Copper nanoparticles (Cu-NPs) have a wide range of applications as catalysts, in scientific fields as diverse as drug discovery and materials science. The natural abundance of copper, and its relatively low cost, makes it a viable alternative to catalysts made from rare and expensive precious metals, such as platinum and palladium. However, the synthesis of Cu-NPs usually involves high temperatures and toxic solvents. Additionally, Cu-NPs produced via conventional synthesis tend to agglomerate and oxidise, and require the use of inorganic chemicals to maintain their catalytic activity. New research, published in Small, details proof-of-concept experiments demonstrating that the metal-reducing bacterium Shewanella oneidensis offers a greener route to Cu-NP synthesis, and the potential to reclaim copper from wastewater streams.
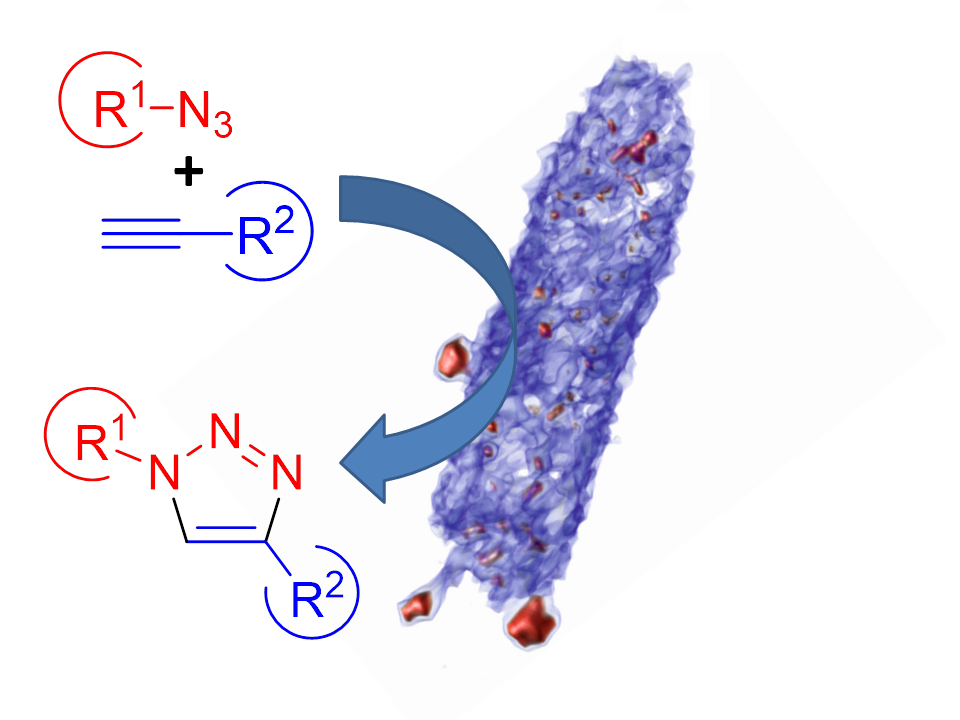 Figure 1: Schematic representation of ‘click-chemistry’ reactions catalysed by biosynthesised copper nanoparticles. The image shows a 3D representation of a Shewanella oneidensis cell (blue) and associated copper nanoparticles (red) reconstructed from serial block-face scanning electron microscope images.
Figure 1: Schematic representation of ‘click-chemistry’ reactions catalysed by biosynthesised copper nanoparticles. The image shows a 3D representation of a Shewanella oneidensis cell (blue) and associated copper nanoparticles (red) reconstructed from serial block-face scanning electron microscope images.
