Related publication: Vita N, Platsaki S, Basle A, Allen SJ, Paterson NG, Crombie AT, Murrell JC, Waldron KJ, Dennison C. A four-helix bundle stores copper for methane oxidation. Nature 525, 140-143, doi:10.1038/nature14854 (2015).
Keywords: Copper; Copper storage; Methanotrophs; Methane oxidation; Particulate methane monooxygenase.
Methane is a more potent greenhouse gas than carbon dioxide, and atmospheric levels are increasing. Bacteria called methanotrophs can oxidise methane, using an enzyme called particulate methane monooxygenase (pMMO). pMMO requires copper in order to oxidise methane, and there are abundant pools of soluble copper within methanotrophs. Identifying the proteins that can bind copper in methanotrophs is important for potential uses of these organisms in reducing atmospheric methane, and in the industrial production of bioenergy, bulk, and fine chemicals.
Data from the MX beamline I02 at Diamond Light Source enabled the decoding of the structure of the copper-free form of the previously unknown copper-binding protein Csp1 from a methanotroph. The tuneable microfocus MX beamline (I24) at Diamond was used to determine how the structure of the protein changed when it was loaded with copper. X-ray absorption near-edge spectroscopy experiments on the same beamline also yielded the oxidation state of the Csp1-bound copper.
The data from these two beamlines shows that this new copper-storage protein forms a four-helix bundle, a structural feature found in many proteins. Csp1 is made up of a tetramer of these four-helix bundles, which together can bind up to 52 copper ions in a previously unseen manner: the binding happens mainly via the Cys amino-acids that point into the core of each four-helix bundle. Csp1 is the first example of a protein that stores a metal within an already established protein-folding motif (the four-helix bundle).
MX Village | Beamlines I02/I24
Methanotrophs (methane-oxidising bacteria) are nature’s primary mechanism of mitigating the release of the potent greenhouse gas methane to the atmosphere. The first step in the metabolism of methane for carbon and energy by methanotrophs is performed by methane monooxygenases (MMOs)1. Almost all methanotrophs possess the copper-requiring particulate pMMO that is housed on specialised intracytoplasmic membranes (Fig. 1). The fine structure of these membranes remains unknown and it is therefore currently not clear if they are discrete from the plasma membrane and whether pMMO is a very rare example of a bacterial cytoplasmic copper enzyme. Some methanotrophs are able to switch to using a soluble iron sMMO when copper is limiting. To satisfy their copper requirements methanotrophs secrete small modified peptides called methanobactins (mbtins) that bind copper with high affinity2,3. The structures of Cu(I)-mbtins have previously been determined using data collected on I02 and I042,3.
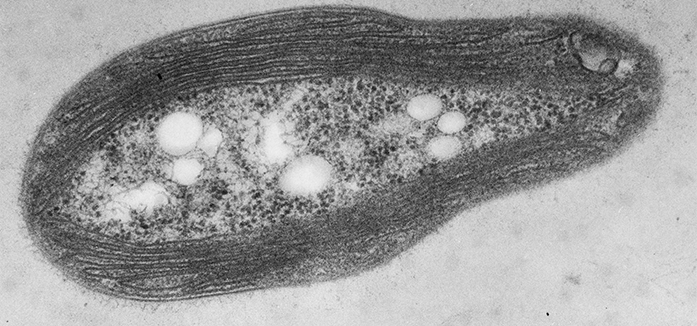
Figure 1: The structure of a methanotroph. Thin-section transmission electron micrograph of the methanotroph Methylosinus trichosporium OB3b showing the intracytoplasmic membranes around the periphery of the cell that house pMMO. It is currently not known if these are connected to the periplasm or form discrete compartments.
