Keep up to date with the latest research and developments from Diamond. Sign up for news on our scientific output, facility updates and plans for the future.
Gram-negative bacteria that are extensively resistant to multiple antibiotics are an ever-increasing global threat. Colistin is an antibiotic used as the last line of defence against highly resistant Gram-negative bacteria. However, in recent years bacteria have emerged with resistance to this crucial antibiotic.
Resistance to colistin is determined by an enzyme called ‘mobilised colistin resistance’ (MCR-1), which has rapidly disseminated worldwide and is found in clinical strains of E. coli and Klebsiella pneumoniae. MCR-1 modifies the outer membrane of Gram-negative bacteria, preventing it from interacting with colistin, but despite its huge importance, the mechanism by which this happens remains to be established.
To shed some light on the workings of MCR-1, the crystal structure was solved using the Macromolecular Crystallography (MX) beamlines I04-1, I03 and I02. MCR-1 was found to be a zinc metalloprotein, with an alkaline phosphatase fold containing three disulphide bonds. These high-resolution data provided a detailed view of the active site, identifying areas of the enzyme that were critical for function. Moreover, these results allowed a computational prediction of the mechanism of action, and determined that a single zinc ion may be sufficient for MCR-1 to function. Knowing these important details could enable the rational development of novel antimicrobials to effectively combat colistin-resistant bacteria in the future.
Bacterial diseases, particularly those caused by Gram-negative bacteria (GNB), are a serious and increasing clinical threat worldwide. Treatment of GNB infections is complicated by their intrinsic resistance to many antibiotic classes and ready acquisition of resistance to additional agents. There are now some bacteria for which there are few treatment options remaining, with only a handful of antibiotics that are considered reliable for therapy. The polymyxin colistin is one such key `last-resort’ antibiotic, used to treat infections by multidrug-resistant GNB pathogens. Colistin is a positively-charged polycationic peptide that binds to and disrupts the outer membrane of GNB by interacting with the negatively-charged lipid A headgroup. Until recently, resistance to colistin was considered rare, limited to changes in expression of chromosomal proteins that collectively modify the lipopolysaccharide, the component of GNB outer membranes. However, in November 2015 our team described colistin resistance mediated by MCR-1, a plasmid-encoded phosphoethanolamine (PEA) transferase1. Since this initial report, it has now been reported in over 40 countries, and is found in numerous bacterial strains, mainly E. coli and Klebsiella pneumoniae. In addition, a number of homologues have been further identified, including MCR-22. MCR- 1 has now been reported in bacteria producing other resistance determinants, such as carbapenemases, which can result in essentially untreatable bacterial infections. However, expression of MCR-1 in E. coli comes at significant cost to the host, highly attenuating virulence and resulting in decreased growth rate, cell viability, and degradation of the cell membrane3.
MCR-1 is an integral inner-membrane enzyme that catalyses the transfer of positively-charged PEA onto a phosphate of the glucosamine headgroup of lipid A, which is subsequently incorporated into the outer membrane. This confers resistance to colistin by reducing the net negative charge and preventing colistin from binding to, and subsequently inserting into, the outer membrane. The catalytic mechanism of MCR-1 (and other bacterial PEA-transferases) remains to be established.
