Keep up to date with the latest research and developments from Diamond. Sign up for news on our scientific output, facility updates and plans for the future.
Every year is an exciting one at the Membrane Protein Laboratory (MPL) with so many positive results and amazing collaborations. Over the past years, the MPL has become a well-established user facility providing a state-of-the-art pipeline from protein production to high throughput protein crystallisation for the membrane protein structure determination community. Its proximity to Diamond’s beamlines has greatly facilitated excellent working relations and collaborations between the beamline scientists, and both the MPL staff and users. Today, more than 18 membrane protein structures are as a result of the MPL and more than 30 publications acknowledge the use of the facility.
This year the MPL reports two more structures resulting from the use of the MPL. The first is the structure of the Lipid A phosphoethanolamine transferase from Neisseria meningitidis (NmEptA) which has resulted from a collaboration between the MPL, University of Western Australia and several other institutions in Australia and USA1 (Fig. 5). The family of Lipid A phosphoethanolamine transferases in Gram-negative bacteria confers bacterial resistance to innate immune defensins and colistin antibiotics. This structure provides a strong basis for a structure-guided approach to develop small molecule inhibitors to combat multidrug resistance in pathogenic Gram-negative bacteria. Currently multidrug resistance Gram-negative bacteria are estimated to cause 700,000 deaths per year globally with a prediction that this figure could reach 10 million a year by 2050. The MPL has contributed to the establishment of the protein purification protocol, initial crystallisation, and initial crystal hit screening. During the project, the MPL hosted the Principle Investigator, Professor Alice Vrielink, for two months, in addition to her research assistant who also visited the MPL for another three months
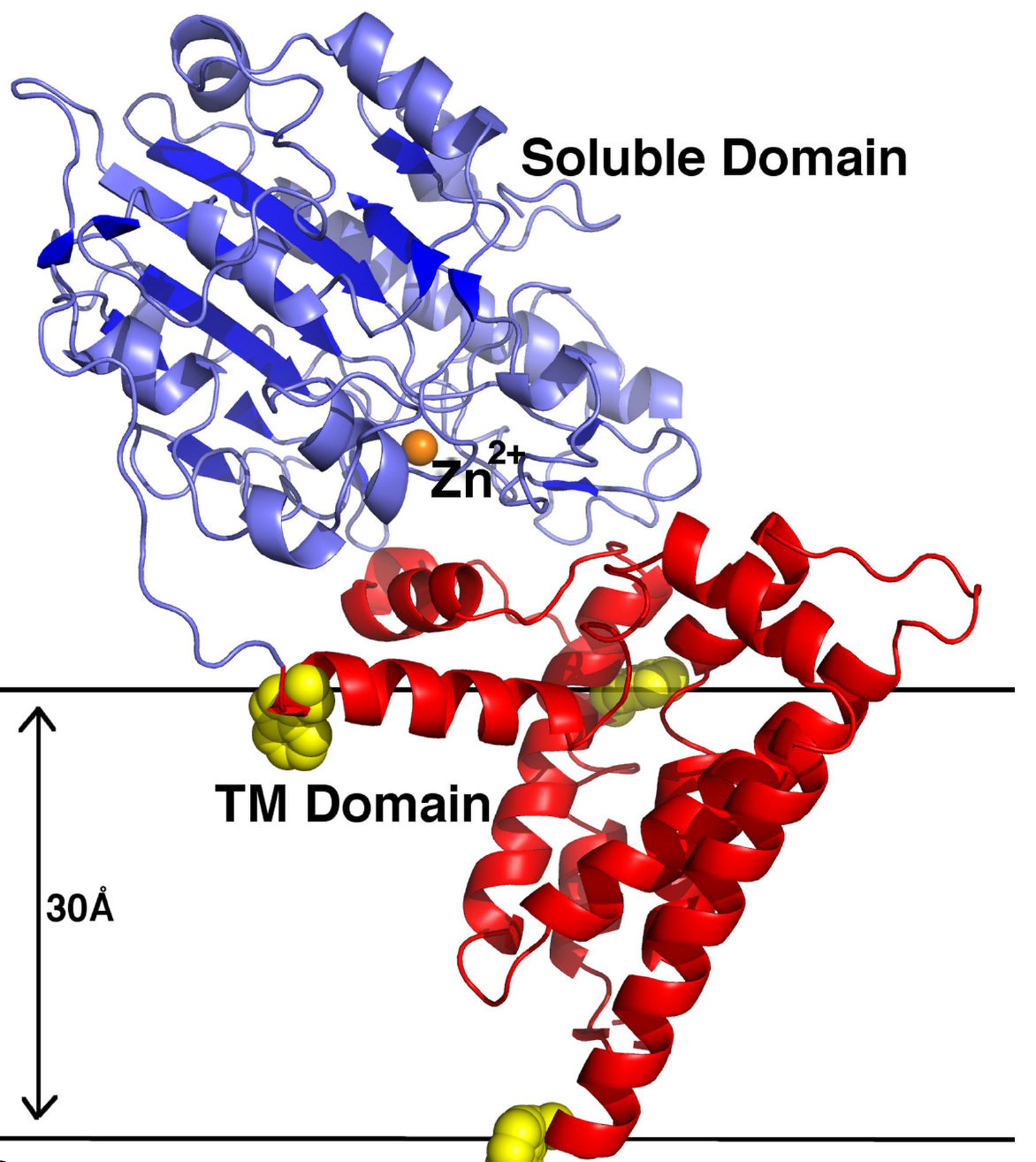 Figure 1: A ribbon representation of NmEptA. The amino terminal TM domain is shown in red and the carboxyl terminal soluble domain is shown in blue. The side chains of the three TM domain tryptophan residues (Trp126, Trp148 and Trp207) are shown as yellow spheres.
