Using X-ray crystallography to solve the structure of proteins that carry unpaired electrons, called radicals, has proven challenging by traditional crystallography methods because they become chemically altered once irradiated with X-rays. Researchers overcame this hurdle using a rapid form of the technique in which they captured protein structure within 1,000,000,000,000,000th of a second, too quick for the radical to change. They found that the unpaired electron had a large influence over the protein’s structure, and they hope their work will inform the design of therapeutics that target radical proteins.
Some enzymes in the body carry radicals, chemical groups with highly reactive unpaired electrons, to catalyse biochemical reactions, but it has proven challenging to study the structure of these enzymes with the radicals intact. The X-ray crystallography techniques conventionally used to study protein structure introduce ‘X-ray damage’ that would neutralise radicals and alter the protein structure. To study how radicals influence proteins, researchers turned to a ribonucleotide reductase enzyme subunit called R2 that uses a radical to synthesise DNA bases. The team previously used X-ray crystallography at beamline I24 and small angle X-ray scattering at beamline B21 of the Diamond Light Source to solve the structure of this enzyme without safeguarding the radical. In the recent study, they harnessed X-ray free electron laser (XFEL) serial femtosecond crystallography at the Linac Coherent Light Source in collaboration with Diamond’s XFEL Hub to zero in on the radical. With XFEL, they used X-rays to rapidly capture the structure of the protein within femtoseconds — 1015 times quicker than a second and too quick for X-rays to neutralise the radical or distort protein structure. By comparing the enzyme with and without the radical, they revealed that the presence of the unpaired electron greatly influences the structure of the enzyme’s active site. Their research will allow them to explore the workings of this and related enzymes in finer detail and holds promise for designing drugs that target radical enzymes in cancer cells and infectious microbes.
Free radicals — molecules carrying a lone, unpaired electron — can cause damage to cells if they enter the body because they readily react with proteins and DNA. But not all radicals are bad. Some found naturally inside your body are essential for synthesising the four bases that make up DNA — A, C, G, and T. These radicals are found at the active site of so-called ribonucleotide reductase enzymes, which kickstart the process of synthesising bases.
Researchers have attempted to explore the structure of these essential enzymes in the past, but they were unable to capture the radical at work. Scientists have tried using cryo-electron microscopy but were unable to determine the structure in enough detail to see the radical. X-ray crystallography has been used at higher resolutions, but it comes at a different cost: significant X-ray damage to the active site made it impossible to study the enzyme with an intact radical, leaving scientists in the dark about the radical’s role.
In the new study, researchers turned to X-ray free electron laser (XFEL) serial femtosecond crystallography, an alternative X-ray technique. With regular X-ray crystallography, a large crystal of purified protein is typically illuminated with an X-ray beam, and an X-ray diffraction pattern is detected after a few minutes or hours, by which time X-ray damage may have distorted the protein. With XFEL however, many tiny microcrystals of protein are illuminated with an X-ray beam, and the diffraction pattern is captured only a few femtoseconds later — 1015 times shorter than a second. Because the data is captured so quickly, the radicals don’t have time to react or change and X-ray damage won’t occur: this is called the “diffraction-before-destruction” principle. Professor Martin Högbom of Stockholm University, who led the study, said that “there are a lot of enzymatic systems that utilise radicals in biology,” and now we have a technique to study them.
This was the first time that researchers used an XFEL to study a radical in a ribonucleotide reductase enzyme, and the researchers had to find the simplest form of the enzyme to boost their chances of success. They turned to an enzyme called R2 from the bacterium Mesoplasma florum whose structure they previously determined without capturing the radical using small-angle X-ray scattering (SAXS) and X-ray crystallography at Diamond Light Source. They discovered that it has an unusual property: unlike other ribonucleotide reductase enzymes, this one doesn’t involve metal ions at its active site. This made the enzyme easier to work with because the researchers didn’t need to worry about combining the purified protein with metal ions in the correct amounts before working out its structure.
