Dr Eugen Stulz, University of Southampton
The structure determination of biomolecules, in particular of DNA and of proteins, is of utmost importance to understand their function in chemistry and biology. In recent years, DNA has become attractive as template for the creation of new materials in nano-biotechnology. The availability of synthetic DNA from automated synthesis has allowed using the sequence specific recognition of the complementary strand, together with a clever design of the DNA sequence, to form two and three dimensional arrays such as grids, lattices, cubes and tetrahedra. Recent developments in the formation of DNA origami structures now allows these assemblies to be used as platforms for further functionalisation, for example to assemble proteins in very specific arrays. Because the use of standard spectroscopy, such as UVvis absorption or NMR, has limited application to analyse these materials, circular dichroism (CD) spectroscopy plays a vital role to determine structure and stability of the constructs. The beamline B23 with its excellent state-of-the-art CD spectrometer enables the detailed analysis of these materials, which are based on both natural and synthetically modified DNA, on a level hitherto inaccessible. Here we will describe some of our experiments in analysing specific DNA strands using the VUV region, the study of synthetic DNA which has been highly modified with chromophores (porphyrins), and of the determination of secondary DNA structures such as G-quadruplexes, where the experiments have provided invaluable data.
One of the most influential discoveries of the past century is arguably the determination of the structure of double-stranded DNA (dsDNA) by Watson, Crick, Wilkins and Franklin.[1] Since then, the synthesis of DNA has reached a level of sophistication where organic chemists can synthesise almost any modified nucleotide and incorporate it, site-specifically, into oligodeoxynucleotides (ODNs). The well-known structure of dsDNA (i.e. distinctive helicity, interior base-stacking region, and major and minor grooves) allows for a reasonable prediction of the structure of modified DNA. The reliability with which a specific DNA sequence recognises its complementary sequence has been used in the past ten years or so to create new nanometrescale two- and three-dimensional objects such as grids and lattices on surfaces,[2] nanoscale patterns through folded DNA,[3] bipyramids,[4] cubes and cages,[5] based on native or modified DNA strands.[6] Two-dimensional DNA structures on surfaces can be used to further position proteins or nanoparticles in order to form grids that may be of use in diagnostics.[2a, 7] The utility of novel DNA architectures in supramolecular chemistry, biology and nano-technology clearly demonstrates the versatility of this approach.[8]
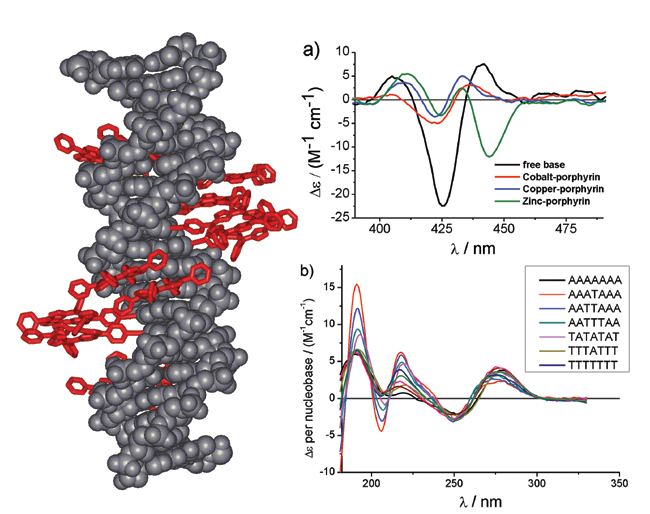 |
| Figure 1: Structure of a porphyrin modified DNA, where the porphyrin-nucleoside has been attached to the complementary strands of DNA to give a zipper porphyrin array. The CD spectra show the differences obtained from various metallated porphyrins. |
The combination of DNA modification with a programmed architecture will certainly be one of the future strengths of this approach. We are using DNA as a supramolecular scaffold to create novel functional molecules on the nanometre scale. In particular, multiporphyrin arrays have been synthesised which are promising candidates for electronic wires, light harvesting antenna systems in artificial photosynthesis and for opto-electronic applications. The attachment of up to 11 porphyrins in a row represents the highest amount of a large hydrophobic metal complex that was incorporated into DNA. The CD spectra show that the constructs retain a B-type DNA structure which is reasonably close to the natural DNA strand. However, depending on the nature of the modification and the amount of substitution some deviations can be observed. By attaching the porphyrins onto complementary strands to form a zipper type array, where the porphyrins form a stacked aggregate upon DNA hybridisation, does retain the natural B-DNA structure better. Of particular interest is the excitonic coupling between the chromophores which become chiral when attached to the nucleosides. The CD signatures in the visible range are very characteristic to the specific type of array. Porphyrins are also excellent ligands for a variety of metals, and we have inserted zinc, copper and cobalt into the porphyrin-DNA strands. In some circumstances the metallation cannot be discerned very clearly by absorption spectroscopy. Here CD spectroscopy proves again to be a very versatile analytical method as the different metallo porphyrin-DNA arrays give rise to very distinctive CD signals (Figure 1a).