Figure 1: A ribbon representation of NmEptA. The amino terminal TM domain is shown in red and the carboxyl terminal soluble domain is shown in blue. The side chains of the three TM domain tryptophan residues (Trp126, Trp148 and Trp207) are shown as yellow spheres.
The second structure reported this year is the Phospho-N-acetylmuramoyl-pentapeptide translocase (MraY) transmembrane enzyme essential in the bacterial peptidoglycan synthesis pathway in complex with the nucleoside antibiotic tunicamycin'(Fig. 4). MraY is also a promising target for developing novel antibiotics2. This work was supported by the NanoMem project consortium (FP7-PEOPLE-2011-ITN) which the MPL was also part. The MPL has helped this project with the crystallisation and data collection strategy.
References:
1. Anandhi A., Evans G.L., Condic-Jurkic K., O’Mara M.L., John C.M., Phillips N.J., Jarvis G.A., Wills S.S., Stubbs K.A., Moraes I., Kahler C.M., Vrielink A. Structure of a lipid A phosphoethanolamine transferase suggests how conformational changes govern substrate binding. Proceedings of the National Academy of Sciences 114.9, 2218-2223 (2017).
2. Hakulinen J.K., Hering J., Brändén G., Chen H., Snijder A., Ek M., Johansson P. MraY-antibiotic complex reveals details of tunicamycin mode of action. Nature Chemical Biology 13(3), 265-267 (2017).

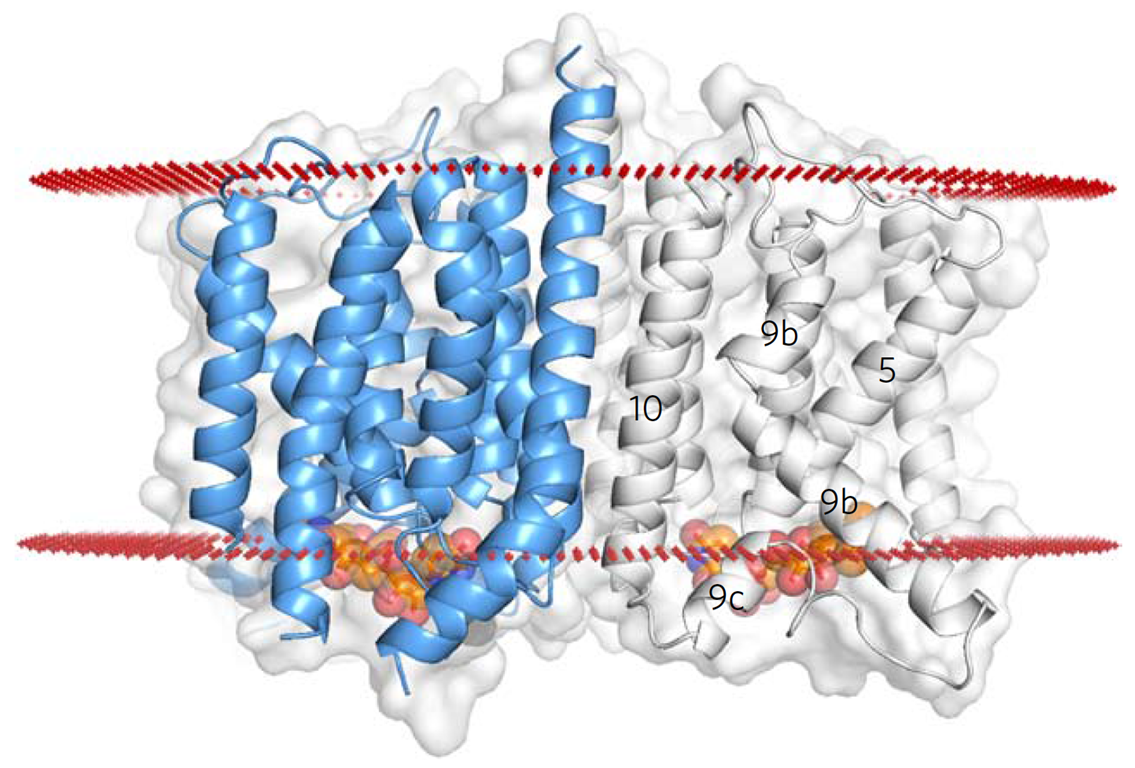
Figure 2: The MraY tunicamycin complex structure. The bound tunicamycin shown in orange.

