Despite decades of study, the mechanisms that HIV employs at certain steps of its replication cycle remain relatively nebulous. One such step involves the viral regulatory protein, Rev, whose principal function is to affect the nuclear export of a majority of the viral mRNA transcripts that are produced during infection. Rev achieves the specific transport of the viral RNAs by oligomerizing onto the Rev response element, a highly-structured RNA motif within an env intron. Without such RNA shuttling functionality, the major viral proteins are not produced. Rev is therefore both essential for HIV infection and a promising target for novel drug development for the treatment of AIDS. No small-molecule Rev inhibitors have been developed to date primarily because of the lack of structural data on Rev itself. The regulatory protein has thwarted all efforts toward crystallization until recently as a result of its propensity to polymerize into long, helical filaments in vitro. Consequently, we have engineered an antibody to inhibit Rev polymerization, purified and crystallized the Fab-Rev complex, and determined the structure of the Rev dimer with bound Fab at 3.2 Å resolution, which has facilitated the development of structural models for Rev-Rev multimerization and Rev-RNA association, critical molecular events in the HIV replication cycle.
After HIV enters a newly infected cell, the viral genome is integrated into the host cell genome as a single 9-kb provirus. Yet in order to produce 15 viral proteins from one ORF, the virus must employ a complex pattern of downstream RNA processing whereby over 30 different mRNA transcripts are alternatively spliced. Rev is a 13 kDa regulatory viral protein that facilitates nuclear export of the majority of these transcripts, specifically the unspliced and partially-spliced viral mRNAs that are ultimately used both as genomic RNA for packaging into assembling virions and for translation into many of the viral proteins. In the absence of Rev, these intron-containing transcripts would never be exported to the cytoplasm due to eukaryotic splicing regulation. Since Rev’s function is so closely linked with the production of the structural and enzymatic viral proteins, Rev itself is viewed as eliciting the transition from early to late phase HIV infection.
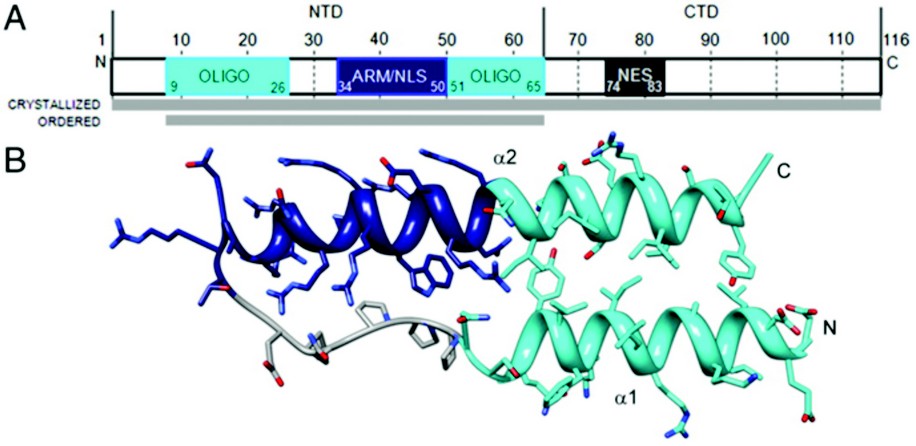
Figure 1: Rev monomer. (A) Domain organization of Rev. Oligomerization motifs depicted in cyan, arginine-rich motif (ARM)/nuclear localization sequence (NLS) in navy blue, and nuclear export signal (NES) in dark gray. Although full-length Rev was crystallized, only the N-terminal domain was ordered. (B) Ribbon representation of Rev monomer (top view) showing the helix-loop-helix motif with coplanar helices.
Rev targets viral mRNAs by recognizing a ~350 nucleotide, highly-structured region within an env intron known as the Rev response element (RRE). Rev contains an arginine-rich RNA-binding motif (ARM) that specifically binds to a purine-rich bulge within stem loop IIb of the RRE with high affinity. This interaction nucleates assembly of an RNP complex of 200-300 kDa containing 8-10 Rev molecules that are recruited, one at a time, in a highly cooperative process mediated primarily through Rev-Rev interactions. The fully-formed Rev-RRE complex then binds a collection of host cell proteins through its nuclear export signal (NES), such as Crm-1 (exportin-1), GTP-bound Ran, and many others, which together promote nuclear export of the RRE.
