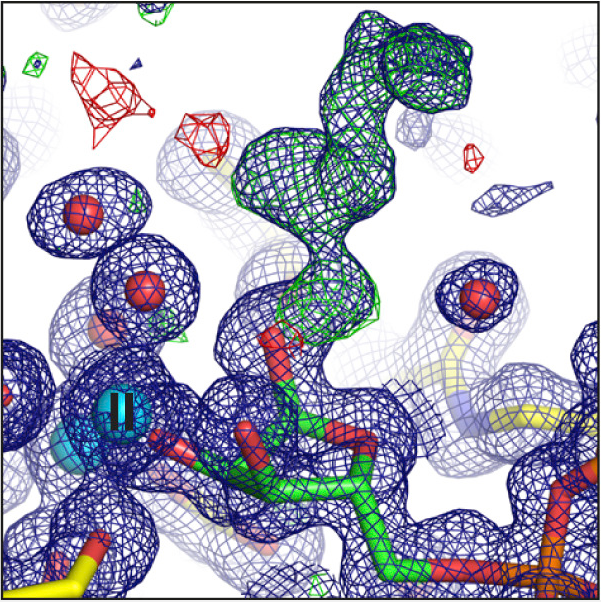
Rack JGM et al. (ADP-ribosyl)hydrolases: Structural Basis for Differential Substrate Recognition and Inhibition. Cell Chemical Biology 25.12 (2018). DOI:10.1016/j.chembiol.2018.11.001.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.