Annual Review: MX
Aug 5, 2016
Aug 5, 2016
Related publication:
Yang Y, Darbari VC, Zhang N, Lu D, Glyde R, Wang Y-P, Winkelman JT, Gourse RL, Murakami KS, Buck M, Zhang X. Structures of the RNA polymerase-σ54 reveal new and conserved regulatory strategies. Science 349, 882-885, doi:10.1126/science.aab1478 (2015).
Keywords:
RNA polymerase; Transcription; Sigma factor; Regulation.
RNA polymerase (RNAP) is a fundamental piece of cellular machinery required for making messenger RNA from DNA in all cell types. Despite the large amount of structural information that has been obtained over the last 15 years, the means by which RNAP is regulated is still not completely understood. In bacteria, the RNAP is assisted by proteins called sigma factors, which recruit RNAP to specific regions on the DNA called promoters. One such sigma factor, known as sigma54 (σ54), recruits RNAP during special circumstances like stress and infection and inhibits transcription until additional activators intervene.
Using Diamond Light Source’s Macromolecular Crystallography (MX) beamlines (I03 and I04), the interplay between σ54 and RNAP was explored. The crystal structure of RNAP in complex with σ54 was resolved to 3.8 Å, and it demonstrated that tight inhibition between the two proteins arose from large regions of interaction. All of the domains of σ54 were involved with preventing the activation of the polymerase. Furthermore, by comparing σ54 with other functionally-related regulatory factors in bacteria and eukaryotes, a number of regulatory hotspots within RNAP were discovered that might be targeted through a diverse range of strategies. This structural study serves to improve our mechanistic understanding of this fundamental cellular machine and also offers new insights into how RNAP, an antibiotic target, could be inhibited.
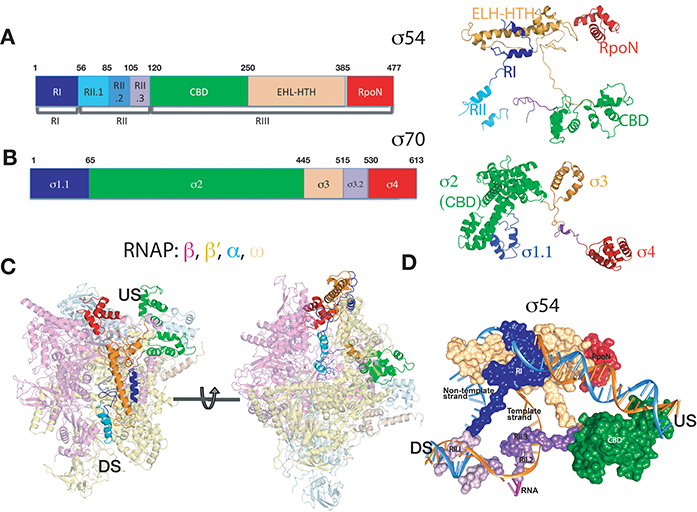
Figure 1: Crystal structure of σ54-RNAP. a) σ54 coloured by functional domains; b) σ70 coloured by functional domains. Same colour was used in similar functional domains between σ54 and σ70 ; c) σ54 (same colour as in (a)) in the context of RNAP (in semi-transparent); d) σ54 in surface representation overlapped with the DNA transcription bubble. Blue – non-template strand, gold – template strand. US – upstream, DS- downstream relative to transcription start site and direction.
The RNAP enzymes in bacteria, eukaryotes and archaea are closely related in structure and function, but become specialised through the use of accessory factors that control the RNAP during various stages of transcription1. Over the last 15 years, significant progress has been made towards understanding the transcription apparatus, including the high resolution structural determinations of bacteria, yeast and archaeal RNAPs. The significance of the work has been recognised by many awards including the Nobel Prize to Roger Kornberg on yeast RNA polymerase II. However, despite the advance, a detailed understanding on how RNAP is activated is still limited.
