Keep up to date with the latest research and developments from Diamond. Sign up for news on our scientific output, facility updates and plans for the future.
Related publication: Abascal-Palacios G., Ramsay E. P., Beuron F., Morris E. & Vannini A. Structural basis of RNA polymerase III transcription initiation. Nature 553, 301 (2018). DOI: 1038/nature25441
RNA polymerase (Pol) III is a major determinant of lifespan in eukaryotes (organisms whose cells have a nucleus enclosed within membranes, unlike bacteria). The level of Pol III transcription is tightly linked to the rate of growth, as it is known to play a role in cancer, and neurodegenerative diseases. A better understanding of the structure of Pol III could lead to new therapies for these diseases. However, Pol III is the most complex nuclear RNA polymerase.
The eukaryotic nuclear genome is transcribed by the multisubunit enzymes RNA Polymerase (Pol) I, II and III, which catalyze DNA-dependent RNA synthesis. Pol III transcribes genes encoding short non-coding essential RNAs, such as the entire pool of transfer RNAs (tRNAs), the 5S ribosomal RNA (rRNA) and the spliceosomal U6 small nuclear RNA (snRNA). Pol III genes are essential in all cells and involved in fundamental processes such as ribosome and protein biogenesis, RNA processing, and protein transport. Pol III transcription is tightly co-regulated with Pol I activity, accounting together for up to 80% of nuclear gene transcription in growing cells1. Pol III activity influences TORC1- dependent lifespan in yeast, flies and worms and, given its conservation, it is likely to exert the same effect in vertebrates2. Pol III deregulation has been associated with a series of neurogenerative diseases and with cancer onset3.
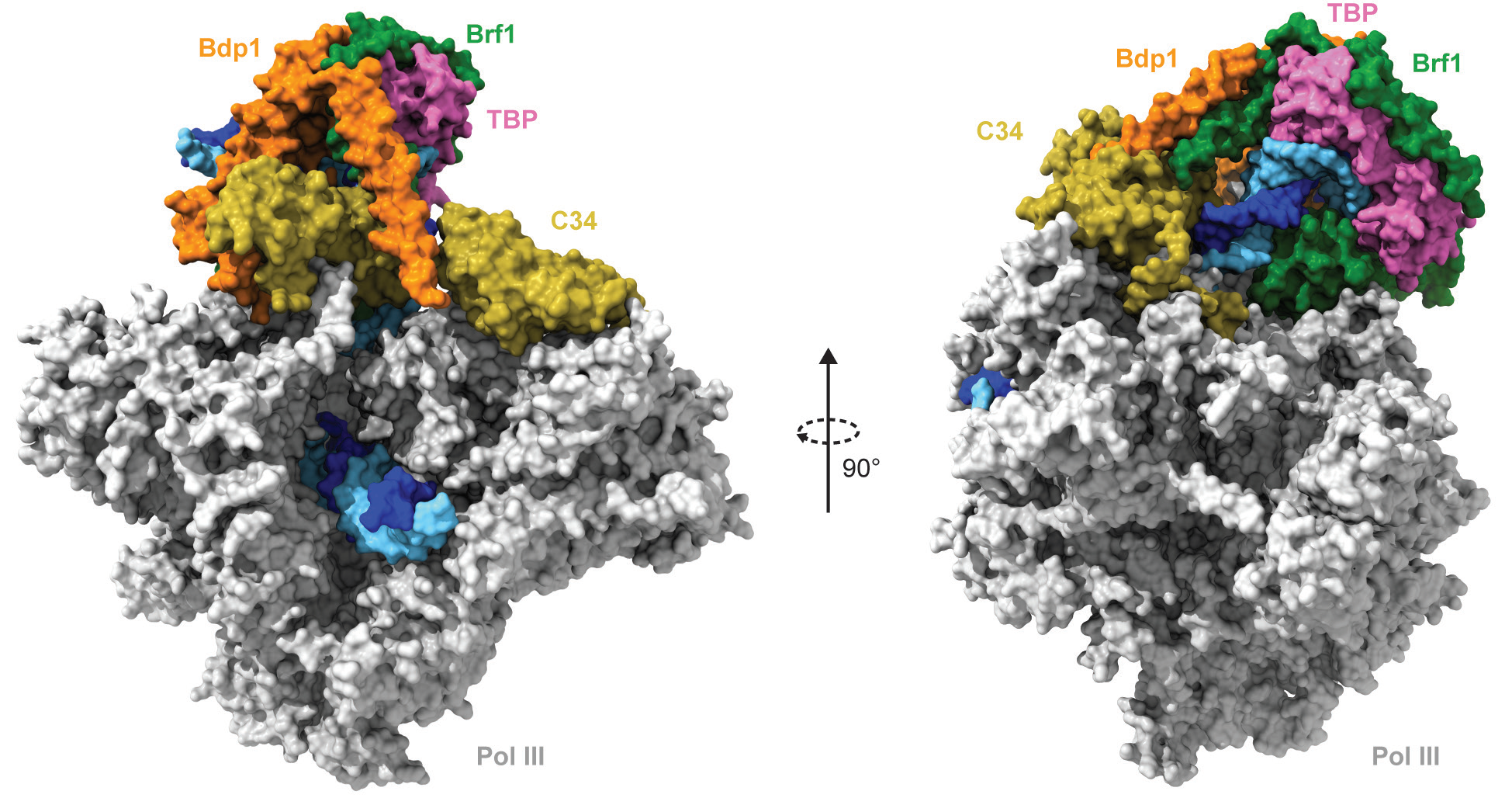
Pol III, with its 17 subunits and nearly 700 kDa mass, is the most complex nuclear RNA polymerase. Recruitment of Pol III at its target promoters rely on a specific subset of transcription factors, including TFIIIA, TFIIIB and TFIIIC. TFIIIA and TFIIIC complexes can be regarded as ‘peripheral’ recruiting factors for the core transcription factor TFIIIB. At all Pol III-transcribed genes, TFIIIB ultimately recruits Pol III resulting in the formation of a closed pre-initiation complex (PIC) and subsequently, assists Pol III in melting the DNA and inserting the template strand at the active site, which is the transition to an open PIC1,3. While Pol I and Pol II requires additional transcription factors, Pol III only requires TFIIIB in order to drive specific transcription from a TATA-box containing U6 promoter in vitro and does not require the ATP-dependent helicase activity of TFIIH for the closed- to open-PIC transition in vivo, as in the case of Pol II1,3.
