The membrane of a cell is a double-layered structure that is composed of different types of lipids, which can help to support a variety of membrane proteins embedded within them. One such protein, the antibacterial peptide ABC transporter, McjD, is heavily dependent on its neighbouring lipids. McjD transports toxic peptides out of bacterial cells to help protect them, but the specific lipids that assist this activity have not yet been identified.
The Microfocus Macromolecular Crystallography and Circular Dichroism beamlines, I24 and B23, at Diamond Light Source were used to solve the structure of McjD in complex with membrane lipids. The researchers also removed and replenished the lipids from the protein to see how they affected the overall structure. Removal of the lipids caused the structure of McjD to deteriorate and the protein lost some of its activity; an effect which was revered upon re-addition of the lipids. Mass spectrometry was used to identify the mystery lipids that are important for its activity and it was discovered that they each possessed differing structures and chemical properties, which worked together synergistically to support McjD.
The results from the study are invaluable for understanding the function of the ABC transporters. Many of these proteins are involved in bacterial multi-drug resistance, so this information is vital for prolonging the usability of our antibiotic arsenal.
The lipid bilayer consists of a mixture of lipids with different chemical and structural properties that regulate the function of membrane proteins; these lipids are either annular, masking the protein hydrophobic surface and are usually not essential for protein function, or specific lipids, essential for protein function. Loss of specific lipids (eg. purification) usually results in aggregation or reduced transport and enzymatic activity. Adenosine triphosphate (ATP)-binding cassette (ABC) transporters are one of the largest superfamilies found in eukaryotes, archaea and prokaryotes, and they are associated withmulti drug resistance in both bacterial and eukaryotic cells1. ABC transporters are powered by the hydrolysis of ATP. In order to understand how specific lipids modulate the activity of ABC exporters, the antibacterial peptide ABC transporter McjD from E. coli was selected as a case study. McjD is involved in the export of the antibacterial peptide microcin J25. The previously determined structure had shown that detergent purified McjD hydrolysed ATP at a lower rate compared to proteoliposome reconstituted protein, suggesting that the lipid environment was important for its activity2.
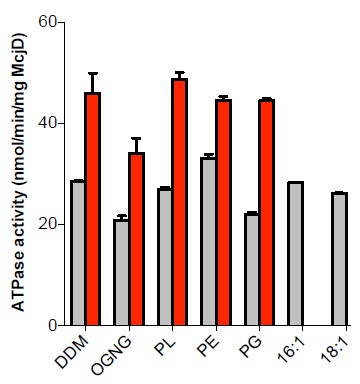
This study sought to establish what lipids are associated with McjD and how they affect its function and structure3. Using tandem mass spectrometry of detergent purified (lipidated) and delipidated samples, 11 lipid species were identified, 2 negatively charged (phosphatidylglycerol (PG)) and 9 zwitterionic (phosphatidylethanolamine (PE)), for the lipidated protein. Delipidation of McjD resulted in the loss of several zwiterrionic lipids. The effect of the different lipid types identified on the ATPase activity of McjD was investigated by partially delipidating it. The basal ATPase activity of delipidated McjD was reduced by 30% relative to the lipidated protein (Fig. 1). When the delipidated protein was supplemented with detergent-destabilised small unilamellar vesicles composed of E. coli polar lipid extract (PL) its basal ATPase activity was restored. The PL extract is a mixture of PE, PG and cardiolipin (CARD), and the ATPase activity of delipidated McjD was measured in the presence of different detergent-destabilised small unilamellar vesicles composed of different lipids. PG was not able to fully restore the basal ATPase activity, whereas PE was able to restore it. The ligand Hoechst stimulated the lipidated and delipidated McjD by 2.1 and 1.6 fold, respectively, relative to the basal ATPase activity of the delipidated McjD (Fig. 1). When delipidated McjD was supplemented with lipids, the ATPase activity in the presence of Hoechst almost doubled relative to the basal ATPase activity of delipidated McjD.
