Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()
In 1962 the Nobel Prize in Chemistry was jointly awarded to Max Perutz and John Kendrew for determining the structures of two proteins, haemoglobin and myoglobin using a technique called X-ray diffraction. Since then over 70,000 protein structures have been solved, leading to an increased understanding of processes in the human body, how diseases take hold, and how the impact of diseases can be lessened. Diamond is adding to this vast lexicon of knowledge, with scientists using the facility to find over 500 of these important structures.
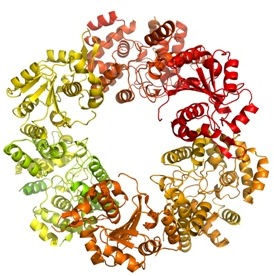
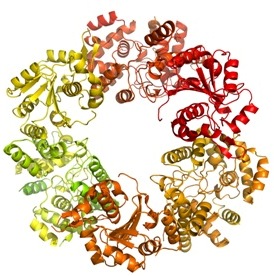 |
| The trimer of the Lassa nucleoprotein, part of the Lassa virus which causes up to 500 000 cases of acute viral haemorrhagic fever and 5000 deaths each year across West Africa. |
Proteins are biological molecules involved in almost every cellular process, determining how the human body and every other organism functions and flourishes, or gets sick and dies. Each protein folds in a unique way, the structure of which provides information on how it works. But finding this structure is a complex process. The protein is produced, crystallised and illuminated by X-rays. The interactions between the X-rays and the crystal form a pattern that can be analysed to deduce the protein structure. The technique is known as crystallography.
In 1971 a group including Max Perutz decided that a repository was needed to store the increasing number of structures solved by groups around the world. In the UK the Cambridge Crystallographic Data Centre (CCDC) was already assembling and archiving structures gathered in this way, and across the pond Brookhaven National Laboratory (BNL) were doing the same thing. By October 1971 the Protein Data Bank was established, jointly managed by CCDC and BNL. It was to become the single place where all protein structures were deposited.
“This was really significant. The people that founded the technique were instrumental in setting up the Protein Database, and they did this right at the beginning before the community had time to adopt different standards for how to report protein structures. We are all benefiting from this today.”Diamond scientist and protein crystallographer Thomas Sorensen
Now there are over 70,000 structures in the database. Since the 1980s synchrotrons have played a vital role in providing intense sources of X-rays that enable scientists to solve increasingly complex structures with increasing efficiency. Over 45,000 structures have been solved by the worldwide synchrotron community. Many of these structures were chosen because of the role they play in the development and prevention of disease, to enable the development of new drugs which neutralise viral proteins or prevent them taking hold in the body. Drugs including Herceptin and Tamiflu relied on synchrotron studies as part of the development process.
 Listen to Diamond users discuss their research in the structural biology podcast
Listen to Diamond users discuss their research in the structural biology podcast
Thomas Sorensen says, "We are very pleased to reach 500 protein structures but this is really just the beginning for Diamond. Two new dedicated crystallography beamlines have come online since the facility opened and design/construction for an additional beamline is well underway with first users scheduled for Autumn 2013. Adding in our continued development of existing beamlines, we are providing a facility that enables our users to solve increasingly complex structures."
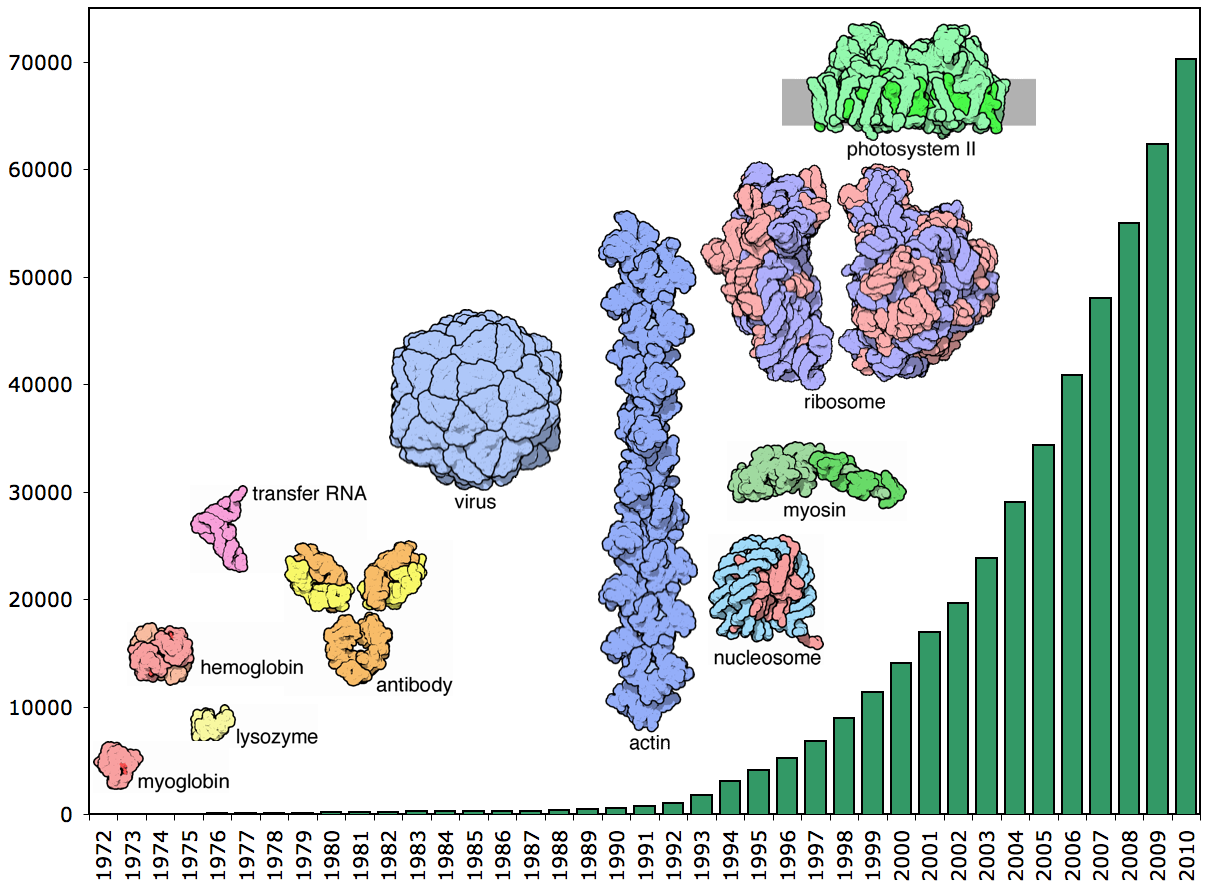
