A recently published article in the journal Nature highlights the collaborative environment for research at Diamond Light Source. New insights into the mechanisms of bacterial amyloid secretion were obtained by a team comprising researchers from seven institutions collecting data from four of Diamond’s macromolecular crystallography (MX) beamlines: I02, I03, I04 and I24.
Bacteria are particularly resourceful when it comes to protecting themselves, as evidenced by growing concern over the resistance of microbes, including bacteria, to antibiotics. Antimicrobial resistance, which was announced as the subject of the Longitude Prize 2014, reduces the effectiveness of treatment and leads to the increased occurrence of disease due to longer infectious periods.
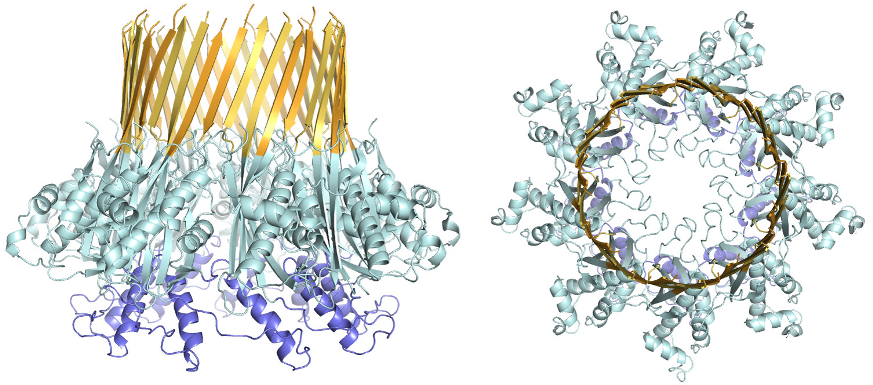
Atomic structure of the curli transport channel CsgG, viewed from the side (left) and top (right.)
Bacteria are able to create a biofilm to protect themselves from hazards in the surrounding environment. Such biofilms consist of a group of microorganisms in a matrix of extracellular polymers that act as a protective barrier, in which bacteria can weather out onslaughts from antibiotics or the immune system. By studying the bacterial biofilm structure, scientists aim to inhibit growth of components that contribute to biofilm formation. For many bacteria, sticky protein fibres called ‘curli’ are a major constituent of the biofilm matrix. Using a dedicated protein transport system, bacteria deliver curli building blocks from the inside to the outside of the bacteria, after which they self-assemble into an extensive network of curli fibres that aid the production of the protective biofilm capsule.
Dr Parveen Goyal and Dr Han Remaut of Vrije Universiteit Brussel and VIB, an inter-university institute of life sciences, led the investigation into the three-dimensional structure of the protein transport system that leads to the assembly of curli on the bacterial cell surface. “The structure determination of transmembrane protein complexes is often a lengthy process” Dr Remaut explains. “Frequent access to the Diamond MX beamlines, which are at the forefront of the field in terms of technical performance and data acquisition tools, allowed us to screen through large sample numbers and find those rare crystals of good enough quality to allow structure determination.”
Using beamlines I02, I03, I04 and I24, the group have been able to create the first three dimensional image of the CsgG protein, the pore through which the building blocks of curli fibres cross the bacterial wall. Based on the structure of this curli secretion channel, a detailed model for its mode of action can be produced. With these new insights, ways can be sought to prevent bacteria forming these virulence-associated fibres.
In addition to the fundamental importance of this discovery, mapping out the assembly mechanism of curli is of interest in biotechnological applications. The curli subunits can serve as building blocks for physically robust nanowires, whose properties can be modified by adding foreign peptides and proteins on the curli subunits. Also the biofilm properties can be of interest. Although antimicrobial resistance is currently high on the scientific agenda, biofilms of non-pathogenic bacteria find applications for example in wastewater treatment. Research is ongoing to interpret the skilful strategies of the microbial world.
For more information about the MX beamlines, see the individual web pages or contact the relevant Principal Beamline Scientist:
I02: Thomas Sorensen: [email protected]
I03: Katherine McAuley: [email protected]
I04: Dave Hall: [email protected]
I24: Robin Owen: [email protected]
Goyal P., Petya V. Krasteva P. V., Van Gerven N., Gubellini F., Van den Broeck I. Troupiotis Tsaïïaki A., Jonckheere W., Péhau-Arnaudet G., Pinkner J. S., Chapman M. R., Hultgren S. J., Howorka S., Fronzes R. and Remaut H. Structural and mechanistic insights into the bacterial amyloid secretion channel CsgG. Nature (2014). DOI: 10.1038/nature13768
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.