Dr James Huntington, University of Cambridge
Ordered protein aggregation is thought to be the cause of several diseases including Alzheimer’s, Huntington’s, Parkinson’s, the prion encephalopathies, and many more. However, the molecular basis of the intermolecular linkage of the underlying protein (Aβ1-42, huntingtin, α-synuclein, and prions) is not known for any of these ‘conformational diseases’ [1]. The SERPIN family of proteins also cause disease when mutations lead to their intracellular aggregation. The process has become known as polymerisation, due to the ordered ‘beads-on-a-string’ appearance by electron microscopy and laddering on non-denaturing gels [2]. The deposition of serpin polymers in the endoplasmic reticulum of secretory cells presumably occurs via a folding intermediate and efforts to prevent or reverse the process depend on knowledge of the folding pathway and the intermolecular linkage. We recently solved the crystal structure of a serpin dimer that suggested an unexpected ‘domain-swapping’ mechanism of polymerisation [3]. This structure explained the hyper-stability of the intermolecular linkage and why polymerisation occurs during folding in vivo, or upon mild denaturation in vitro. The closed dimer can be ‘opened up’ to form a linear dimer with persistent donor and acceptor ends that can propagate infinitely. The hyperstability of the serpin polymers may play a role in their accumulation in the endoplasmic reticulum, and contribute to their toxicity. There are several crystal structures of domain swapped proteins, and it has been suggested that domain swapping might be the molecular basis behind certain of the conformational diseases (especially the prion ecephalopathies) [4,5]. However, the serpins are the only known example of proteins that become hyperstable upon domain swapping. This feature sets them apart, explains their accumulation and suggests a mechanism of toxicity.
The serpins constitute a family of proteins, most of which are serine protease inhibitors [6]. They share a common mixed α/β fold composed of nine helices, three β-sheets, and a 20-residue loop, known as the reactive centre loop (RCL). In the native state, the main β-sheet (sheet A) contains five strands, and the RCL begins just after the fifth strand of sheet A (s5A, Fig. 1A). The unusual thing about the serpins is that the native conformation is not its most stable. Insertion of the RCL into β-sheet A as its 4th strand converts serpins into a hyperstable state (ΔΔ G of approximately -32kcal/mol) [7]. This can occur in the presence of proteolytic nicking of the RCL to form the ‘cleaved’ state, or in its absence resulting in the ‘latent’ state. This stored energy is crucial for the inhibitory function of serpins, as illustrated by the crystal structure of the serpin-protease complex (Fig. 1B) [8]. The RCL acts as bait and allows the chemistry of proteolysis to commence, and at the acyl enzyme intermediate stage the serpin rapidly incorporates the RCL into β-sheet A, flinging the protease from one end of the serpin to the other. The resulting distension of the catalytic loop effectively re-zymogenises the protease, preventing the breakdown of the covalent (ester) bond between the protease and the serpin. Thus, the functioning of serpins requires a folding pathway that bypasses the hyperstable RCL-inserted state in favour of a metastable native state. Perhaps unsurprisingly, this requirement renders serpins highly sensitive to mutations, almost anywhere (Fig. 1A), that perturb the folding pathway and lead to the loss of secreted functional serpin [9].
Electron micrographs of serpin polymers, obtained from patient samples or formed in vitro using mild denaturation (typically heat, chaotropic agents, or low pH), revealed a ‘beads-on-a-string’ morphology, with occasional selfterminating ‘necklaces’ [10]. This material ran on native PAGE as discrete ladders, and was resistant to denaturation by either heat or urea, and could only be broken into monomers by the combination of SDS and prolonged boiling. Peptides based on RCL sequences were found to anneal into β-sheet A of native serpins in the s4A position, suggesting that RCL incorporation in trans might be the basis of the hyperstable protomer-protomer interaction [2, 1]). This ‘loop-sheet’ mechanism has gained prominence in the field and was widely accepted, in spite of serious conceptual and practical limitations (e.g. requires folding to the native state before formation of polymers; loop-sheet polymers would be rigid, and RCL incorporation in trans cannot fully occupy the s4A position).
We set out to determine the molecular basis of serpin polymerisation by X-ray crystallography, but knew that to do so would require the isolation of a compact ‘closed’ polymer (like the necklaces seen by EM). Such a dimer had been found in the blood plasma of patients with a mutation in the serpin antithrombin (AT) [12], and although we were unable to obtain sufficient amounts of this material for crystallisation, incubation of purified wild-type AT at pH 5.7 and 37°C produced a dimer with similar properties. Crystals were eventually obtained using the in vitro-made closed dimer. An initial low resolution dataset was collected on Diamond Beamline I02, and later a better quality dataset was obtained (to 2.8 Å) on Beamline I04. The structure was solved by molecular replacement using the program Phaser [13], with latent AT providing the best model.