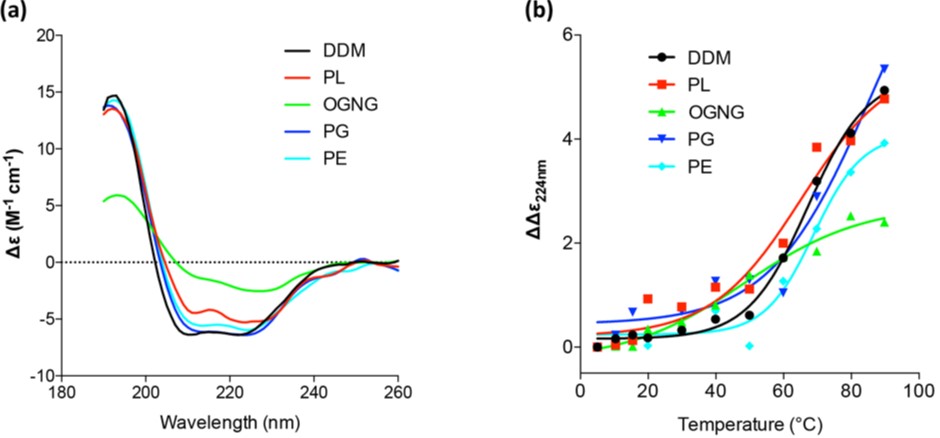
Since the activity of the delipidated protein was restored in the presence of various lipids, the effect of different lipid environments on the overall protein architecture was studied by measuring the secondary structure and melting temperature, Tm, of McjD using Synchrotron Radiation Circular Dichroism (SRCD) at B23. The far ultraviolet Circular Dichroism (UV-CD) spectra oflipidated McjD showed a typical α-helical structure, with an α-helix content of ca. 68% (Fig. 2a) that is similar to our published crystal structure in dodecyl-maltopyranoside (DDM). Delipidated McjD resulted in a more ”aggregated” structure with a higher content of β-strand (26%) and a significant reduction in a-helical content (29%) compared to lipidated McjD. McjD supplemented with PL, PE and PG showed similar CD features to that in DDM, with a comparable a-helix content of 64-68 %, suggesting that the lipids are capable to revert the loss of the secondary structure (Fig. 2a). The thermal stability of McjD in the different lipid environments was measured as a function of secondary structure loss upon heating (Fig. 2b). Lipidated McjD has a Tm of 67 °C and delipidation significantly reduces its Tm to 48 °C. Delipidated McjD supplemented with PL, PE and PG has a Tm of 65 °C, 69 °C and 74 °C (and a second transitions at 13 °C) respectively. The α-helix component in McjD in the various lipids (PL, PG and PE) gave higher Tm (>70 °C) compared to McjD in DDM (66 °C), suggesting that other secondary structure components of McjD were likely perturbed before the α-helix content was affected.
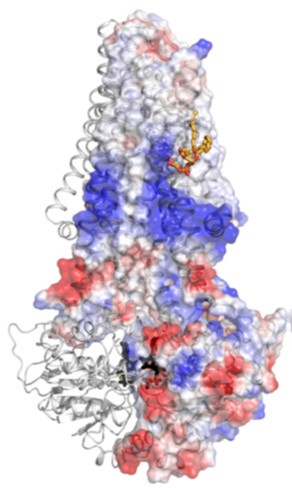
Previous studies have shown that inclusion of low-resolution diffraction spots in the refinement and map calculations can allow the identification of bound lipids to membrane proteins; using this technique, a structurally resolved specific lipid was determined in the McjD structure that was assigned as PG (18:1/18:1) based on the above data (Fig. 3). In the refined structure, the PG lipid is weakly associated with residues from TMs 3 and 4 and probably stabilises.
In conclusion, this study has demonstrated the synergistic role of specific lipids on the modulation of the structure and activity of the ABC exporter McjD. Partial delipidation of McjD reduced its activity and affected its overall structure that could be restored with the addition of different classes of lipids. The data suggest that the role of PE is to provide overall structural stability to McjD whereas PG is involved in stabilising different transitions.
References:
Funding acknowledgement:
The Oxford University Mass Spectrometry facility is funded by ERC IMPRESS (26851) and the Medical Research Council (98101). C.V.R. is a Royal Society Research Professor. Work in DPTs group is supported by the Canadian Institutes for Health Research (MOP-62690). Part of this work was also supported by the Biotechnology and Biological Sciences Research Council (BB/H01778X/1 to KB).
Corresponding author:
Dr Konstantinos Beis, Imperial College London, [email protected]
Related publication:
Mehmood S, Corradi V, Choudhury H, Hussain R, Becker P, Axford, D, Zirah S, Rebuffat S, Tieleman D, Robinson C, Beis, K. Structural and Functional Basis for Lipid Synergy on the Activity of the Antibacterial Peptide ABC Transporter McjD. Journal of Biological Chemistry 291, 21656-21668 doi: 10.1074/jbc.M116.732107 (2016).
Publication keywords:
ABC transporter; Circular dichroism (CD); Lipid; Membrane protein; Membrane transport; Molecular dynamics; Structural biology; Structure-function; Delipidation; Non-denaturing mass spectrometry
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.