Copper-loaded mbtins are taken up by methanotrophs3,4, but their intracellular fate is not known. Whilst trying to locate internalised Cu(I)- mbtins using metalloproteomic analyses of soluble extracts from the model switchover methanotroph Methylosinus trichosporium OB3b a number of copper pools were observed. This was unexpected as M. trichosporium OB3b is predicted to possess only a single soluble copper protein. Detailed analysis of the most abundant copper peak lead to the identification of a previously unstudied 122 amino-acid protein with 13 Cys residues. Cloning of the gene followed by over-expression in Escherichia coli allowed purification of large quantities of the protein. In vitro studies showed that this predominantly α-helical protein forms a tetramer in solution that can tightly bind 12-14 Cu(I) ions per monomer (Fig. 2a) using Cys ligands, with an average Cu(I) affinity of approximately 1 × 1017 M-1. This indicated a role in copper storage, and the protein was therefore named Csp1 (copper storage protein 1).
The crystal structure of metal-free (apo) Csp1 was determined from data collected on I02 and shows the protein to be a four helix bundle with all 13 thiolate groups pointing into the core and the absence of disulfide bonds. Apo-Csp1 forms a tetramer in the asymmetric unit and the same arrangement is present in the crystal structure (Fig. 2b) of the protein loaded with Cu(I), which was determined using single-wavelength anomalous dispersion (SAD) data collected on I24 at the copper edge. The calculated anomalous difference density matched the additional electron density found within the core of the copper protein structure. To verify the oxidation state of the copper in the Csp1 crystals, X-ray absorption near edge spectroscopy was carried out on I24. The spectrum (Fig. 2c) has a distinctive peak at 8984 eV, indicative of Cu(I) with a low coordination number. The structure of Cu(I)-Csp1 confirms that the core of the four-helix bundle fills with 13 copper ions (Fig. 2b), with the majority of these bound by two Cys ligands. At the more open end of the bundle are Cu(I) sites at which His36 and Met48 are ligands (Fig. 2d). Met48 bridges between two Cu(I) ions and along with the nearby Met40 and Met43 may assist in recruiting the metal. The metal core of Cu(I)-Csp1 exhibits a rich array of coordination chemistry that has not been seen previously.
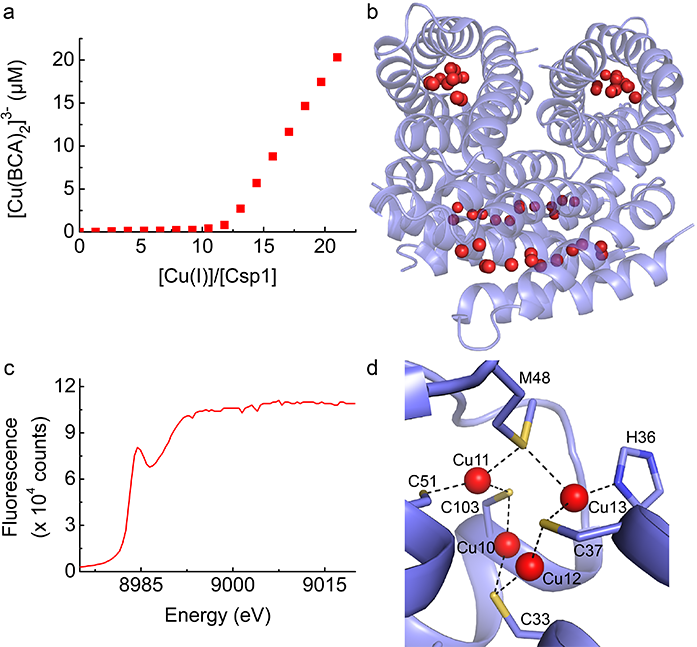
Figure 2: Copper binding by Csp1. a) A plot showing the formation of [Cu(BCA)2]3- upon addition of Cu(I) to apo-Csp1 in the presence of approximately 25 equivalents of the high-affinity chromophoric Cu(I) ligand bicinchoninic acid (BCA). The [Cu(BCA)2]3- complex starts forming after Csp1 has been fully loaded with Cu(I); b) The structure of the Cu(I)-Csp1 tetramer with 13 Cu(I) ions (red spheres) bound within the core of each four-helix bundle monomer; c) The X-ray absorption near edge spectrum of a crystal of Cu(I)-Csp1; (d) A more detailed representation of the structure of the Cu(I) sites close to the more open end of the four-helix bundle of Cu(I)-Csp1.
