Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()
Tuberculosis (TB) is one of the deadliest infectious diseases worldwide that is spread in the air like the common cold. TB is caused by the bacterium Mycobacterium tuberculosis (Mtb) and was responsible for approximately 1.6 million deaths in 2021. Mtb will usually attack the lungs but can attack any part of the body. Drug resistant TB strains are spreading and present a major concern. New work and a paper by a team of scientists from Manchester, Cambridge and Huddersfield Universities using a structure-guided approach, combined with biophysical characterization obtained a series of compounds with activity against clinically relevant drug-resistant isolates. These will support further development of much-needed additional treatment options against Mtb.
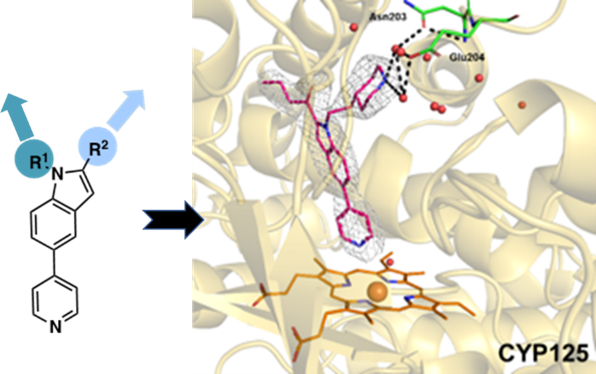
During infection the TB bacteria can utilize lipids (cholesterol and fatty acids) from the human host to act as nutrients to maintain and fuel the infectious state. The teams’ paper, just published in Chemistry Europe, outlines this collaborative project and the results of their research into a set of enzymes involved in a key step in the breakdown of host cholesterol used by the TB bacteria. Called “Structure Based Discovery of Inhibitors of CYP125 and CYP142 from Mycobacterium tuberculosis,” (https://doi.org/10.1002/chem.202203868) ; their work involves characterising the enzymes using the UK’s national synchrotron, Diamond Light Source X-ray macromolecular crystallography beamlines to view their structures in molecular detail.
Corresponding author, Dr Kirsty McLean Senior Lecturer in Cell Biology/Biochemistry in the Department of Biological and Geographical Sciences, University of Huddersfield explained that these enzyme structures are used to design inhibitor molecules that can bind to the enzymes and prevent them doing their job - essential for TB infection. The inhibitors molecules initially came from screening a library of small chemical compounds and were then gradually built up and synthesised chemically designing them to sit specifically into the enzyme structure. This process iteratively combines protein structure knowledge and chemical synthesis to design a succession of better inhibitors. The leading compounds were tested against clinically active and drug resistant strains of Mycobacterium tuberculosis and revealed anti-TB activity.
Dr McLean adds;
Using a structure-guided approach, combined with biophysical characterization, compounds with micromolar range in-cell activity against clinically relevant drug-resistant isolates were obtained. These will support further development of much-needed additional treatment options and provide routes to probe the role of CYP125 and CYP142 in Mtb pathogenesis. This study should incite further development of an alternative range of anti-TB inhibitors
Dave Hall, Science Group Leader for Diamond’s Macromolecular Crystallography Group continues;
Much of the team’s X-ray diffraction data was collected and managed using tools for remote access to our beamlines and new shift modes. Our staff worked closely with the group across various of our beamlines including I03, I04, I04-1 and I24. We are delighted to have helped move this important research on to the next stage.
With the emergence of extensive drug resistance, novel therapeutic agents are urgently needed, and continued drug discovery efforts required. Host-derived lipids such as cholesterol support Mtb growth, and are also suspected to function in immunomodulation, with links to persistence and immune evasion. Mtb cytochrome P450 (CYP) enzymes facilitate key steps in cholesterol catabolism and thus present potential targets for inhibition. The authors present a series of compounds based on an ethyl 5-(pyridin-4-yl)-1H-indole-2-carboxylate pharmacophore which bind strongly to both Mtb cholesterol oxidases CYP125 and CYP142.
Katariya, M. M., McLean, K. J., et. al. Structure Based Discovery of Inhibitors of CYP125 and CYP142 from Mycobacterium tuberculosis. Chem. Eur. J. 2023, e202203868. DOI: https://doi.org/10.1002/chem.202203868
X-ray data collection and structure solution: X-ray diffraction data was collected at various beamlines at Diamond Light Source in Oxfordshire UK, and was indexed and integrated using the DIALS pipeline.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.