Two wrongs to make it right
Sep 26, 2017
Sep 26, 2017
A variant of the NQO1 gene is linked with increased susceptibility to cancer partly because it reduces the affinity between NQO1 and its molecular partner FAD. However, the addition of a second change to the NQO1 sequence reverses this effect, restoring the normal FAD-binding affinity and functionality of NQO1. In a study published in FEBS Letters, a team led by Dr Angel Pey of the University of Granada unravels how these two changes affect the interaction between NQO1 and FAD despite being far from the FAD-binding site. Using the High Throughput Small Angle Scattering beamline (B21) at Diamond Light Source, together with molecular dynamics and thermodynamics analyses, they discovered that the two changes shift the equilibrium conformation of NQO1 between FAD-binding competent and non-competent forms, altering the protein’s affinity for FAD and its activity in cells. By combining several different approaches to investigate the malleability of the structure of NQO1, this study clarifies how single amino acid changes can significantly alter a protein’s structural and mechanistic features.
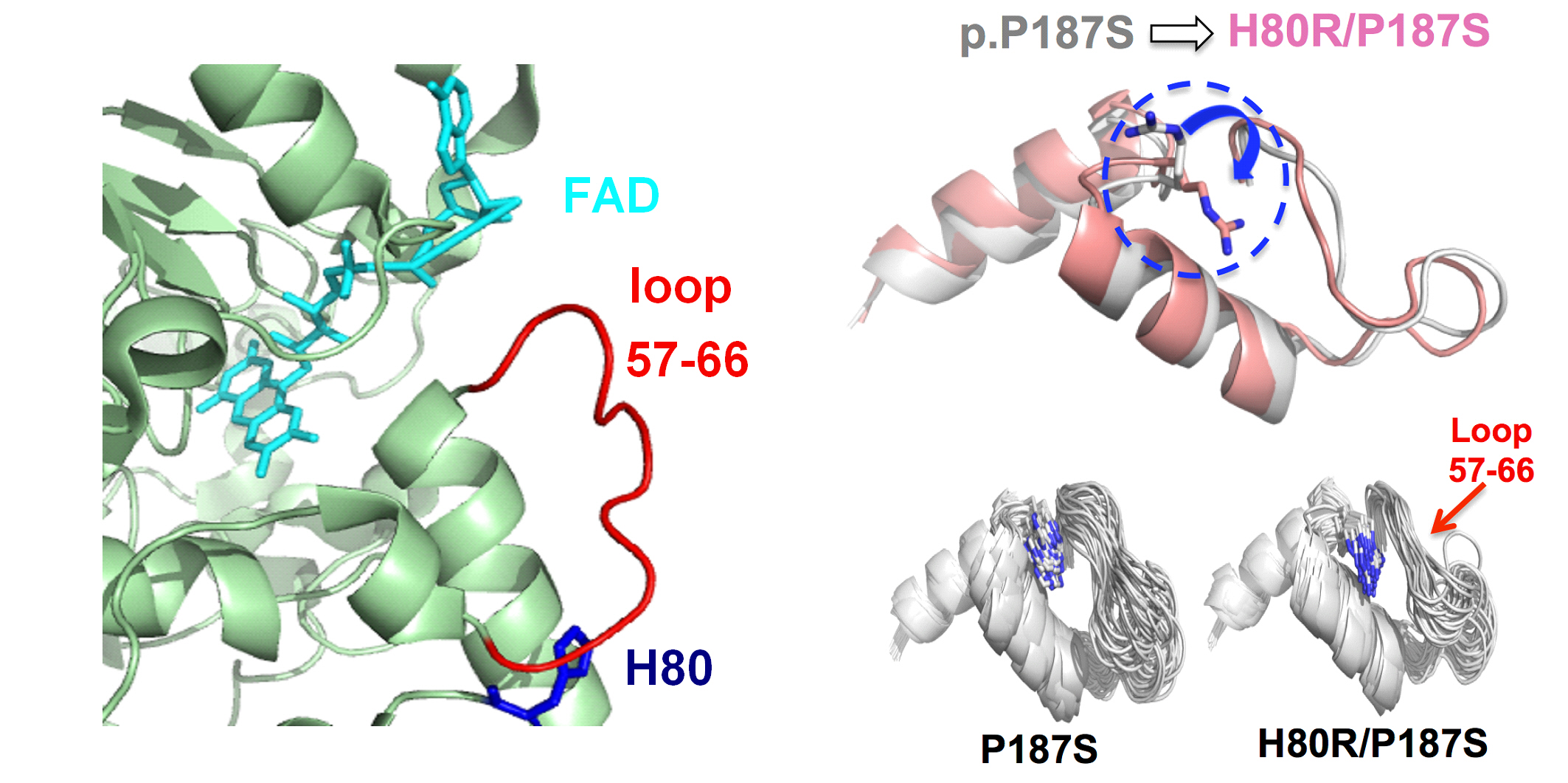
A flexible problem
The NQO1 gene encodes an antioxidant enzyme that stabilises transcription factors involved in regulating the cell cycle and also plays a role in the activation of several anti-cancer drugs. Some people carry a version of the protein with a single amino acid change, known as P187S, which increases their risk of developing cancer. NQO1 activity depends on binding the molecule FAD, and the P187S change reduces FAD binding and accelerates the protein’s degradation.