Excess copper is potentially toxic to cells and in eukaryotes metallothioneins, relatively short Cys-rich polypeptides, fold around copper clusters to safely store the metal. Related copper-binding bacterial metallothioneins have only been found in certain mycobacteria. Storing any metal within a well characterised protein-folding motif, such as a four-helix bundle, is unprecedented in biology. The Csp1 tetramer can store up to 52 Cu(I) ions and the presence of a predicted twin arginine translocase targeting signal peptide indicates folding, and potentially also metal binding, before export from the cytosol. Deletion of the genes for Csp1 and the close homologue Csp2 results in a mutant strain of M. trichosporium OB3b that switches to using sMMO more quickly than the wild type when copper becomes limiting, consistent with a role for these proteins in copper storage for methane oxidation. Mbtin has a Cu(I) affinity that is 4-5 orders of magnitude tighter than that of Csp1 and can readily remove all Cu(I) from Csp1. M. trichosporium OB3b produces mbtin when copper levels fall and under these conditions mbtin may be involved in removing Cu(I) from Csp1. Whether Csp1 and Csp2 locate to the periplasm, or to discrete pMMOhousing intracytoplasmic compartments (Fig. 1), remains to be determined. Discovering and understanding the function of proteins such as Csp1 that are involved in copper handling by methanotrophs is important for developing biotechnological applications of these organisms. This could include their use in approaches for reducing greenhouse gas activity as well as utilising methane as a feedstock for the production of biofuels and also bulk and fine chemicals.
A third Csp (Csp3) is present in M. trichosporium OB3b, but this lacks a signal peptide and must therefore be cytosolic. Bioinformatic studies have shown that Csp3 is present in a wide range of bacteria. This is another highly interesting outcome from this study as it is currently thought that bacteria do not maintain a cytosolic pool of copper due to the potential toxicity of this metal. Bacteria usually possess the copper-transporting P-type ATPase CopA that removes the metal from the cytosol, but the widespread presence of cytosolic Csp3s would imply a need to safely maintain a certain level of copper in this compartment, potentially for yet to be discovered copper proteins.
References:
- Hakemian, A. S. & Rosenzweig, A. C. The biochemistry of methane oxidation. In Annual Review of Biochemistry. Vol. 76, 223-241, doi: 10.1146/annurev.biochem.76.061505.175355 (2007).
- El Ghazouani, A. et al. Variations in methanobactin structure influences copper utilization by methane-oxidizing bacteria. Proceedings of the National Academy of Sciences of the United States of America 109, 8400- 8404, doi:10.1073/pnas.1112921109 (2012).
- El Ghazouani, A. et al. Copper-Binding Properties and Structures of Methanobactins from Methylosinus trichosporium OB3b. Inorganic Chemistry 50, 1378-1391, doi:10.1021/ic101965j (2011).
- Balasubramanian, R., Kenney, G. E. & Rosenzweig, A. C. Dual Pathways for Copper Uptake by Methanotrophic Bacteria. Journal of Biological Chemistry 286, 37313-37319, doi:10.1074/jbc.M111.284984 (2011).
Funding acknowledgements:
This work was supported by Biotechnology and Biological Sciences Research Council (grant BB/K008439/1 to C.D. and K.J.W.) and Newcastle University (part-funding of a PhD studentship for S.P.).
Corresponding author:
Professor Christopher Dennison, Newcastle University, christopher.dennison@ncl.ac.uk.


 Diamond Annual Review 2016
Diamond Annual Review 2016