Bacterial biosynthesis of Cu-NPs
If we can harness the metabolism of metal-reducing bacteria, this gives us a route to cheap, simple and environmentally-benign nanoparticle synthesis. This is the first study to investigate the bioreduction of soluble copper(II) ions and the synthesis of Cu-NPs using anaerobic metal-reducing bacteria, organisms that exist naturally in anaerobic sediments, and gain there energy by transferring electrons from organic matter to metals in the sediments. Shewanella oneidensis is one of the most versatile and well-studied species of metal-reducing bacteria, able to reduce a wide range of metals under laboratory conditions. It was first isolated in 1988, by Professor Ken Nealson, from sediments in Lake Oneida in New York (from where it takes its name). It was chosen for these experiments due to its versatility as a metal-reducer and having its entire genome sequenced. The resulting availability of mutant strains allows investigation of the pathway involved in metal reduction (e.g. the enzymes involved). Identifying the electron transfer pathway involved in Cu reduction could lead to efficiency improvements in future. The results demonstrate that it is possible to use Shewanella oneidensis for the bioreduction of copper(II) ions, forming elemental Cu(0) nanoparticles, which in itself is surprising as many forms of copper are known to be toxic, being used as a disinfectant and a fungicide, and has been investigated for use in anti-microbial surfaces.
This new process ticks all the boxes for ‘green synthesis’, as it is able to produce Cu-NPs at room temperature, in water. In addition, during the catalysis tests the Cu-NPs were not separated from the biomass, and the bacterium acted as a support matrix for the nanoparticles - removing the need for inorganic additives and making the Cu-NPs more reactive. Finally, the catalyst can be easily filtered out using a centrifuge, allowing it to be reused.
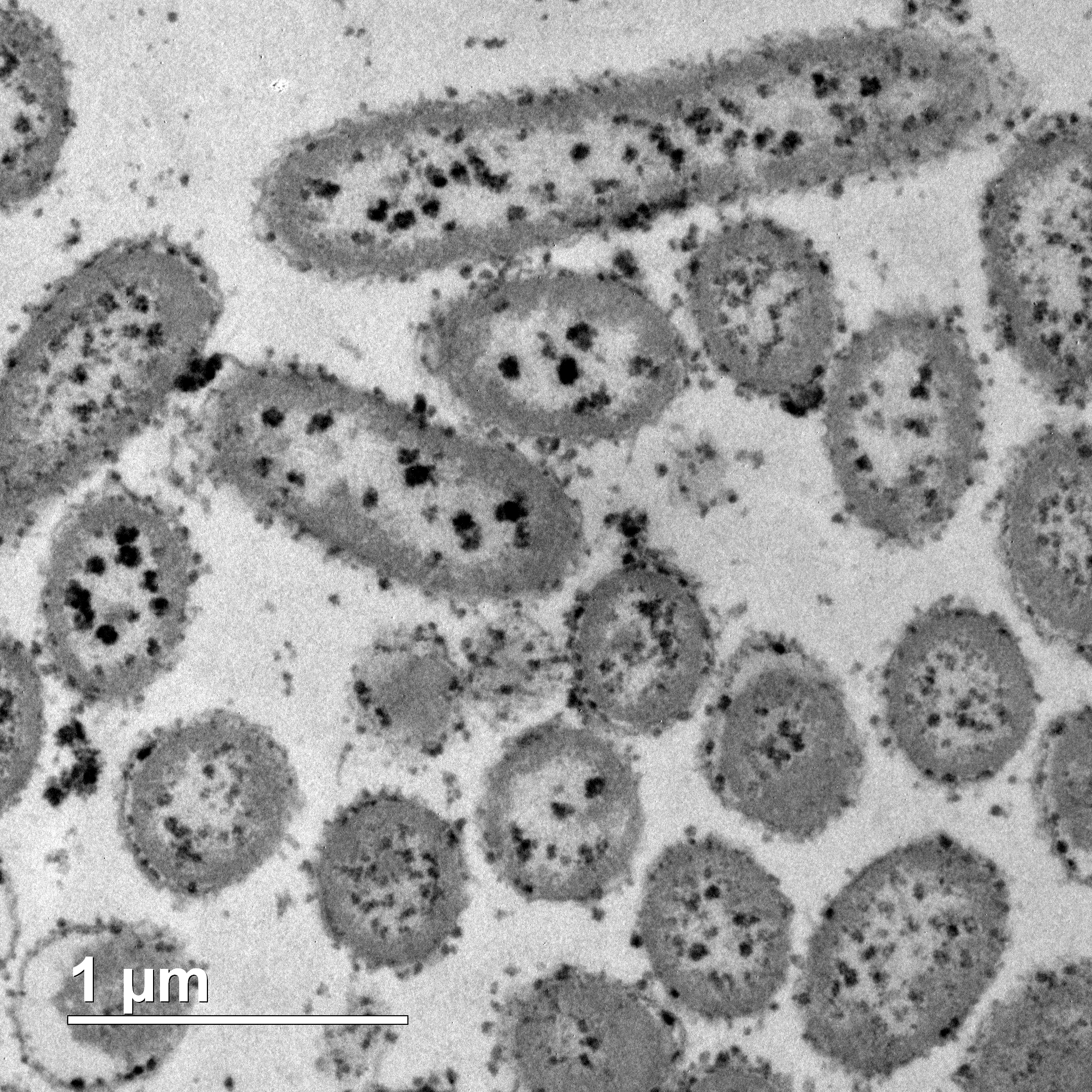 Figure 2: Thin section TEM image of Shewanella oneidensis and biosynthesised copper nanoparticles.
Figure 2: Thin section TEM image of Shewanella oneidensis and biosynthesised copper nanoparticles.
“Where there's muck there's brass”
