A new approach to inhibit the flu virus
The World Health Organization estimates that worldwide, annual influenza epidemics result in about 3‑5 million cases of severe illness and about 250,000 to 500,000 deaths. Influenza A viruses are the most common cause of seasonal flu in humans and represent a significant risk to public health.
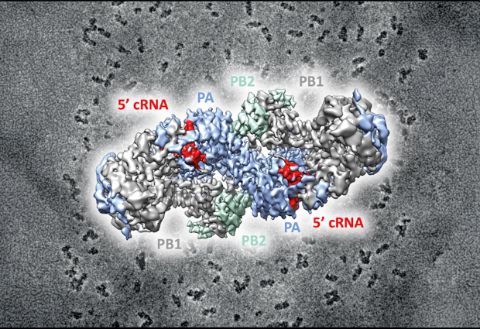
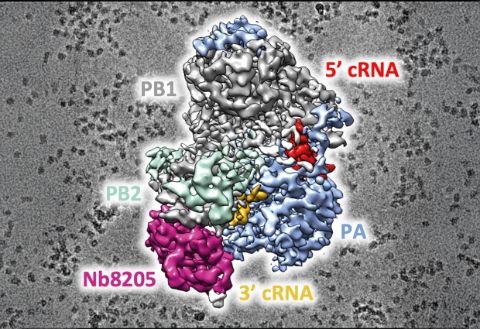
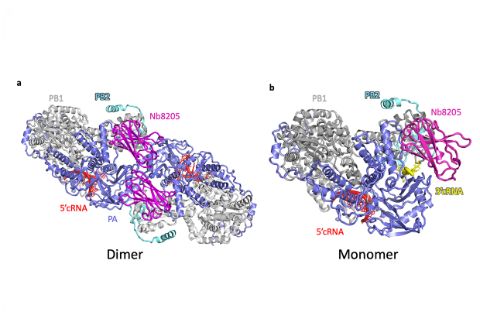
When the influenza virus infects a host cell, it starts to make copies of itself as the disease spreads. The protein driving this behaviour is the viral RNA polymerase which replicates the viral RNA genome and makes RNA templates for protein synthesis. In order to understand how this process works, you need to work out the structure of the RNA polymerase at the atomic level.