The inner workings of a lethal giant freshwater prawn virus have been revealed by an international team of researchers using data gathered at Diamond Light Source. The results reveal a possible new class of virus and presents the prospect of tackling a disease that can devastate prawn farms around the world.
The detailed structure of a virus that can devastate valuable freshwater prawn fisheries has been revealed by an international team using image data collected in the Electron Bio-Imaging Centre (eBIC) based at Diamond Light Source. The researchers produced high-resolution images of virus like particles, VLP’s, composed of virus shell proteins which they compared with lower resolution images of the complete virus purified from prawn larvae. They found strong similarities between the two suggesting that the more detailed VLP images are a good representation of the intact virus. This research, exposing the inner workings of the MrNV, could make it easier to develop ways of combating the economically important disease, but also suggests that it belongs in a new, separate, group of nodaviruses.

The researchers used the rapidly developing technique of cryo-electron microscopy, cryoEM, which has the ability to produce very high-resolution images of frozen virus particles. Images so detailed that the positions of individual atoms could be inferred. Recent breakthroughs in this technique have transformed the study of relatively large biological complexes like viruses allowing researchers to determine their structures comparatively quickly. The data to produce the MrNV structure described here was captured in two days at the eBIC facility.
The virus, Macrobrachium rosenbergii nodavirus or MrNV, causes white tail disease that infects the larvae of the giant freshwater prawn M. rosenbergii. An outbreak can kill all the larvae it infects. The prawn is widely cultivated in Asia and the Americas, with some production in Africa and the Middle East. Adult males can reach up to 35cm and are an increasingly valuable food source. Fresh water prawn farming has grown dramatically in the last two decades, global production is more than 200,000 tonnes a year, and is considered less environmentally harmful than the equivalent salt water prawn farming. Outbreaks of MrNV are a significant threat to both individual farms, those employed in the industry and food security more widely.
At present there is no vaccine or treatment available to prevent outbreaks of the disease, outbreaks of which have occurred across much of China and South-East Asia. It is well established that a detailed knowledge of the structure of a virus provides researchers opportunities to develop vaccines and treatments. It is hoped that the results from this international team of researchers might lead to new ways to tackle MrNV and also provide new insight into this family of viruses.
MrNV is a relatively simple virus consisting of a capsid protein that makes up its shell, and a small fragment of RNA that carries its genetic information and an enzyme responsible for helping it infect its host. The researchers were able to isolate the gene for the capsid protein and put it into another virus which was in turn grown inside insect cells in the laboratory. This produced large amounts of the protein that they were able to extract and purify and is a common, well-established method for studying viruses. Furthermore, when grown in this way the capsids protein molecules spontaneously organised themselves inside the insect cells into virus like particles that resembled the naturally occurring virus. Again a common and well established protocol.
As a result of this work, the researchers had sufficient quantities of VLP’s to start to determine their structure. This was done by the rapidly developing technique of cryogenic electron microscopy, or cryoEM.
David Bhella, programme leader in the Medical Research Council’s University of Glasgow Centre for Virus Research, who led the research, described how the samples were all prepared in his Glasgow laboratory. A solution containing the VLP’s was spotted onto a small TEM grid overlain with a thin perforated carbon film, and then frozen quickly to around -190°C. They were then transported frozen to the eBIC facility at Diamond ready for imaging. Dr Bhella and his Malaysian colleague Kok Lian Ho went with the samples as Dr Ho was keen to learn more about the cryoEM process.
However, Dr Bhella said that one of the strengths of the eBIC facility is the staff are very skilled and you can just send the samples on the grids, greatly speeding up and simplifying the process. And the high degree of automation means that data can be collected rapidly both night and day.
