First structure of a DNA crosslink repair ligase determined
Nov 4, 2019
Nov 4, 2019
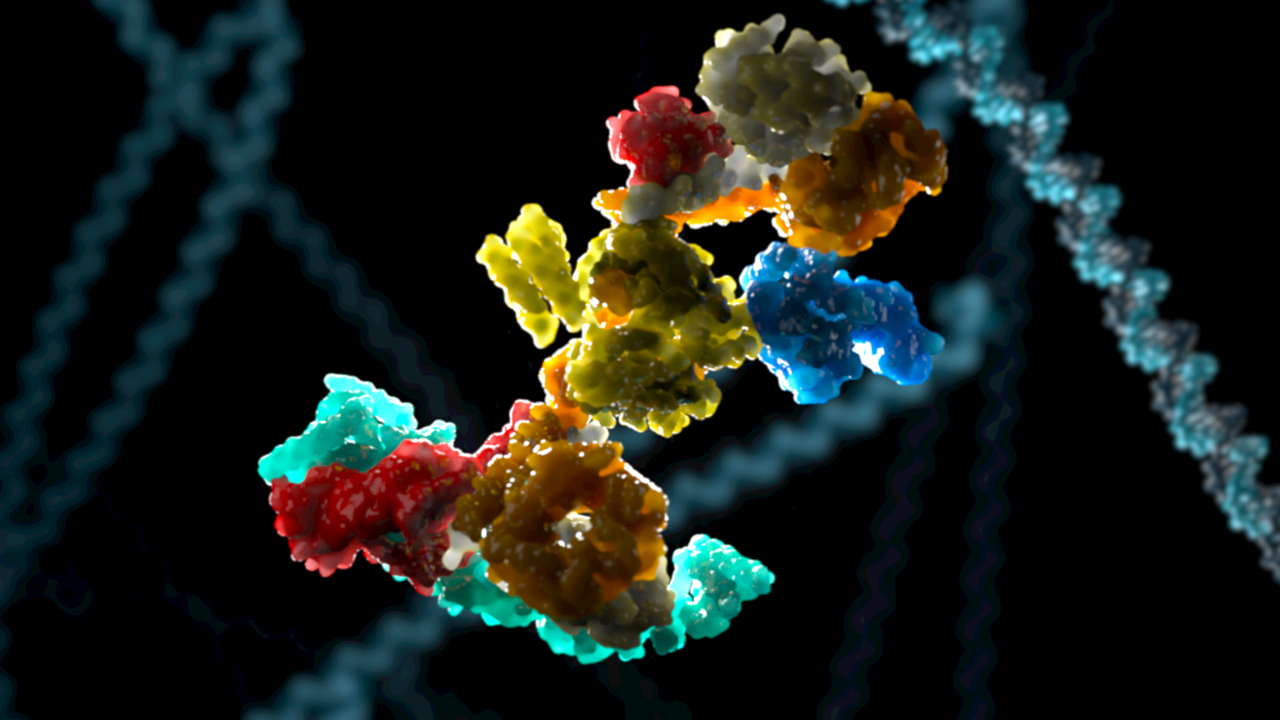
The FA core complex, image credit Phospho Biomedical Animation
Diamond’s Electron Bio-Imaging Facility (eBIC) has been used to generate the first 3D structure of the Fanconi anaemia (FA) core complex, a multi-subunit E3 ubiquitin ligase required for the repair of damaged DNA. The work, led by Dr Lori Passmore from the MRC Laboratory of Molecular Biology and a team of researchers, has been published today in Nature, and their research provides the molecular architecture of the FA core complex and new insights into how the complex functions.
The FA pathway senses and repairs DNA crosslinks that occur after exposure to chemicals including chemotherapeutic agents and alcohol, but also as a result of normal cellular metabolism. The megadalton FA core complex acts as an E3 ubiquitin ligase to initiate removal of these DNA crosslinks, helping to repair the damage caused. The research team used eBIC’s imaging facilities to make a major breakthrough in understanding the FA core complex by determining its structure using an integrative approach including cryo-electron microscopy and mass spectrometry.
Dr Peijun Zhang, Director of eBIC notes that:
Enabling cutting-edge research like this is exactly why we established eBIC, to provide scientists with state-of-the-art experimental equipment and expertise in the field of cryo-electron microscopy, for both single particle analysis and cryo-electron tomography. Determining the structure of the FA core complex for the first time is a fantastic achievement for the MRC research team.

The FA core complex is comprised of eight stably-associated subunits. Progress in understanding the mechanistic basis of ubiquitination has been slowed by the challenges in obtaining pure protein in sufficient quantities, an incomplete description of subunit functions and interactions, and a lack of high-resolution structures. In addition, many subunits do not have homology to proteins of known structure.
Dr Lori Passmore of the MRC Laboratory of Molecular Biology says:
The FA core complex is disrupted in diseases including Fanconi Anaemia and cancer, but it is also of immense interest due to its role in a fundamental DNA repair pathway, and because the mechanisms of monoubiquitination are unknown. Thus, understanding the structure and function of the FA core complex has been a central question for many years.
“FA core complex was a challenging structure because of its inherent flexibility, therefore, we required a large dataset and that’s where eBIC came in and helped us in collecting excellent data which we able to merge to our existing in-house data”, explained Dr Shabih Shakeel.
The structure of the FA core complex revealed several surprises. Firstly, the structure was asymmetric - it contained two copies of three subunits and one copy of the remaining five. This was unexpected because previous studies of subcomplexes showed them to be symmetric and containing two copies of each protein. Secondly, the two copies of FANCL (the RING finger subunit) were in different conformations, implying two different functions. Lastly, the majority of the FA patient mutations mapped to peripheral regions which are less important for the structural integrity compared to the central regions of the structure.
To learn more about Diamond’s eBIC facility, please visit: https://www.diamond.ac.uk/Instruments/Biological-Cryo-Imaging/eBIC.html
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Copyright © Diamond Light Source. Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.