The crystallographic fragment screen against the SARS-CoV-2 Nsp3 macrodomain encompassed multiple fragment libraries: the DSI-poised library, MiniFrags (Astex), FragLites & Peplites (CRUK Newcastle Drug Discovery Unit (Newcastle University)), York3D (University of York), SpotFinder (Hungarian Academy of Sciences) the Edelris Keymical library and the EU OPENSCREEN fragment library.
The screen identified 88 binding events in the four sites of known biological interest (along with the usual artefactual hits that bind to the surface and crystal contacts):
- 54 fragments (58 binding events) bind to the adenine subsite of the active site, indicating how to compete with the natural ligand, i.e. ADP-ribose
- 9 fragments (10 events) bind to the proximal ribose subsite of the active site, again indicating how to compete with the natural ligand, i.e. ADP-ribose
- 7 fragments (8 events) bind near the catalytic loop indicating how to compete with the macrodomain interaction with ADP-ribosylated host proteins
- 8 fragments bind near a loop previously-observed to be involved in pyrophosphate binding
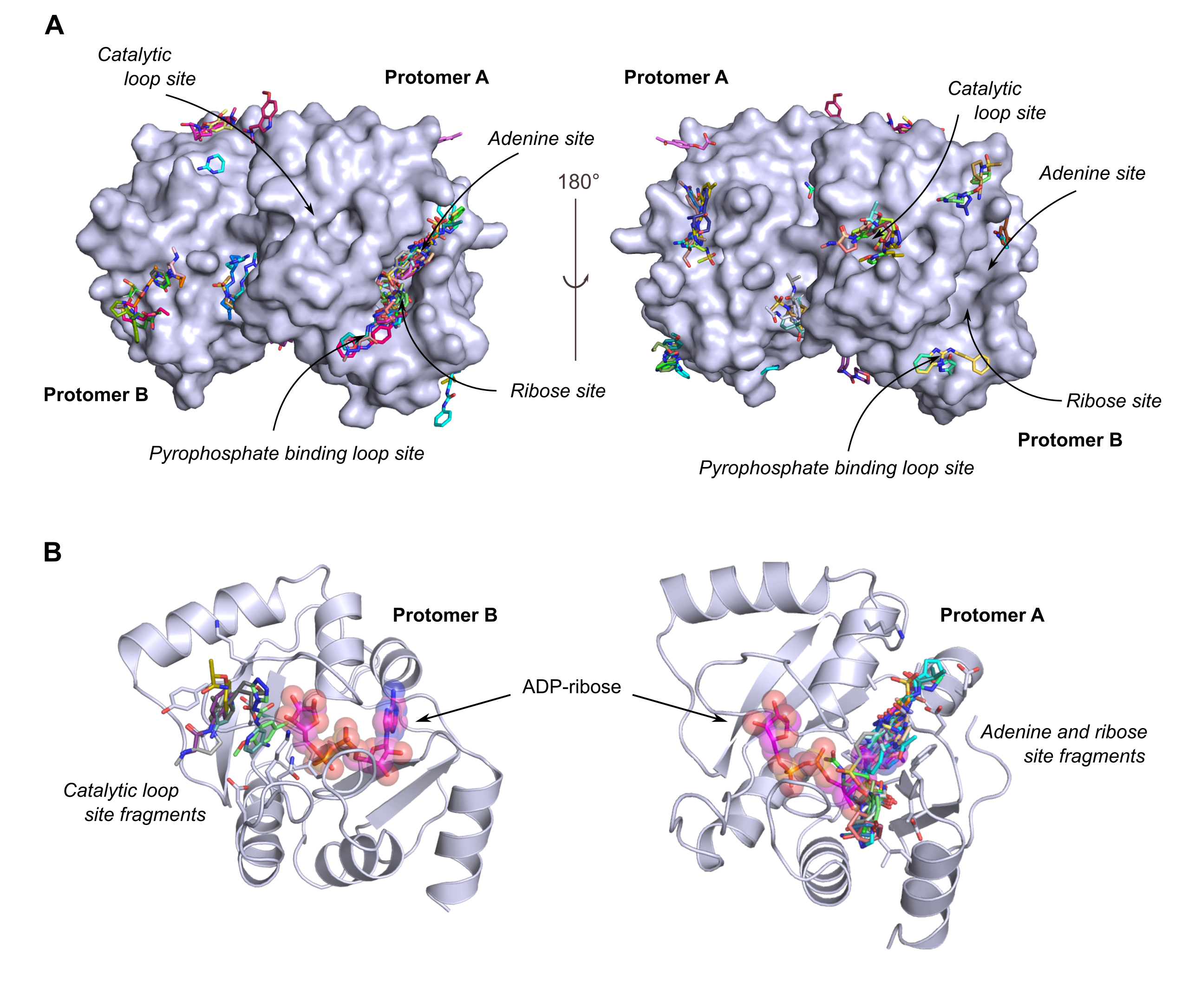
Figure 2. SARS-CoV-2 Nsp3 macrodomain fragment hits. (A) Fragment hits were identified at different sites around the macrodomain crystallographic dimer. Of particular interest are fragments binding to site 1 as being the adenine binding site of the natural ligand ADP-ribose and fragments binding to site 4 as being the substrate binding site which enables macrodomain interactions with ADP-ribosylated host proteins. The macrodomain is shown in surface representation and the fragments as atom-coloured stick model. (B) Overlay of fragments targeting the adenine (right) and substrate (left) binding site of the Nsp3 macrodomain. The natural ligand ADP-ribose in stick/sphere representation was superimposed to the selected structures.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.