The HIV Rev structure was first solved via 3.2 Å crystallographic data collected on a single rod-shaped crystal of the Fab-Rev complex on Diamond Light Source’s beamline I02 . Molecular replacement solutions of the Fabs were used for initial phasing and the Rev structure was subsequently built de novo. The P1 crystal’s unit cell contains six Fab fragments and six Rev monomers, arranged as heterotetramers, each comprising a Rev dimer flanked by two Fabs. Although full-length Rev was present in the Rev-Fab crystals, only residues 9-65—the N-terminal domain (NTD)—were found to be ordered. The NTD adopts a helix-loop-helix motif (helical hairpin), consistent with previous proposals (Fig. 1). One prong of the hairpin consists of an a-helix (a1), residues 9 to 24, followed by the loop region from residue 25 to 33; a longer a-helix (a2), residues 34 to 65, comprises the second prong. The two faces of this planar platform are denoted A and B. Helix a1 and the C-terminal half of a2 are amphipathic, with hydrophobic and polar surfaces on opposing sides of the helix. The N-terminal half of a2 is the RRE-binding ARM, densely populated with Arg and Glu residues. Nonpolar residues cluster around the juxtaposed surfaces of a1 and a2 to form a hydrophobic core that serves at least two purposes: (i) stabilization of the hairpin; and (ii) presentation of hydrophobic surface patches, one on each face of the hairpin that mediate interaction with other Rev monomers.
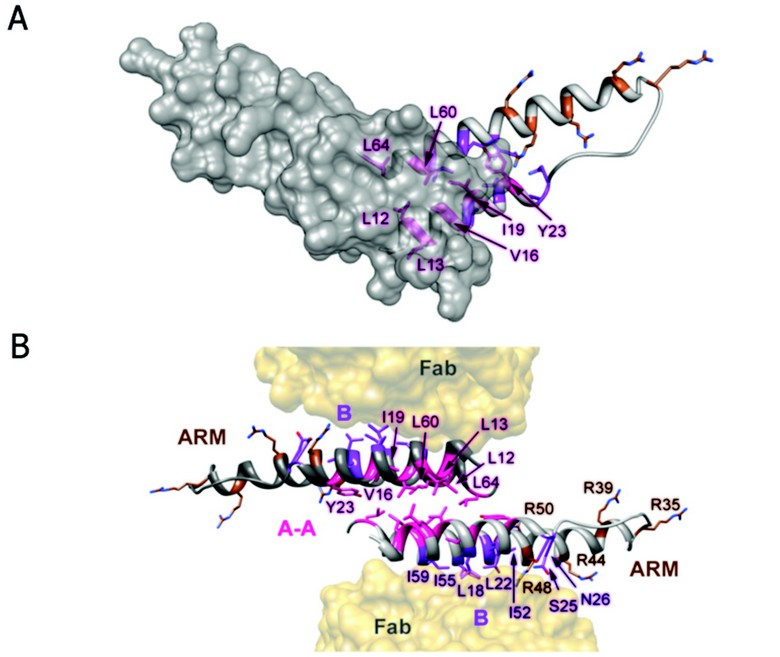
Figure 2: Rev dimer. (A) Top down view of Rev dimer depicting residues involved in A-A interface in pink, residues implicated in B-B oligomerization and RRE binding in purple and brown, respectively; one Rev subunit is shown by isosurface rendering. (B) Ribbon representation of Rev dimer, depicting the A-A dimer interface and the B oligomerization faces contacting the Fabs. The oligomerization surfaces and ARM motifs are segregated, consistent with the cooperative model of Rev multimerization on the RRE (coloring as in A).
The Fab-Rev crystal structure reveals molecular dimers of Rev flanked by Fabs (Fig. 2). The Rev dimer interface is formed by the hydrophobic patch on surface A at the pronged end of the hairpin, where the subunits overlap by two helical turns. The two subunits are related by an almost exact twofold axis (between 177 and 180º) and rest across each other at an angle of ~140º. In the dimer, identical A-surfaces mate—i.e., in a head-to-head as opposed to a head-to-tail interaction—while the B surfaces interact with Fabs. Indeed, a Rev oligomerization model based on the analysis of assembly-deficient mutants supports the idea of an alternating series of symmetrical interactions: A-A, B-B, A-A, etc. It is noteworthy that the hydrophobic patch on surface B coincides with the Rev epitope for this Fab (Fig. 2). In the absence of Fab, Rev dimers could assemble further via B-B interactions as well. The hydrophobic patches required for dimerization, and (by inference) higher order assembly, localize to the pronged end of the Rev monomer and are thus segregated from the RNA-binding ARM at the other end. As the hydrophobic residues L12, L13, V16, I19, L60, and L64 consistently bury more than 50% of their accessible surface areas at the A-A interface, it is clear that they make key stabilizing interactions, consistent with a wealth of biochemical and genetic data on the functional relevance of specific residues in Rev oligomerization. However, biophysical analysis of such mutants has shown that reduced RRE-binding and nuclear export activities are more closely correlated with destabilization of Rev tertiary structure than simply the ablation of oligomerization faces A and B.
