A team of scientists from the University of Oxford have successfully mapped the structure of the deadly Ebola virus molecule that drives the attack strategy and leads to fatal infections in humans. Using the powerful X-rays produced at Diamond Light Source, the researchers solved the structure of the molecule in its native state, and also when bound with the anticancer drug toremifene and ibuprofen.
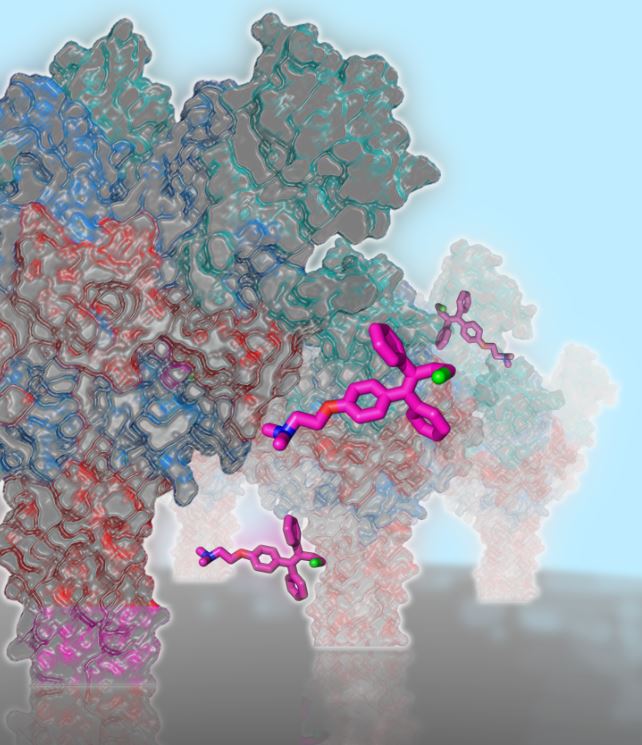
A schematic showing Ebola glycoproteins on the membrane being targeted by Toremifene
Ebola virus disease is a severe, often fatal illness in humans and Ebola viruses (EBOVs) are responsible for repeated outbreaks of fatal infections, including the recent epidemic in West Africa, which resulted in more than 28,000 cases and 11,000 deaths.
Currently, there are no approved therapeutic drugs or vaccines for the disease, however, there is clearly an urgent need to find lead compounds for drug development or repurpose drugs for the treatment of EBOV disease.
EBOV is an enveloped virus where the membrane envelope contains glycoprotein (GP), a molecule that is solely responsible for the three key stages of the viral attack. GP is therefore a primary target for the development of antiviral drugs.
Prof Dave Stuart, MRC Research Professor at the Division of Structural Biology at the University of Oxford and Diamond Light Source, explains, “Exploring the potential of drugs that are already licensed makes sense because they have already been through clinical trials so the timeframe for bringing them to market is usually much quicker. A set of potential inhibitors for EBOV had been identified, including an anticancer drug toremifene and the painkiller ibuprofen, however, their mechanism of action was largely unknown.”
“Using the crystallography beamlines at Diamond, we solved the structure of EBOV GP in its native state and also bound with toremifene and ibuprofen. The native structure gives a more complete and accurate picture of the molecule itself and the conformational changes that take place when an antibody is introduced and receptor binding takes place.”
“Our results, which we have published today in Nature, were unexpected. The binding site is different to that predicted for ibuprofen. We were able to pinpoint the binding site of toremifene and ibuprofen on the surface of the molecule and reveal that the drugs decrease the stability of the viral GP, triggering premature release of one of its subunits and, crucially, preventing viral fusion. Toremifene was shown to lock in more tightly than ibuprofen. This information can guide the design of more powerful anti-EBOV drugs. My group will be continuing this project at Diamond on the new XChem fragment screening facility, which should help us to further accelerate the rational drug design process. We hope this research will lead to new drugs that can help to save lives if we are faced with further outbreaks of the disease in the future.”
Prof Dave Stuart