RNAP is promiscuous in binding to DNA and in bacteria, RNAP relies on dissociable sigma factors for recruitment to specific promoter regions for transcription2. Often a number of sigma factors exist in each bacterium. For example, there are seven sigma factors in E. coli with sigma70 (σ70) as the primary sigma factor responsible for transcribing housekeeping genes. Other sigma factors are utilised in transcribing genes in response to environmental changes including stress responses and pathogen virulence. Based on the functional similarities, sigma factors have been divided into two classes. The majority of sigma factors belong to σ70 class, characterised by recognising -35 and -10 promoter regions (numbering refers to the nucleotide upstream from the transcription start site) where transcription can proceed spontaneously after RNAP has formed a closed promoter complex (RPc)3. Activators of the σ70 type RNAP holoenzymes often increase holoenzyme affinity with cognate promoters. Contrasting σ70, the σ54 factor (also called sigmaN), is responsible for transcribing a wide range of stress responsive genes and forms a unique class of holoenzyme. It recognises -24 and -12 promoter regions, forms a stable RPc and rarely spontaneously isomerises to transcriptionally competent open complex (RPo), and so σ54-dependent systems show a major dependence on their cognate ATP-hydrolysing activators that are bound to upstream enhancer DNA4. These activators are thus called bacterial enhancer-binding proteins
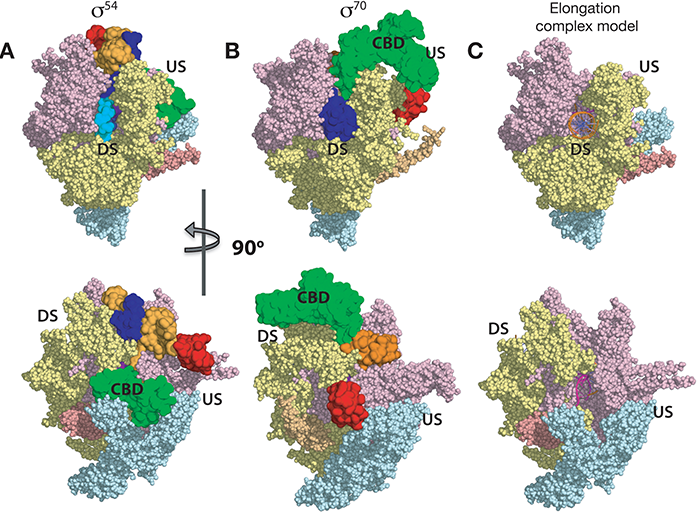
Figure 2: Comparison of σ54-RNAP and σ70-RNAP. Same color scheme as in Fig. 1. a) σ54-RNAP holoenzym; b) σ70-RNAP holoenzyme, σ54 and σ70 in surface representation, and RNAP in sphere representation; c) Elongation complex model showing the locations of DNA and synthesised RNA.
σ54 thus tightly inhibits bacterial RNAP core enzyme, which is itself competent for RNA synthesis. Moreover, the bacterial enhancer-dependent system represents a simplified system that is reminiscent to eukaryotic RNAP II, which requires TFIIH for open complex formation and shows enhancerdependent activation. The enhancer-dependent activators of the σ54-RNAP system can be considered as combining the functionality of TFIIH and enhancer-binding activators, which also require the mediator complex, into one simple system for activating and converting RNAPs from RPc to RPo. The bacterial σ54-RNAP system thus represents an ideal model system in studying RNAP inhibition as well as the transcription initiation process.
The crystal structure of σ54-RNAP holoenzyme, which amounts to 450 kDa (see Fig. 1), reveals some unexpected structural and functional features. This is the first non- σ70 bacterial holoenzyme structure and comparisons establish the novelty of the structures for function. Our structure reveals that σ54 contains four domains, with each making unique contributions to maintain RNAP in the stable RPc (Fig. 1). Specifically: Region I (RI) interacts with Region III (RIII) to form a lid that blocks the DNA template strand from entering the RNAP active cleft (Fig. 1B-C). Region II.1 (RII.1) blocks the downstream DNA channel while RII.2 and RII.3 occupy the space where the template strand and newly synthesised RNA would subsequently be located (Fig. 1D). Our structural comparisons with σ70 reveal that homologous functions between σ54 and σ70 are achieved by very different structural elements. For example, the RNAP corebinding domain (CBD) of σ54 lies upstream, also blocking the RNA exit channel (Fig. 2) while that of σ70 is located downstream. Remarkably the area on RNAP, where the σ54 CBD binds to, is occupied by the main DNA binding domain of σ70 in the σ70-RNAP holoenzyme. These differences explain a wealth of biochemical and functional data. Pervasively, we find many of these regulatory strategies σ54 uses to control RNAP are shared by a range of other regulatory factors, from the bacterial σ70 to eukaryotic TFIIB and components in RNA Polymerase I. Our structures thus uncover regulatory hotspots within the evolutionally conserved RNAPs that are targeted by a wide range of regulatory elements and provide mechanistic insights into how RNAP is controlled in general.
Our structure provides a molecular basis for how RNAP is tightly inhibited by σ54 and reveals a number of novel regulatory strategies, some of which are conserved in the holoenzymes from other systems. Our findings form foundations for developing antibiotics as a number of the regulatory sites are not currently targeted. Furthermore, σ54 is widely used for controlled expression of genes important for bacterial pathogenicity, so understanding how they are controlled can provide means to control pathogenicity, which could well provide alternative avenues to counter antibiotic resistance.
Mechanistically, we are currently focusing on elucidating how the inhibition by σ54 is released by its cognate activator proteins, which belong to the AAA ATPase family, consisting of a large family of proteins that utilise ATP binding and hydrolysis to carry out a diverse array of functions. How AAA proteins interact and act upon its substrates is still poorly understood and our studies should thus not just provide mechanistic insights into transcription initiation/activation, but also the mechanisms of action of AAA proteins in general.
References:
Funding acknowledgements:
The majority of this work was funded by BBSRC project support to XZ and MB. VC is funded on a Wellcome Trust Investigator Award to XZ while RG is funded by a BBSRC DTP studentship.
Corresponding authors:
Robert Glyde, Dr Vidya Darbari, Professor Xiaodong Zhang, Imperial College London, [email protected].
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.