Number of structures in the PDB from 1972 - 2010. Image courtesy of the RCSB Protein Data Bank.
 Understanding retroviral DNA integration
Understanding retroviral DNA integration
Understanding how viruses such as HIV function is vital in the search for more effective antiviral medicines, as well as opening up novel possibilities in gene therapy. A collaborative study involving Diamond’s Macromolecular Crystallography beamlines revealed important mechanistic aspects of retroviral replication. More...
Improving antibiotics
%20sm.png) In general, the antibiotics that are prescribed by your GP work. Complete the prescribed course and your symptoms have gone. But in some cases, the bacteria causing the illness have developed a resistance to the drugs. We need to find a way to improve these drugs so that they can overcome this resistance. But we need to know how our drugs work before we can improve them, and one way to do this is by producing a 3D molecular image of the drug/inhibitor in action. Knowing the 3D structure of an antibiotic bound to its target protein elucidates the molecular mechanism – revealing how the drug works and how it could be modified to overcome resistance. More...
In general, the antibiotics that are prescribed by your GP work. Complete the prescribed course and your symptoms have gone. But in some cases, the bacteria causing the illness have developed a resistance to the drugs. We need to find a way to improve these drugs so that they can overcome this resistance. But we need to know how our drugs work before we can improve them, and one way to do this is by producing a 3D molecular image of the drug/inhibitor in action. Knowing the 3D structure of an antibiotic bound to its target protein elucidates the molecular mechanism – revealing how the drug works and how it could be modified to overcome resistance. More...
More effective chemotherapy
Cancer comes in many forms, but it is characterised by cells rapidly dividing out of control. Chemotherapy refers to a group of drugs which use a range of mechanisms to try and block the cells from dividing. A group of scientists from Cancer Research UK have been using Diamond to study the structure of a protein called FANCL which is known to play a role in initiating DNA repair. Understanding how this protein functions could help make some forms of chemotherapy more effective. More...
The cause of Lassa fever
 The Lassa virus remains invisible to the human defence system and causes thousands of deaths each year. Treatment is currently limited and there is no effective vaccine In addition, the virus is a biological threat agent, as Lassa fever could transmit from person to person. It had been known that the nucleoprotein (NP) of Lassa virus has an essential role in immune suppression in sufferers but how this protein worked on a molecular level was still a mystery. More...
The Lassa virus remains invisible to the human defence system and causes thousands of deaths each year. Treatment is currently limited and there is no effective vaccine In addition, the virus is a biological threat agent, as Lassa fever could transmit from person to person. It had been known that the nucleoprotein (NP) of Lassa virus has an essential role in immune suppression in sufferers but how this protein worked on a molecular level was still a mystery. More...
View the MX Structure Galleries I02 | I03 | I04 | I24 | I04-1
![]() Follow the MX Beamlines on Twitter
Follow the MX Beamlines on Twitter
Background on the PDB from The Protein Data Bank: a historical perspective, Helen M Berman, Acta Cryst (2008). A64 88-95. DOI: 10.1107/S0108767307035623
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.