Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()
Cryo-EM requires protein samples to be frozen before they are imaged using an electron microscope. This revolutionary technique and its application to structural biology was the focus of the Nobel Prize for Chemistry in 2017 and has contributed to a wave of new structural information for proteins that were difficult or impossible to prepare for X-ray crystallography. As cryo-EM became more popular, new methods and tools arose to improve the technique and make it more efficient. Now, many people are working on the way that samples are prepared so that this is easier to get high resolution structural information from each experiment. Once samples are prepared, there are a number of automated tools that have been developed for picking the frozen particles to be processed. It was while investigating these tools that PhD student Mateusz Olek and his supervisor Peijun Zhang with help from Yuriy Chaban and Donovan Webb, uncovered an unusual problem. Their findings have been published in the journal Structure.
The automated particle pickers are designed to analyse an image and automatically choose the best particles to select for experiments. While studying the performance of these tools, the team noticed that there were portions of the image with no selections. These voids were not easily explained, especially when the team visually inspected the images and saw clearly that particles were present. For some reason, the automated particle pickers were blind to certain regions of the image. This problem could have repercussions for cryo-EM experiments. If viable particles are being left behind by automation software, scientists may not be able to collect all the data that they need for an experiment.
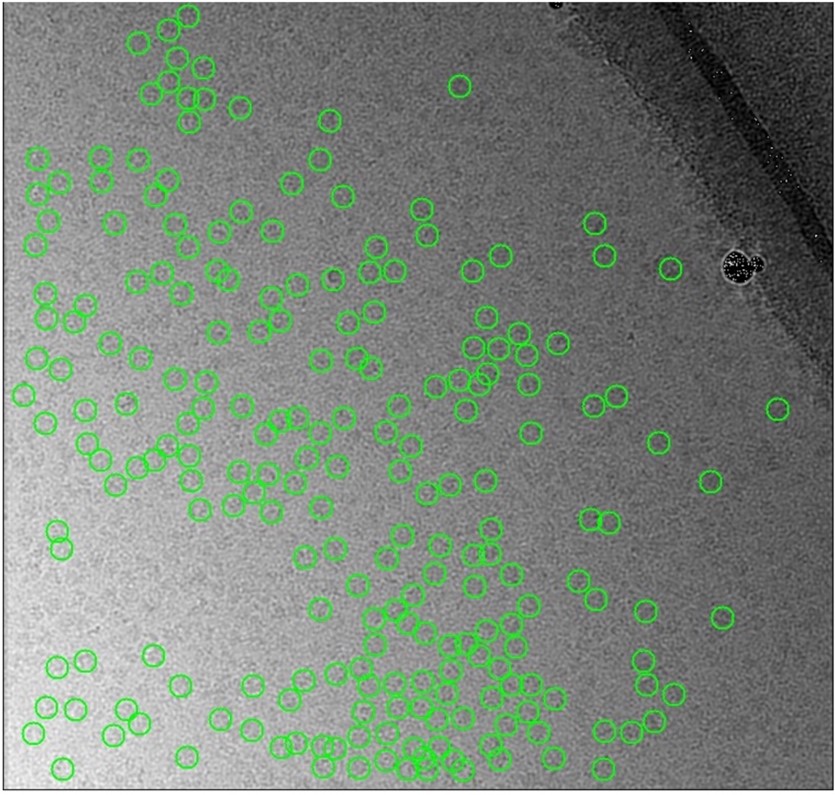

The team quickly noticed that the background of the images was not consistent. Some portions were darker and others were lighter. This could account for some of the failings in automation software which rely on measuring the contrast of a protein particle against the background. To do cryo-EM, protein particles are suspended in a thin film of ice, so the team concluded that inconsistencies in the background of the image were related to different thicknesses of the ice. This raised several problems for researchers using cryo-EM. Firstly, one obvious solution would be to try to make the ice film more uniform. Unfortunately, while great efforts are being made to improve the preparation of cryo-EM samples, it is still very difficult to generate a uniform film of ice. Often a gradient of ice thickness can be seen in a cryo-EM sample.
Mateusz and the team started from scratch to develop a new method to combat the ice problem. They began by segmenting different images and analysing the background. This allowed their picker to identify particles irrespective of the background coming from the ice. This innovation meant that in a given experiment, researchers could now reliably gather more structural information by analysing more protein particles. However, this was not the end of the road. While collecting more particles is extremely valuable for researchers using cryo-EM, the amount of ice affects the quality of the particles, and therefore the quality of cryo-EM maps that can be reconstructed. Mateusz and the research team knew that they could take the software further, by giving information instantaneously to the researchers about the quality of the protein particles that were being imaged.
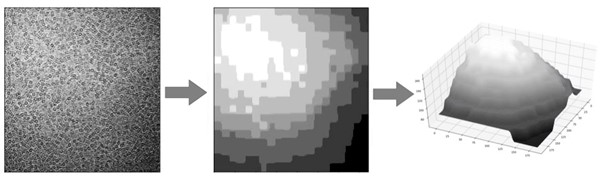
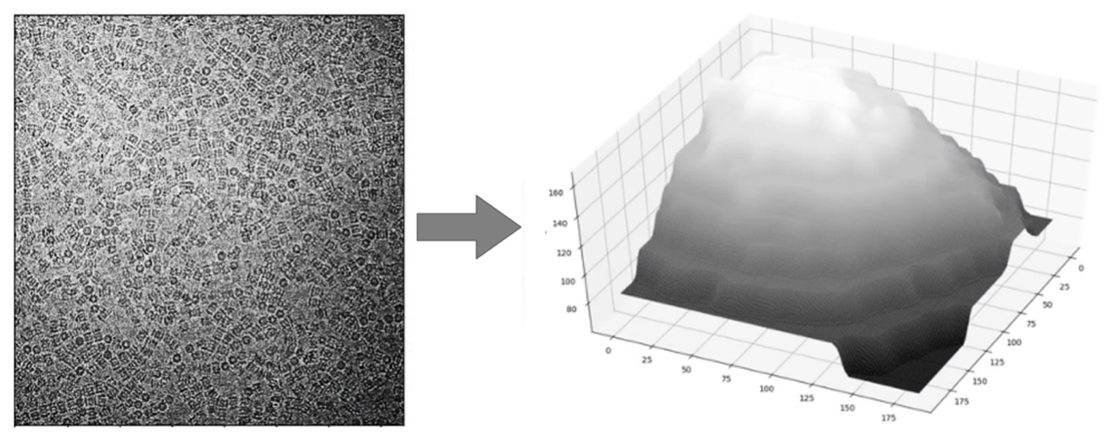

