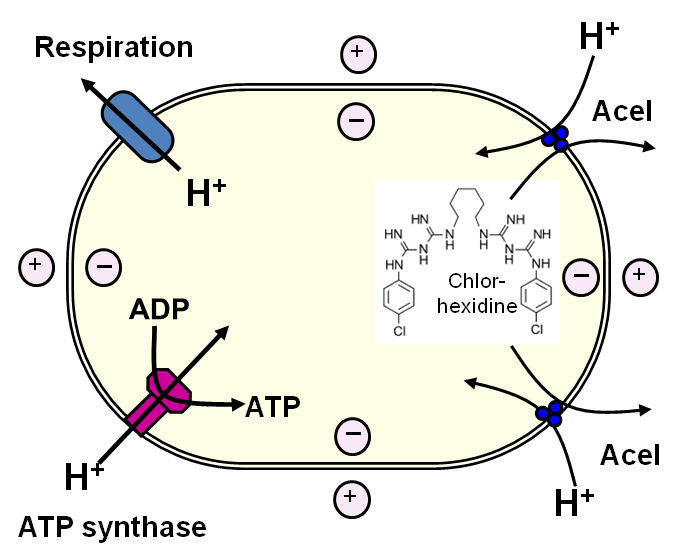
Research performed at Diamond Light Source has helped to understand how a bacterium responsible for many hospital-acquired infections is able to resist one of the most widely used antiseptic agents. By using ultraviolet light to perform circular dichroism measurements of the proteins within the cells, the scientists were able to characterise a previously unknown method by which the antiseptic could be actively transported out of the cell.
 The work involved contributors from the Department of Chemistry and Biomolecular Sciences, Macquarie University, Australia; the Astbury Centre for Structural Molecular Biology and School of Biomedical Sciences, University of Leeds, United Kingdom; and the School of Biological Sciences, Flinders University, Australia.
The work involved contributors from the Department of Chemistry and Biomolecular Sciences, Macquarie University, Australia; the Astbury Centre for Structural Molecular Biology and School of Biomedical Sciences, University of Leeds, United Kingdom; and the School of Biological Sciences, Flinders University, Australia.Hassan, K. A., Jackson, S. M., Penesyan, Patching, S. G., Tetu, S. G., Eijkelkamp, B. A., Brown, M. H., Henderson, P. J. F., Paulsen, I. T. Transcriptomic and biochemical analyses identify a family of chlorhexidine efflux proteins. PNAS 110. Issue 50, 20254–20259 (2013), doi:10.1073/pnas.1317052110
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.