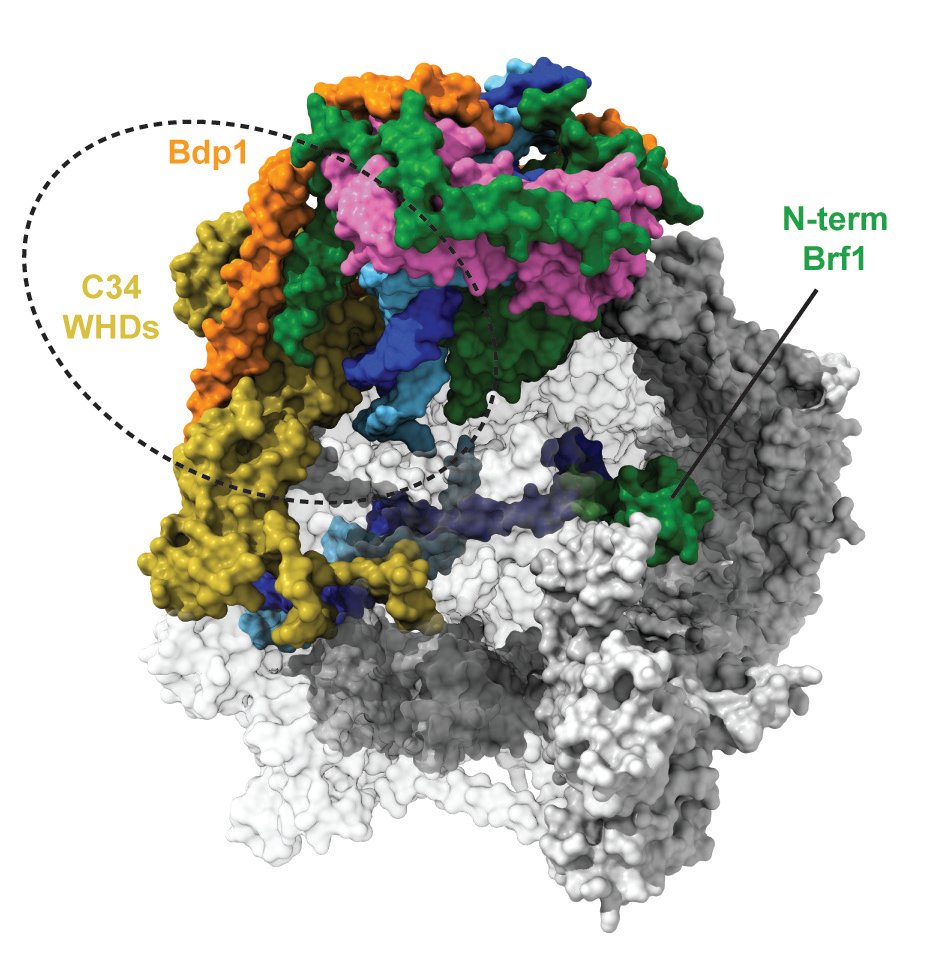
Cryo-EM analysis of a core Pol III PIC, reconstituted in vitro using endogenous S.cerevisiae Pol III, recombinant TFIIIB and a 70 base-pairs double-stranded DNA scaffold encompassing the yeast U6 snRNA promoter, resulted in reconstructions of Pol III PIC in different states in the 3.4-4.0 Å resolution range. In particular, an atomic model of a core Pol III PIC in its open state, where the DNA has been spontaneously melted and loaded in the active site of the Pol III enzyme could be built (Fig. 1).
References:
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.