Public Engagement at Diamond
Take a virtual tour of Diamond and learn more on how you can visit us here.
Find out how science gets done!
We have developed new scientific animated videos, demonstrating the work that has been done at Diamond's XChem facilities. Click here to explore the full series.
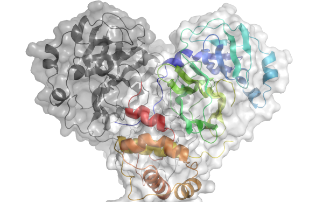
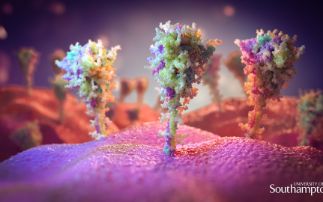
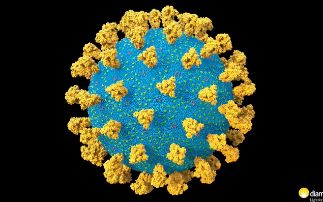
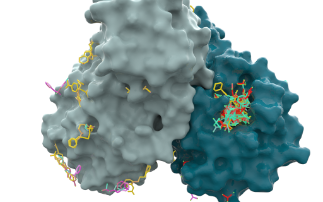
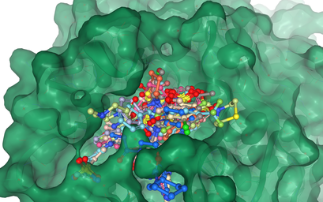
Diamond is currently working on a wide range of COVID-19 related projects that teach us more about the virus and how we can combat it. We are looking at the fundamental interactions of the virus, from which it is hoped new therapies can be developed. But we are also studying how existing drugs, that have already been tested and approved for other diseases, can be repurposed and used to treat patients. The array of specialised facilities at Diamond allow for many different techniques to be used, from looking at the structure of the virus and fitting drugs into it, like a tiny jigsaw puzzle, to taking direct images of the virus without its infectious component, so we can see how it interacts with drugs.
Shining our expertise on Coronavirus
Diamond in the News
Diamond’s work on COVID-19 related projects has been covered widely by the media. Here are some media highlights.
- On the 13th July, a number of media outlets, including BBC News, Sky News and The Independent published a piece outlining how Antibodies derived from llamas have been shown to neutralise the SARS-CoV-2 virus in lab tests.This fascinating research is the result of a collaboration between scientists from Diamond Light Source, the Rosalind Franklin Institute, Oxford University and Public Health England.
- On the 26th June, Channel News Asia broadcasted a video explainig how scientists are crowdsourcing a potential COVID-19 treatment.
- On the 21st June, the South African tv channel DStv released a short documentary showing how Diamond's powerful x-rays are helping to identify weak spots in the structure of SARS-CoV-2.
Have you seen Diamond in the news recently? Share your favourite stories with us on Twitter or Facebook.
Research updates from Diamond:
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.