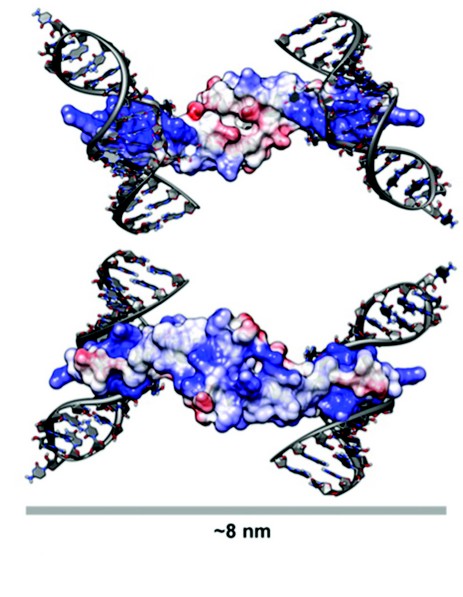
Figure 3: A Model for Rev Oligomerisation. Rev dimer in the context of Stem Loop IIB (SLIIB) high-affinity RRE-binding site. The two views are related by a 180° rotation about a horizontal axis. SLIIB RNA nucleotides from NMR structure of SLIIB bound to Rev alpha helix (res 33–55) were superposed onto the Rev dimer (rmsd of 0.8 Å over main chain atoms of residues 37–55). The width of the dimer-RNA structure above is 8 nm, consistent with the 8nm diameter of Rev-RNA filament assemblies as observed in EM. Molecules are colored according to surface charge, from acidic (Red) to basic (Blue).
The Rev dimer structure also provides insight into Rev-RRE interactions, which was modeled by superposing the NMR structure (PDB ID: 1ETF) of the stem loop IIb-ARM complex onto the Rev crystal dimer. After superposition, the RNA helix is positioned such that there are no steric clashes with the Rev dimer and there is ample clearance for the binding of additional dimers (Fig. 3). The two ARMs per dimer are at opposite ends of this elongated molecule, and when one subunit is modeled as RRE-bound, the other subunit cannot bind to an adjacent groove of the same RNA helix, given the orientation of RNA in the NMR structure. As such, two stem loop IIb RNAs were modeled onto the dimer, one on each side, the dyad axis aligning the two RREs into a track upon which additional Rev molecules could feasibly bind. In this model, stacked Rev dimers present arrays of ARMs spaced 20-25 Å, consistent with such an array making repetitive interactions with successive major grooves along the same side of an RNA helix. Because it has been shown that the ARM can bind RNA with different helical faces, Rev has some flexibility with which to bind additional RNA grooves as it multimerizes. While the structure of the RRE is largely unknown, it is plausible that its tertiary structure provides an opposing second RNA helix for binding on the opposite side of dimer from the initial stem loop IIb interaction.
Future work will endeavor to study the assembled structure of the full Rev-RRE ribonucleoprotein complex that forms to effect nuclear export of viral mRNAs. It will be interesting to see how the RNA structure limits association of only 8-10 Rev molecules, either by altering the oligomerization interface structures or possibly by capping the ends of the RNA-bound Rev oligomer. The Rev structure is also being utilized toward efforts for structure-based drug design of Rev inhibitors of oligomerization, potentially giving rise to a novel class of HIV small molecule therapeutics.
DiMattia, M.A., Watts, N.R., Stahl, S.J., Rader, C., Wingfield, P.T., Stuart, D.I., Steven, A.C. and Grimes, J.M. Implications of the HIV-1 Rev dimer structure at 3.2 Å resolution for multimeric binding to the Rev response element. Proc. Natl. Acad. Sci. USA. 107:5810-5814 (2010)
Additional Reference
Stahl, S.J., Watts, N.R., Rader, C., DiMattia, M.A., Mage, R.G., Palmer, I., Kaufman, J.D., Grimes, J.M., Stuart, D.I., Steven, A.C., Wingfield, P.T. Generation and characterization of a chimeric rabbit/human Fab for co-crystallization of HIV-1 Rev. J. Mol. Biol. 397:697-708 (2010).
Funding Acknowledgement
This work was supported in part by the Intramural Research Programs of the National Institute for Arthritis and Skin Diseases and the National Cancer Institute, the National Institutes of Health (NIH) Intramural Targeted Antiviral Program, the NIH–Oxford Scholars Program, by the UK Medical Research Council and SPINE2COMPLEXES (LSHGCT-2006-031220).
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.