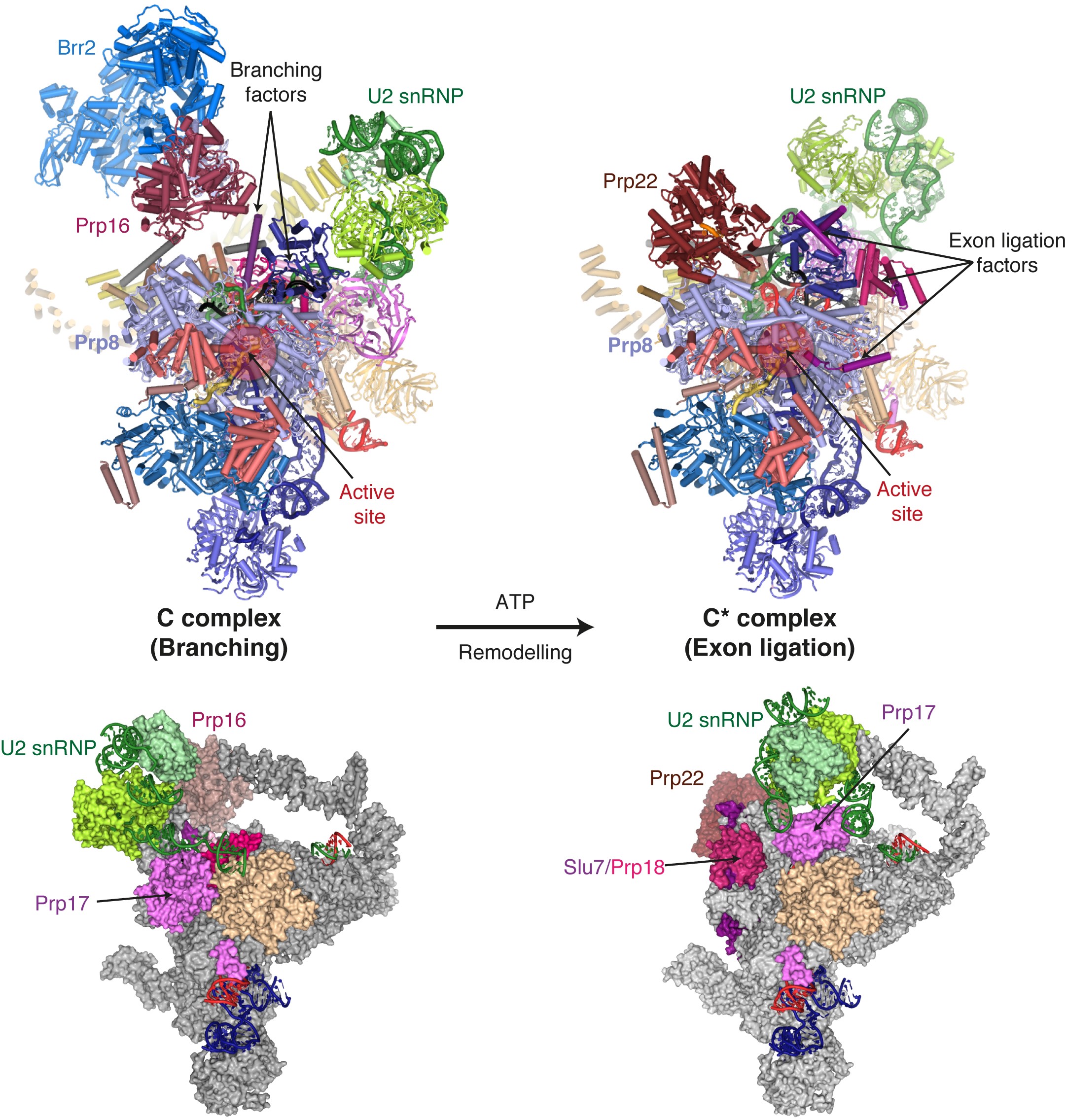
Figure 1: The Prp16-mediated remodelling of the spliceosome through key elements of our new C* structure compared to our previous C complex structure.