In their recent publication, the research team highlighted two major problems that can arise when ice is not the optimal thickness. Firstly, very thick ice creates more background noise in the image. This makes it difficult to get high resolution data from a protein particle. Conversely, if the ice film is too thin, it may not support the proteins properly. This problem is very much dependent on the specific proteins that a researcher is investigating. For example, some proteins are small and tightly bound in a spherical configuration. These proteins can be supported in relatively thin layers of ice. However, many proteins are large with long projecting branches that can fall outside of the ice film if it is too thin. This means that thin ice makes it difficult to image these kinds of proteins, even if the background noise is very low. The optimal ice thickness is something that minimises the background but is thick enough to fully support the protein. Often the optimal ice thickness will depend on the nature of the protein being studied and will change depending on the experiment.
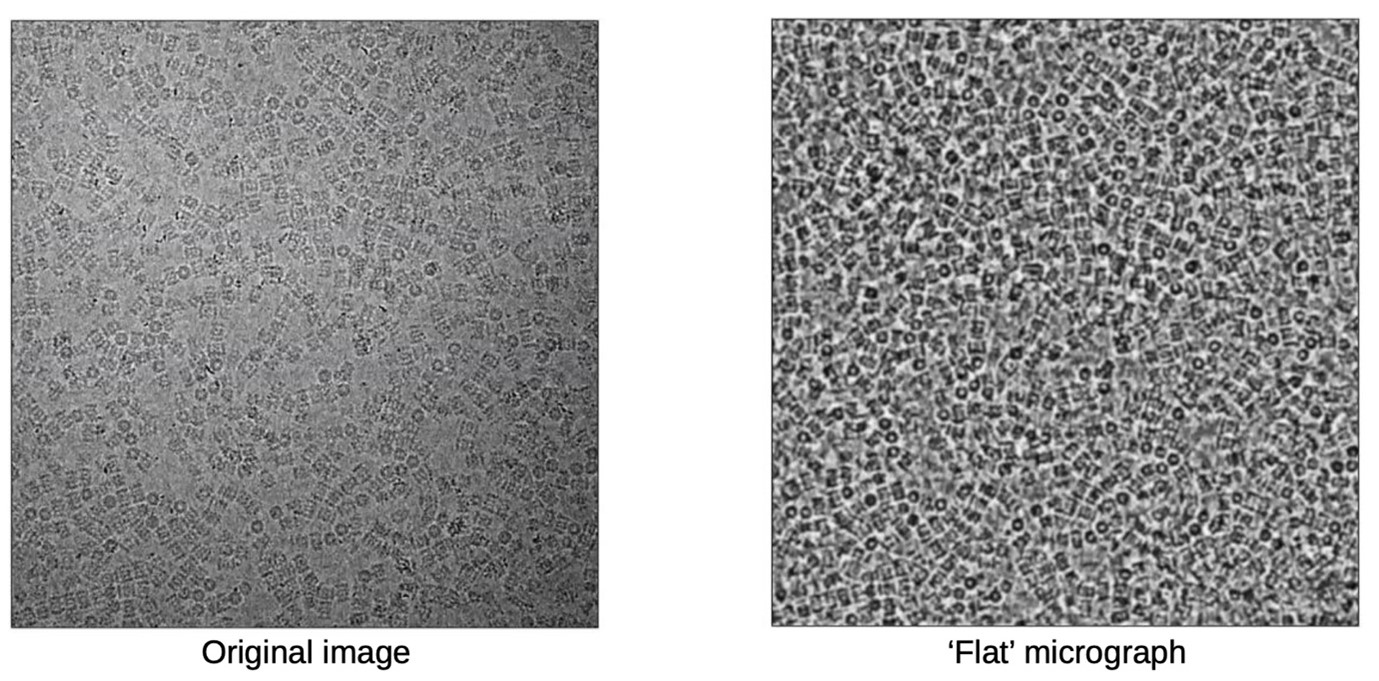

Their new software tool, IceBreaker, could conceivably completely automate the selection of particles. However, the team opted for an approach that gives more flexibility to the researchers using it. Instead of making decisions, the software annotates each particle giving the researcher an indication of the quality. This allows them to steer their experiments in precisely the way they need to achieve optimal results. Sometimes fewer, high-quality particles are enough and sometimes more particles of poorer quality from thicker ice are needed to show unique particle orientations not supported by the thin ice. Using the new IceBreaker software, the researchers are in full control of the data that they are collecting. IcreBreaker is now being implemented as part of the data collection pipeline at Diamond’s electron Bio-Imaging Centre (eBIC) and is freely available on GitHub.
Olek M. et al. IceBreaker: Software for High Resolution Single Particle CryoEM with Nonuniform Ice. Structure (2022). DOI: https://doi.org/10.1016/j.str.2022.01.005
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.