Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()

For the first time, an international team of researchers has visualised the light-sensitive protein archaerhodopsin-3 (AR3) at unprecedented resolution using the I24 and B23 beamlines and supported by the Membrane Protein Laboratory (MPL) at Diamond. The photoreceptor is expressed by Halorubrum sodomense, an organism that grows in the Dead Sea, but is best known for its applications in optogenetics experiments, in which it is used to silence individual neurons and to detect changes in cell membrane voltage. These new structures open the way for the development of new tools and methodologies in the fields of neuroscience, cell biology and beyond.
In their paper, published this week in Nature Communications, the team reports the first ever structure of the ground state of AR3. In this state, the protein is configured to transport one H+ ion across the cell membrane for each photon absorbed. The team were also able to crystallise the photoreceptor in a second conformation, a desensitized state that is adopted by AR3 in the prolonged absence of light.
Corresponding author, Professor Anthony Watts, from the University of Oxford, said:
The superb resolution that we have achieved for these AR3 structures, 1.07 Å for the ground state, is among the highest for a wild-type membrane protein deposited to date in the Protein Data Bank. This quality allows us to visualise directly the complex distribution of water molecules inside the receptor and describe the functional significance of the intricate networks of hydrogen bonds that they form, something that is of importance in many biomolecules – not just photoreceptors. These networks in AR3 involve interactions with the retinal group and amino acid side chains and, in this paper, we’ve shown how they are altered between the ground and desensitised states of the protein.
Explaining the significance of the AR3 protein, joint-first author Dr Juan Francisco Bada Juarez, said:
The exciting developments in optogenetics over the past decade have led to intense interest in AR3 from scientists all around the world. Mutants of the protein are routinely used in neuroscience experiments to selectively silence individual nerve cells and also to detect changes in transmembrane voltage. However, these mutants have been designed without knowledge of the structure of the protein itself. Our data provides structural biologists and protein engineers with the ‘blueprints’ to AR3 and, as a result, we can expect a significant acceleration in the development of new optogenetic tools.
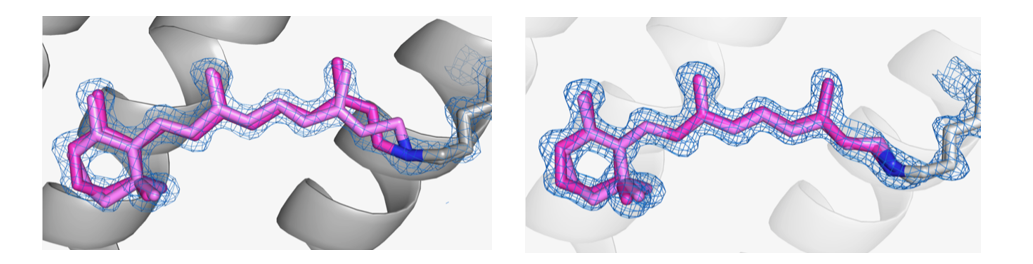%20in%20the%20desensitised%20state%20(top)%20and%20ground%20state%20(bottom)%20of%20AR3..png)
Co-corresponding author Dr Isabel Moraes, from the National Physical Laboratory (NPL), added:
The very high-resolution diffraction data has enabled us to observe in great detail the freedom of motion of several key amino acid and individual water molecules inside the photoreceptor. Solving the AR3 structures was extremely challenging, because of the complexity of the features that we can distinguish at atomic resolution. Many amino acid side chains inside the protein were resolved in more than one position or orientation, indicating their importance for protein function. Knowledge of the dynamics of these groups was key to understanding how the processes of sensitisation and desensitisation take place.
Essential for understanding the workings of the protein, were the high-resolution crystal structures solved by the team. Obtaining such high quality diffraction data would not have been possible without the state-of-the-art microfocus beamline at Diamond. The team worked closely with several experts at Diamond.
Diamond Senior Beamline Scientist, Dr Danny Axford, said:
The microcrystals grown by the Oxford and NPL teams were perfectly suited to the capabilities of the I24 beamline. Using software developed by the DIALS software development group at Diamond, we were able to combine diffraction images from multiple crystals to obtain complete datasets. The combination of precise microfocus X-ray delivery and advanced data analysis enabled us to push the resolution of the resulting structures to the limits.
The CD measurement from Diamond's B23 Beamline allowed the team to quantify the alpha-helical content of the protein (AR3) and correlate with other biophysical information.
In conclusion Prof. Watts added: “This has been a fascinating study. It has been a pleasure to work with so many expert colleagues both in the UK and around the world. Not only do we now have a highly detailed knowledge of the structures of two important functional states of AR3, but the insights gained deepen our understanding of how the processes of sensitisation and desensitisation occur in many other receptor proteins.”
Bada Juarez, J.F., Judge P.J. et al. Structures of the archaerhodopsin-3 transporter reveal that disordering of internal water networks underpins receptor sensitization. Nature Communications (2021) DOI: 10.1038/s41467-020-20596-0
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.