Hardly anything is known about the CD signatures of natural DNA strands in the VUV region below 220 nm, which is the cut-off limit of standard CD spectrometers. With the instrumentation available uniquely at B23, we have observed that the different nucleobases A and T give rise to very different signatures in the region of 180 to 220 nm, whereas in the standard region of 230 to 300 nm there is not much difference visible. A systematic approach is being undertaken, and we have varied the amount of A and T bases within a specific DNA sequence, but much more data will have to be acquired before a final model can be created. It is not quite clear at his point whether the different spectra are purely due to the different chromophoric properties of the nucleobases, and to which extent the neighbouring effect in dipole and excitonic interactions have to be taken into account, together with structural preferences of the DNA. However, a very important result is that we did not observe any DNA degradation upon prolonged exposure of the samples to the high energy light, therefore the DNA can be analysed at the beamline. Figure 1b shows representative spectra that were obtained using B23.
Secondary structures in DNA such as quadruples and i-motifs play an important role in gene regulation and genome maintenance; this is highlighted in last year’s Nobel prize in medicine regarding telomeres and protection of genomic material. These structures can also be used in DNA nanotechnology as reversible switching units, in particular G-quadruplexes. We have for the first time covalently attached porphyrins to the telomere sequence; the porphyrins are well known to stabilise these structures. Using CD spectroscopy we could observe that attaching porphyrins at specific positions within the human telomere sequence stabilises two distinctively different structures of the quartet, in particular a parallel and an anti-parallel structure. These could play important roles in biology and are now selected for further crystallographic and biological studies.
References
[1] J. D. Watson, F. H. C. Crick, Nature, 171, 737; b)R. E. Franklin, R. G. Gosling, Nature, 171, 740; c)M. H. F. Wilkins, A. R. Stokes, H. R. Wilson, Nature, 171, 738; (1953).
[2] a)S. H. Park, C. Pistol, S. J. Ahn, J. H. Reif, A. R. Lebeck, C. Dwyer, T. H. LaBean, Angewandte Chemie-International Edition, 45, 735, (2006); b)H. Yan, S. H. Park, G. Finkelstein, J. H. Reif, T. H. LaBean, Science, 301, 1882 (2003).
[3] P. W. K. Rothemund, Nature, 440, 297, (2006).
[4] C. M. Erben, R. P. Goodman, A. J. Turberfield, Journal of the American Chemical Society, 129, 6992 (2007).
[5] N. C. Seeman, Nature, 421, 427, (2003).
[6] a)M. A. Batalia, E. Protozanova, R. B. MacGregor, Jr., D. A. Erie, Nano Letters, 2, 269, (2002); b)P. K. Lo, P. Karam, F. A. Aldaye, C. K. McLaughlin, G. D. Hamblin, G. Cosa, H. F. Sleiman, Nature Chemistry 2, 319, (2010); c)J. Zimmermann, M. R. J. Cebulla, S. Monninghoff, G. von Kiedrowski, Angewandte Chemie-International Edition, 47, 3626, (2008). [7] Y. G. Ke, S. Lindsay, Y. Chang, Y. Liu, H. Yan, Science, 319, 180, (2008).
[8] a)L. H. Eckardt, K. Naumann, W. Matthias Pankau, M. Rein, M. Schweitzer, N. Windhab, G. von Kiedrowski, Nature, 420, 286, (2002); b)L. Jaeger, A. Chworos, Current Opinion in Structural Biology,16, 531, (2006).
Principal publications and authors
L. A. Fendt, I. Bouamaied, S. Thöni, N. Amiot, E. Stulz, Journal of the American Chemical Society 2007, 129, 15319; bI. Bouamaied, T. Nguyen, T. Rühl, E. Stulz, Organic & Biomolecular Chemistry, 6, 3888, (2008). T. Nguyen, A. Brewer, E. Stulz, Angew. Chem. Int. Ed., 48, 1974, (2009).
Funding acknowledgement
Engineering and Physical Sciences Research Council and Biotechnology and Biological Sciences Research Council.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.