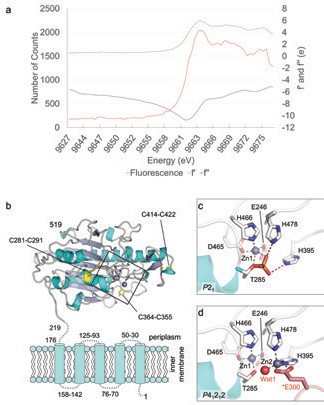
To understand this mechanism, crystal structures of the catalytic domain of MCR-1 were obtained using beamlines I02, I03 and I04-1. To facilitate crystallography the protein was purified lacking the trans-membrane domain, so the soluble, periplasmic domain containing the predicted active site (i.e. the MCR-1 catalytic domain; residues 219 onwards) of the protein was crystallised. A fluorescence scan on beamline I03 of these crystals revealed a strong peak at the Zn-edge, indicating the presence of zinc ions (Fig. 1a). Using this information, the crystallographic phases were determined using four single-wavelength anomalous dispersion (SAD) datasets collected on I03 at 2.50 Å resolution. These phases were used to solve the crystal structure of two higher resolution native datasets that were collected on I04-1 and I02 on two different crystal forms in the space groups P41212 (1.75 Å resolution) and P21 (1.55 Å resolution). The two structures are essentially identical (RMSD 0.37 Å), revealing the MCR-1 catalytic domain to have an overall α-β-α fold, containing three intramolecular disulphide bonds, typical of alkaline phosphatase or sulphatase enzymes, with the N-terminus indicating the likely position of the five transmembrane helices (Fig. 1b). However, the two structures differ in zinc content, with the P21 structure containing one zinc ion (Zn1), coordinated by Asp465, His466, Glu246 and Thr285 in a tetrahedral geometry (Fig. 1c), while the P41212 structure contains density for an additional, also tetrahedral, zinc site (Zn2) coordinated by His395, His478, a water molecule (Wat1) and Glu300 from a symmetry related molecule (Fig. 1d). Furthermore, the active site of the P21 structure contains electron density corresponding to the presence of a phosphate group attached to Thr285, the conserved residue believed to act as the acceptor for the PEA group during the transfer reaction.
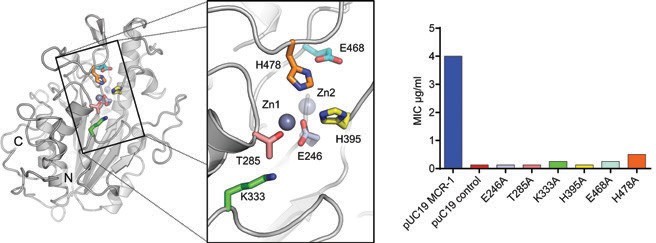
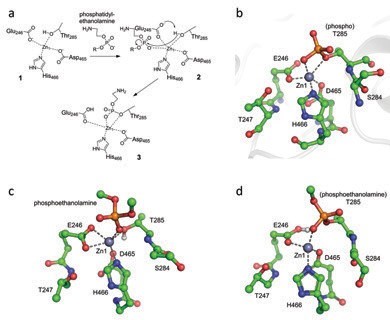
The presence of zinc in the active site is required for activity as MCR-1- positive strains were more susceptible to colistin on EDTA treatment. In addition, mutation of the Zn1 ligand Glu246 or the acceptor Thr285 reduces the colistin Minimal Inhibitory Concentration (MIC) of recombinant E. coli to that of vector-only control (Fig. 2). The effects of mutations at other active site residues are however more variable. Whilst replacement of the conserved His395, part of the Zn2 site (P41212 structure), reduces colistin MIC to basal levels, substitutions of the adjacent His478, or its hydrogen-bonding partner Glu468, give smaller reductions (3 to 4 dilutions). Similarly, the alanine mutation of Lys333 substantially reduces colistin MIC, but not to basal levels. Thus, at positions other than His395, alanine mutants away from the Zn1 site retain some ability to protect expressing strains from colistin action. Hence, while important, these residues are not absolutely essential to MCR-1 function. To further investigate the roles of the zinc sites (particularly Zn2, which is present in only one structure) computational density functional theory models of intermediates in the putative reaction mechanism were calculated for both a mono- and a di-zinc mechanism (Fig. 3). These data indicated that small structural changes required by the reaction are achievable within the constraints of the active site and that, in contrast to alkaline phosphatase enzymes, a single zinc ion may be sufficient to support at least the first stage of the MCR-1 catalysed reaction.
References:
Related publication:Hinchliffe P, Yang QE, Portal E, Young T, Li H, Tooke C, Carvalho MJ., Paterson NG., Brem J, Niumsup PR., Tansawai U, Lei L, Li M, Shen Z, Wang Y, Schofield CJ, Mulholland AJ, Shen J, Fey N, Walsh TR, Spencer J. Insights into the Mechanistic Basis of Plasmid-Mediated Colistin Resistance from Crystal Structures of the Catalytic Domain of MCR-1. Scientific Reports 7, 39392, doi:10.1038/srep39392 (2017).
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.