Diamond’s XFEL Hub contributed to this project by helping create a novel sample delivery strategy to rapidly deliver tiny droplets of a microcrystal slurry of R2 protein to the X-ray interaction region. Like an assembly line in a factory, the microcrystals are shuttled along a conveyor belt called a ‘tape drive’ towards the X-ray beam. “The sample handling at these machines is a research project in itself. To get one structure, you need thousands of microcrystals to be injected into the beam because at every shot the crystal is destroyed,” Högbom said. The major advantage of this system was that the sample delivery and chamber environment could be controlled to avoid neutralizing the radical.
Högbom and his team set out to compare the structure of the enzyme with and without the radical to see how much it influences the shape of the active site. They produced protein samples both with and without the radical before crystallisation and resolved the structures of both down to the ångström scale, which is small enough to detect individual atoms. “This is the first protein radical that I know of that has been properly structurally characterised,” Högbom said. His team found that a triangle of three amino acids (tyrosine, aspartate, and lysine) lying at the heart of the active site were greatly affected by the presence of the radical (figure 1). “You remove one electron, and the whole protein moves,” he said. “It appears spring loaded in that sense.”
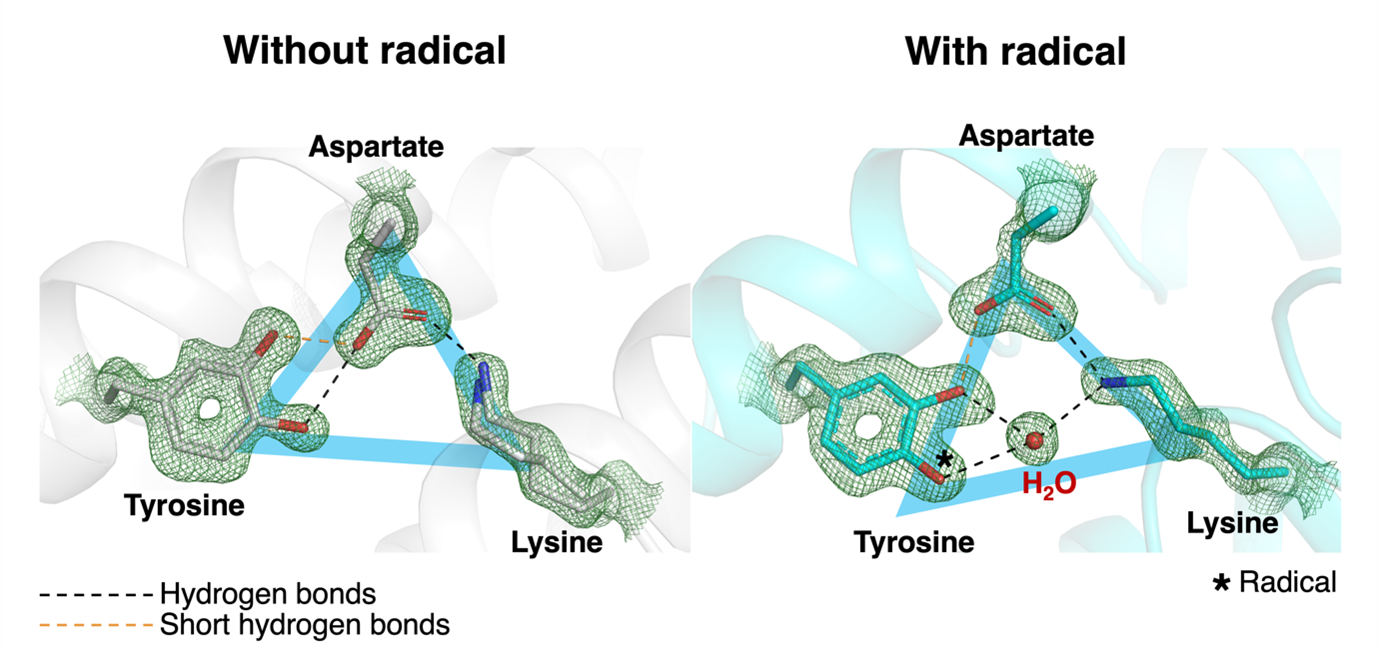
They also observed very short hydrogen bonds connecting aspartate and tyrosine in both the presence and absence of the radical. Such hydrogen bonds have specific properties and can increase the strength of attraction between the two amino acids, which the team proposed could stabilise those residues to prevent them from spuriously interfering with the radical reaction, for example by donating a proton. They also found that amino acids on the outskirts of the triangle played important roles in radical activity (figure 2).
