Prof William Hunter, University of Dundee
Nicotinamidase catalyzes hydrolysis of nicotinamide to nicotinic acid and ammonia, in the NAD(+) salvage pathway. This enzyme activity has a fortuitous medical benefit since in Mycobacterium tuberculosis it converts the prodrug pyrazinamide nto the bacteriostatic pyrazinoic acid. Pyrazinoic acid is active against semidormant tubercle bacilli and together with rifampicin and isoniazid, forms the front line tuberculosis treatment. Studies of the enzyme have revealed aspects of structure and biochemical activity but there were no structural data on how the enzyme binds and processes physiological ligands. High-resolution crystal structures of the enzyme from Acinetobacter baumanii complexed with nicotinic acid and pyrazinoic acid were therefore obtained to provide direct evidence for the interactions that govern specificity for substrate and prodrug, to inform on the enzyme mechanism and to explain how a valued antibacterial agent is activated. The structures revealed that substrate recognition involves interaction between Zn2+, the enzyme is dependent on divalent transition metal cations, and the pyridyl nitrogen and the position of functional groups allowed us to propose a new mechanism for nicotinamidase activity and one which likely applies to the activation of the important anti-tuberculosis drug pyrazinamide.
Two crystal forms were obtained with nicotinic and pyrazinoic acid respectively and the structures determined at 1.65 and 1.70 Å resolution. The structures and the interactions formed by ligands within the active sites are essentially identical. PncA is a dimeric, divalent cation-dependent enzyme and activity has been reported with Fe2+, Mn2+, and Zn2+. Inductively coupled plasma-atomic emission spectrometry (ICP-OES) identified that recombinant AbPncA contained Fe2+ and Zn2+ in an approximate 1:1 ratio with a trace of Mn2+ present. However, anomalous dispersion diffraction measurements are consistent with a higher occupancy of Zn2+ at the active site.
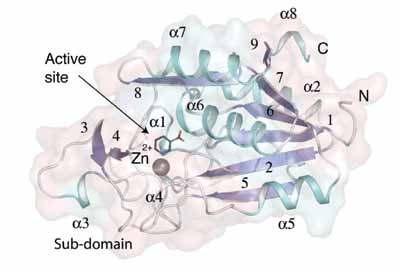 |
| Figure 1. The fold of the AbPncA subunit. β-strands are numbered. |
The enzyme subunit structure is constructed from eight α-helices and nine β-strands (Figure 1). The core of the subunit is a parallel β-sheet of seven strands. Three helices lie on one side of the sheet, with one placed against the other. A sub-domain is placed at one end of the β-sheet. The active site is between the core and the sub-domain (Figure 1), is buried and occluded from solvent by several polypeptide segments. Hydrophobic residues surround the active site and a gross conformational change is required to allow for substrate binding or product release.
The active site Cys159 is located on one side of the active site, which is relatively polar and the Zn2+ positioned on the other. Nicotinic acid binds the cation directly and is positioned between five hydrophobic residues to hold it in place. A further six hydrophobic residues stabilise the nonpolar environment around the nicotinic acid and occlude the active site.
Amidation can involve acidic or basic hydrolysis and the structures of the enzyme:product complex suggest that the latter applies in PncA. Acid hydrolysis would involve nucleophilic water attacking the carbon of a protonated amide. The hydrophobic environment on one side of the amide and close interactions with functional groups on the other renders it difficult to see how water could be positioned to attack C7.
The strictly conserved and essential Cys159, on the polar side of the active site, is positioned to attack the carbonyl carbon and together with the position of the ligands leads us to propose a four-stage mechanism (Figure 2). Substrate binds in the axial position displacing a Zn2+ coordinating water. The equatorial metal ion coordinating waters are also held in position by hydrogen bonding interactions to the enzyme whereas the axial water lacks that restraint. It is the axial water that likely vacates the coordination sphere as substrate binds. This water may be prevented from exiting the active site due to the presence of the hydrophobic lid (the sub domain) covering the active site. The displaced water, or an incoming water would preferentially bind to the hydrophilic side of the active site. Proton abstraction by acidic residue, Asp16, would help to generate a Cys159 thiolate facilitating nucleophilic attack at C7. Two main chain amides create an oxyanion hole to support thiolate attack by stabilizing the tetrahedral intermediate formed during catalysis in a similar fashion to cysteine proteases. Proton donation from Asp16 to substrate would promote C-N bond cleavage and release of NH3 as the tetrahedral-intermediate collapses in stage II; being converted to an acyl-intermediate. Water activation by Asp16, where the amino acid is protonated, generates a nucleophilic hydroxyl to attack the acyl-intermediate producing nicotinic acid (or pyrazinoic acid) and a thiolate in stage III. In stage IV, products are released and the thiolate accepts a proton from Asp16 to regenerate a thiol.
The activation of water to hydroxyl groups by a strong Lewis acid, for example four-coordinate Zn2+ is a commonly invoked feature of zinc-dependent enzymes. In PncA, the metal ion is six-coordinate and this would serve to reduce the Lewis acid strength. In addition, that PncA catalysis is supported by different divalent cations suggests that Lewis acid strength is less important than the structural role provided by octahedral metal ion coordination, precise placement of the substrate and an appropriate ligand exchange rate to support substrate binding with release of a water molecule. Our mechanistic proposal therefore has no requirement for the metal ion to activate water.
 |
|
|
In summary, high-resolution crystal structures determined using synchrotron radiation have revealed how nicotinic acid,and pyrazinoic acid bind AbPncA. The position of functional groups then allowed us to propose a new mechanism for nicotinamidase activity and one that likely applies to the activation of the important anti-tuberculosis drug pyrazinamide.
Principal Publication and Authors
Fyfe PK, Rao VA, Zemla A, Cameron S, Hunter WN. Specificity and mechanism of Acinetobacter baumanii nicotinamidase: implications for activation of the front-line tuberculosis drug pyrazinamide. Angew Chem Int Ed Engl. 48 (48):9176-9 (2009).
Funding Acknowledgement
Biotechnology and Biological Sciences Research Council [BBS/B/14434], The Wellcome Trust [082596 and 083481] and European Commission Seventh Framework Programme (Aeropath; FP7/2007-2013).
Research carried out at Diamond on I04 and at ESRF on ID29.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.