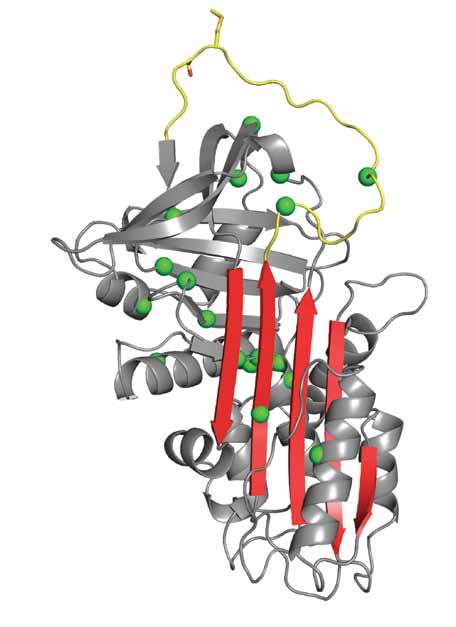 Fig 1.A Fig 1.A |  Fig 1.B Fig 1.B |
| Fig. 1. Serpin structure and mechanism. (A) Ribbon diagram of a native serpin in the classic orientation (RCL, yellow, on top and β-sheet A, red, facing). The strands of β-sheet A are numbered, and the scissile bond (P1-P1’ as sticks) is indicated. Positions of polymer-causing mutations for several serpins are shown as green balls. (B) The final serpin-protease complex has an ester bond between the catalytic Ser195 and the P1 residue (shown), and is stabilized due to the distension of the oxyanion hole (blue balls). The RCL has incorporated as the fourth strand in β-sheet A, resulting in the hyperstable six-stranded conformation. | |
The asymmetric unit contained two latent-like protomers in a back-to-back configuration (Fig. 2A). In contrast to what would be expected for a ‘loop-sheet’ dimer, we found that each monomer had swapped the entire length of both strands 4 and 5A, and the coiled (but stably folded) region just following helix I and preceding s5A. The intermolecular contact was thus a contiguous 50 residue ‘domain’ swap, that resulted in the completion of β-sheet A in the neighbouring protomer. This dimer explained why partial denaturation (unfolding) of serpins is required to form polymers in vitro, since s5A and the coiled region preceding s5A must be extracted from the rest of the molecule in order to participate in the swap. The thermodynamic drive for the observed swap is easy to understand, since the swap results in the mutual completion of β-sheet A with only the short s1C unfolding to permit the return to the parent protomer.
A linear dimer, capable of forming longer polymers (Fig.2B), was made by breaking one linkage and unravelling helix I and some of the coiled region. To test if this limited unfolding occurs in linear polymers of AT and the prototypical serpin α1-antitrypsin (α1AT), we incubated polymers with low concentrations of specific proteases AspN and LysC. Polymers of both AT and α1AT were susceptible to proteolysis under these conditions, whereas native and cleaved controls were not, and specific cleavage was observed for both polymers in the predicted linker region. Other experiments, including disulfide trapping, were conducted in support of the general nature of the two-strand domain swapping mechanism. Although the structure and mechanism of action of serpins have been well characterised, how mutations result in the intracellular accumulation of polymers has remained unknown. Why the accumulation of serpin polymers is cytotoxic is unclear, but our structure suggests that it may be due in part to the hyperstability of the linkage, which might inhibit proteosome-dependent clearance, or alternatively, to the exposure of hydrophobic linker regions, which might mediate the lateral association of polymers into high-order aggregates [14]. Our structure of a self-terminating serpin dimer provides a novel paradigm for serpin polymerisation (extensive domain swapping) and a useful model for the polymerigenic folding intermediate. The challenge now is to determine if this, and/or another domain-swap is responsible
for polymerisation in vivo, and indeed, whether the precise mechanism depends on the serpin and the mutation involved.
|
Fig 2.A |
Fig 2.B |
| Fig. 2. Domain-swapped serpin polymer. (A) Ribbon diagram of the crystal structure of the closed AT dimer, with one protomer coloured red and the other yellow. The domain swap begins just after helix I and continues through the RCL (s4A). (B) Breaking one of the dimer contacts allows the creation of a linear dimer, that can be extended to a pentamer (shown), and beyond. Each protomer is in a different colour. | |
References
[1] Carrell RW & Lomas DA, Conformational disease, Lancet 350:134-138 (1997).
[2] Lomas DA, Evans DL, Finch JT, & Carrell RW, Nature, 357:605-607 (1992).
[3] Yamasaki M, Li W, Johnson DJ, & Huntington, JA, Nature, 455:1255-1258 (2008).
[4] Bennett MJ, Sawaya MR, & Eisenberg, D Structure, 14:811-824 (2006).
[5] Liu Y & Eisenberg, D Protein Sc,i 11:1285-1299 (2002).
[6] Silverman GA et al, J Biol Chem, 276:33293-33296 (2001).
[7] Im H, Ahn HY, & Yu MH, Protein Sci, 9:1497-1502. (2000)
[8] Huntington JA, Read RJ, & Carrell RW, Nature, 407:923-926 (2000).
[9] Gooptu B & Lomas DA, Annu Rev Biochem (2009).
[10]. Lomas DA, Finch JT, Seyama K, Nukiwa T, & Carrell RW, J Biol Chem, 268:15333-15335 (1993).
[11] Schulze AJ et al, Eur J Biochem, 194:51-56 (1990).
[12] Corral J et al, J Thromb Haemost, 2:931-939 (2004).
[13] McCoy AJ, Grosse-Kunstleve RW, Storoni LC, & Read RJ, Acta Crystallogr D Biol Crystallogr, 61:458-464 (2005).
[14] Huntington JA, Sendall TJ, & Yamasaki M, Prion 3 (2009).
Principal Publications and Authors
Yamasaki M, Li W, Johnson DJ, Huntington JA. Crystal structure of a stable dimer reveals the molecular basis of serpin polymerization. Nature; 455(7217):1255-8 (2008).
Huntington JA, Sendall TJ, & Yamasaki M. New insight into serpin polymerization and aggregation. Prion 3 (2009).
Funding Acknowledgement
Medical Research Council, UK.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.