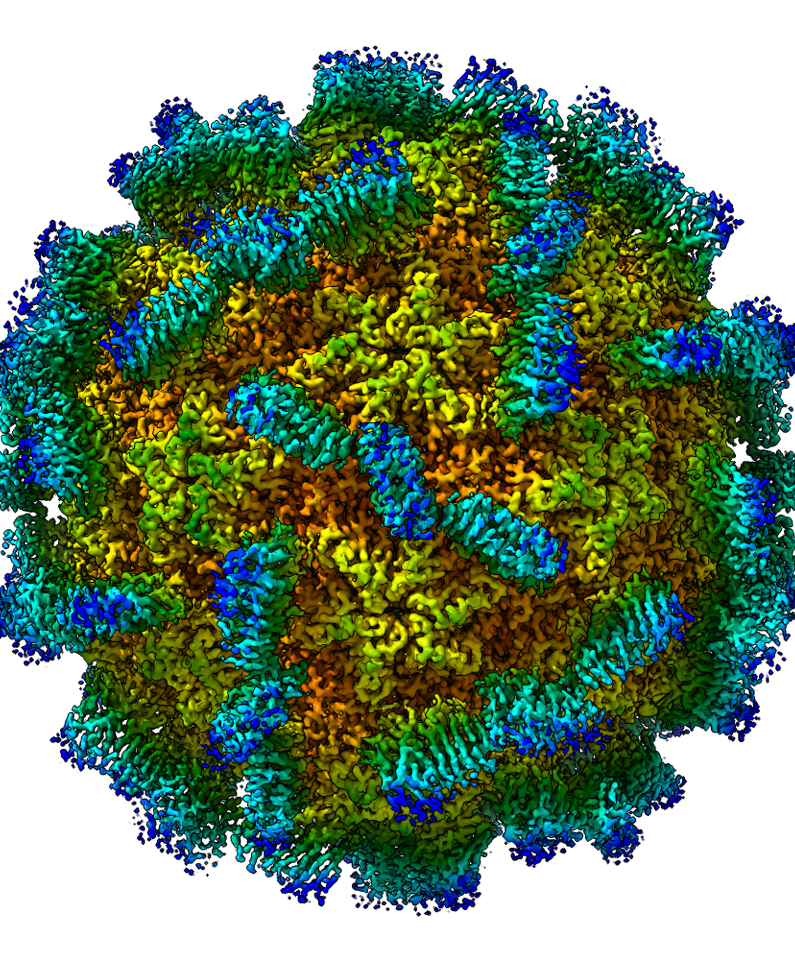
Once at eBIC, the samples were placed in a cryoEM machine, kept at -195°C, and over a period of a two days, images were taken of almost 50,000 individual VLP’s frozen fast to the cryoEM grid. These data were then used by the researchers to produce one composite image that revealed the structure of the VLP’s at very high resolution (see picture).
There was, though, a major outstanding question. These VLP’s were not the full, intact, naturally occurring virus and it was vital to know how closely they resembled the real thing. To find out, the researchers purified active virus from the larvae of infected giant prawns and subjected them to the same type of cryoEM investigation. They were not able to get quite as detailed images, the resolution was around 6 Angstroms [One Angstrom is one ten billionth of a metre] as compared to 3.3 Angstroms, but they were sufficient to show that the VLP’s bore a very close resemblance to the naturally occurring virus particles. An important validation that allows the researchers to be confident that the finer details in their VLP images did indeed match the structure of the native virus.
It is not possible to vaccinate crustaceans like the giant freshwater prawn in the same way as it is possible to vaccinate mammals such as ourselves against various diseases. So it is unlikely that this new MrNV structure will lead to a classic anti-white tail vaccine for the industry. However, developing a drug or other way of combating the disease has just got that bit easier. Anti-viral drugs have to target the virus they are intended to stop without harming their host. To do that researchers need to identify features unique to the virus and a high resolution structure helps significantly with this task.
The structure of the MrNV VLP’s bears some similarities with related viruses but also some striking differences. The key features are a spike on the surface that is likely to be involved in the way the virus binds to and then enters the cells it infects. The spikes of viruses in the same group as MrNV are made up of three capsid protein molecules. However, the MrNV spikes have just two. Also, the MrNV has a metal binding site that more closely resembles those from a different group of viruses, the tombusviruses. Furthermore, the MrNV capsid proteins have a long arm that reach around inside the virus, holding onto other proteins and helping keep the assembly together. This arm, though, is longer than in other viruses and one of the longest ever discovered. These features, as well as others, suggest that the MrNV should in fact be given its own, new, sub classification. Dr Bhella described how virus evolution is hard to understand because their genomes mutate so much. As a result viruses tend to be classified on other aspects of their biology, such as the hosts they infect or the properties of their internal proteins. Or, as might be the case here, the structure of their capsids. At the time of writing this new classification remains just a suggestion and has not been widely agreed.
To find out more about eBIC please contact the Centre Director Peijun Zhang peijun.zhang@diamond.ac.uk
Ho K L, Gabrielsen M, Beh PL, Kueh CL, Thong QX, Streetley J, Tan WS, Bhella D. Structure of the Macrobrachium rosenbergii nodavirus: A new genus within the Nodaviridae? PLOS Biology 16 doi.org/10.1371/journal.pbio.3000038 (2018)
The authors wish to thank Diamond Light Source for access to the cryoEM facilities at the UK National eBIC.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.