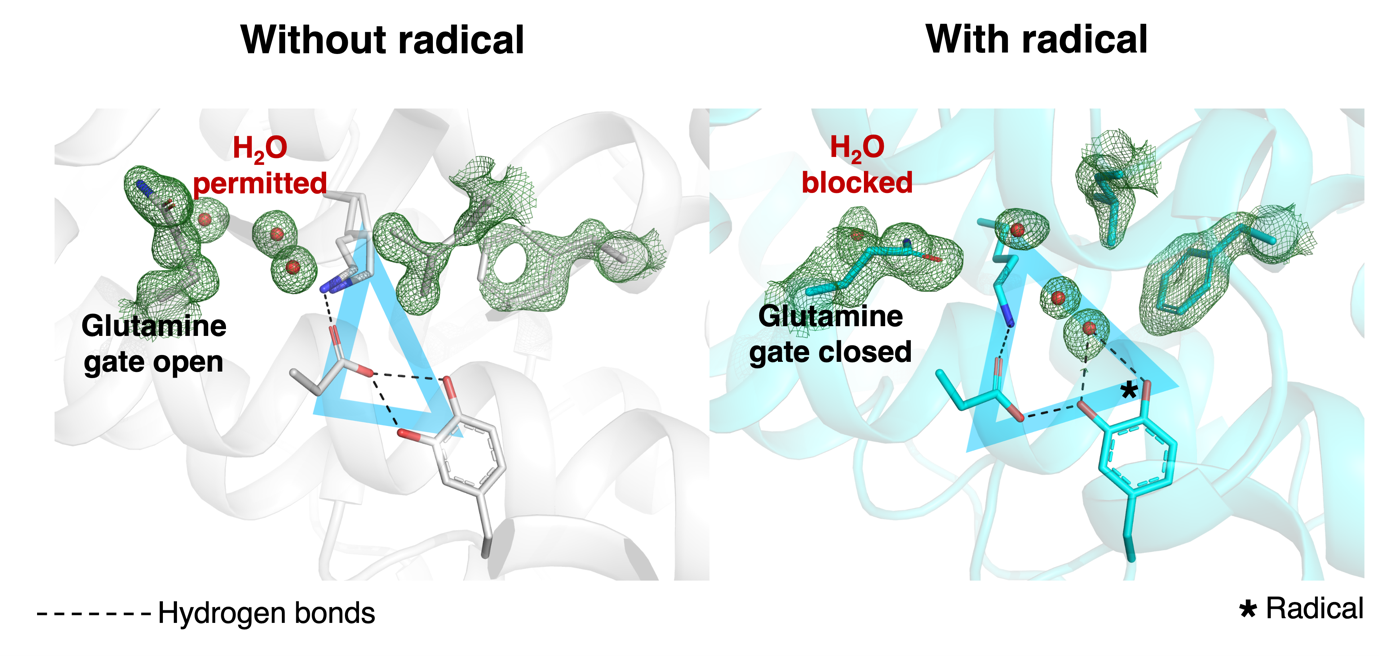
The active site looked strikingly different with and without the radical, highlighting the importance of XFEL techniques for visualising radical biochemistry. Understanding how radicals control biochemical reactions like synthesising DNA bases could be vital to targeting them therapeutically. Enzymes that synthesise bases will be more active in cells that divide rapidly, like cancer cells and infectious microbes, so these enzymes might make good drug targets in principle, Högbom said.
To find out more about the XFEL Hub at the Diamond Light Source, or to discuss potential applications, please contact Principal Scientist Allen Orville: [email protected].
Lebrette H, Srinivas V, John J, Aurelius O, Kumar R, Lundin D, Brewster AS, Bhowmick A, Sirohiwal A, Kim I, Gul S, Pham C, Sutherlin KD, Simon P, Butryn A, Aller P, Orville AM, Fuller FD, Alonso-Mori R, Batyuk A, Sauter NK, Yachandra VK, Yano J, Kaila VRI, Sjöberg B, Kern J, Roos K, Högbom M. Structure of a ribonucleotide reductase R2 protein radical. Science 382(6666), 109-113 (2023). doi: 10.1126/science.adh8160
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.