The NERC-funded experiments used Diamond used X-ray absorption near-edge spectroscopy (XANES) and extended X-ray absorption fine-structure spectroscopy (EXAFS) analysis on B18 (a general purpose XAS beamline) to show that the nanoparticles produced are copper, and to identify its oxidation state. Soft x-ray XAS measurements were done using Diamond’s I10 beamline. These initial investigations used metal salts, but the research team is moving on to look at using industrial wastewater streams. For lead author, Dr Richard Kimber of the School of Earth and Environmental Sciences at the University of Manchester, this is the ultimate goal of the project. He says, “It’s important to recover metals from wastewater, to prevent them from contaminating the environment. What we’re looking at here is a way to create high-value products from waste treatment, so that it will pay for itself.”
Future work will also investigate ways to optimise the system, including determining optimal reaction times and Cu-NP loading, to improve yields. There’s also work to be done on understanding the pathway the bacterium uses to reduce copper. Usually metal-reducing bacteria use common metals (such as iron) for respiration. The initial results suggest that this isn’t the case for Shewanella oneidensis and copper, perhaps unsurprisingly given the toxic nature of the metal. The electron transfer pathway could be part of the bacterium’s detoxification/defence mechanisms, but more work needs to be done. An understanding of the pathway would make it easier to increase the yield of Cu-NPs produced as well as potentially fine-tune their properties. With this in mind, the Manchester-based research team is keen to use the latest advances in synthetic biology to make the next generation of catalysts for industry. Prof Jon Lloyd who leads the research in this area (alongside colleagues in Manchester’s Institute of Biotechnology) notes “this new study gives us a new type of metallic nanocatalyst that we hope will be very useful for the chemical industry, and we are very keen extend the utility of this approach via the incorporation of additional catalytic materials (enzymes and other metallic nanoparticles) into the host cells that we used for this current study. This work forms the basis of a new BBSRC project for the team.”
To find out more about the B18 beamline, or to discuss potential applications, please contact Principal Beamline Scientist Dr Giannantonio Cibin: [email protected]; regarding the I10 beamline, please contact Principal Beamline Scientist Dr Paul Steadman: [email protected]
Kimber, RL et al. Biosynthesis and Characterization of Copper Nanoparticles Using Shewanella oneidensis: Application for Click Chemistry. Small (2018). DOI: 10.1002/smll.201703145
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.