An international team of researchers led by Dr Angel Pey, assistant professor of physical chemistry at the University of Granada, recently published a study1 in Human Molecular Genetics showing that the effects of P187S are counteracted by a second amino acid change in NQO1, H80R. When both P187S and H80R are present, the protein’s ability to bind FAD is rescued and its activity within the cell recovers. Despite their effect on FAD binding, neither of the amino acids altered in these changes are part of the protein’s FAD-binding site, making their connection with FAD-binding affinity a mystery.
A significant challenge in unravelling this mystery is that the unbound form of NQO1 is quite flexible and dynamic. “The unbound state has a lot to say about the effects of the mutation, but it’s extremely difficult to crystallize because it’s too flexible,” explains Dr Pey. Without more precise data about the protein’s structure and its variability, a full understanding of how the amino acid substitutions affect its activity would remain out of reach. “So we had to look for other ways of inferring that information.”
A solution in solution
Working with one of the researchers involved in the earlier study and two new collaborators, Dr Pey brought together several techniques to shed light on the protein’s structure. “I truly believe that from this combination of techniques, structural approaches and thermodynamics and molecular dynamics, you can get a very nice, complete picture of what’s going on,” he says.
The team’s molecular dynamics calculations predicted the distribution of conformations that the unbound protein would take. Using small-angle X-ray scattering measurements with the B21 beamline at Diamond, they were able to avoid the need for crystallisation and probe the structure of the flexible, unbound NQO1 molecules in a solution. Together with a thermodynamic analysis and additional solution spectroscopic analyses, the structural information revealed that the H80R change causes the equilibrium conformation of NQO1 to shift slightly to favour its FAD-binding competent form.
In other words, P187S makes NQO1 molecules more likely to have a shape that isn’t able to bind FAD, and H80R pushes the equilibrium back, making the protein slightly more likely to have a shape which can bind FAD. “The change must be slight, but even this small effect on the equilibrium translates directly into a change in the binding affinity, which then leads to a significant increase in the activity of the protein inside cells,” explains Dr Pey.
A human peculiarity
With a fuller picture of the structural role of these amino acid changes in place, Dr Pey is turning towards understanding their evolution and impact. In the H80R substitution, the amino acid histidine changes to an arginine, stabilising NQO1. However, this histidine appeared in NQO1 during the evolution of primates; while humans have a histidine at this position in the protein, most mammals have an arginine. Dr Pey plans to test whether simply switching the arginine to a histidine in another mammal, such as a mouse, will make its NQO1 as sensitive towards cancer as the human P187S variant.
Understanding the evolution of the different NQO1 variants may help shed light on what function these differences have, if any. Likewise, understanding the mechanistic properties of the variants may eventually make it possible to design molecules that mimic the stabilising variants and help maintain proper NQO1 function. “I usually try to get a deep understanding at all the levels I can and combine our techniques with others like cellular work,” says Dr Pey. “And then with a complete picture of the disease mechanism, we can try to focus on identifying a way – maybe a pharmacological way – of overcoming problems.”
To find out more about the B21 beamline, or to discuss potential applications, please contact Principal Beamline Scientists Dr Robert Rambo: [email protected]
Muñoz, IG et al. A mechanism for cancer-associated inactivation of NQO1 due to P187S and its reactivation by the consensus mutation H80R. FEBS Letters (2017). DOI: 10.1002/1873-3468.12772
1. Medina-Carmona, E et al. Enhanced vulnerability of human proteins towards disease-associated inactivation through divergent evolution. Human Molecular Genetics 26, 3531-3544 (2017).
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.