Related publication:
Controlling a molecular axe
Breaks in double stranded DNA are potentially lethal for a cell, thus different mechanisms for DNA repair have evolved. Homologous recombination is the most precise and preferred mechanism of DNA repair in many bacteria, where the break in the damaged chromosome is restored by recombination with the equivalent region of an intact sister chromosome. RecBCD complex is responsible for processing the broken ends of double stranded DNA to prepare them for DNA strand exchange. Given its complexity and tight regulation, RecBCD is sometimes referred as an example of a ‘smart’ macromolecular machine.
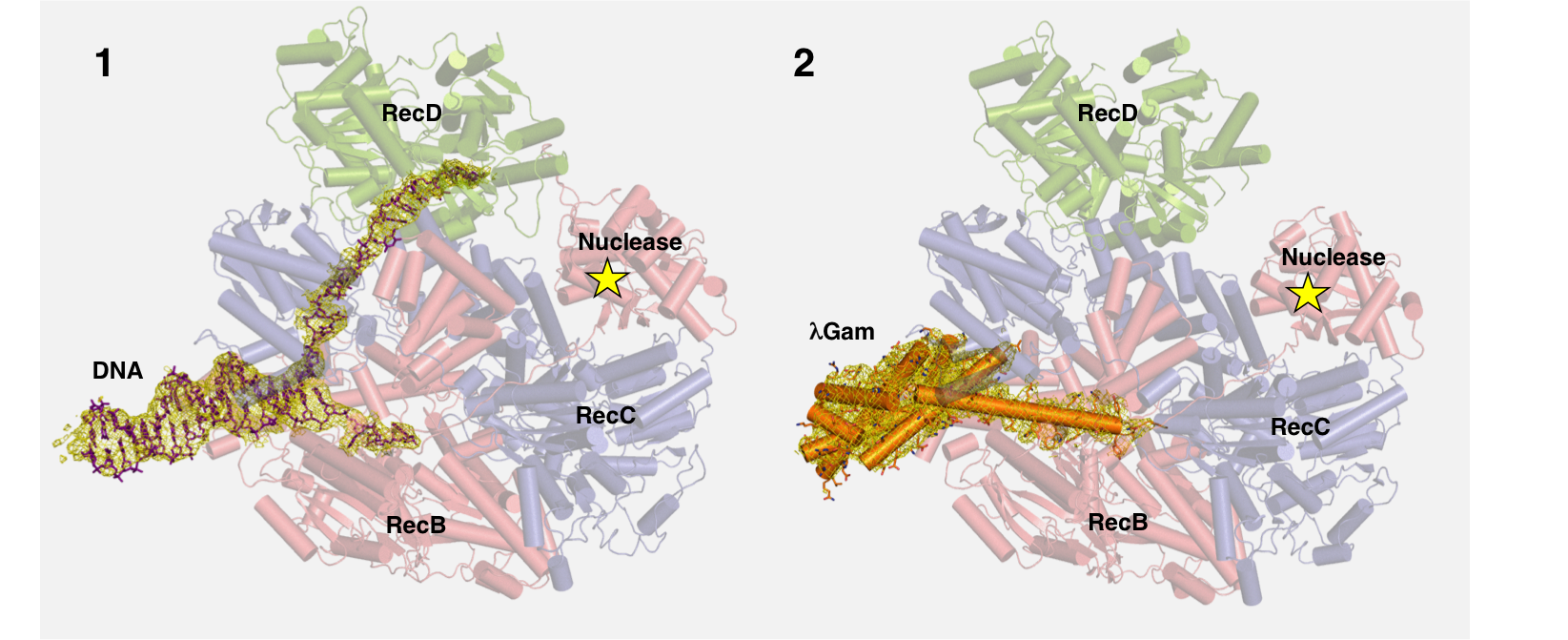 Figure 4a: Structure of DNA-RecBCD complex reveals how propagating strand of DNA causes conformational changes in RecD (green), which lead to activation of nuclease in RecB (indicated by star).
Figure 4a: Structure of DNA-RecBCD complex reveals how propagating strand of DNA causes conformational changes in RecD (green), which lead to activation of nuclease in RecB (indicated by star).
Figure 4b: Protein λGam from bacteriophage λ selectively blocks DNA entrance into RecBCD preventing it from digesting viral DNA. EM density surrounding the DNA and Gam protein respectively are displayed as a yellow mesh, contoured at 4 sigma.
1. Wilkinson M., Chaban Y., Wigley D.B. Mechanism for nuclease regulation in RecBCD. eLife, 5:e18227 (2016).

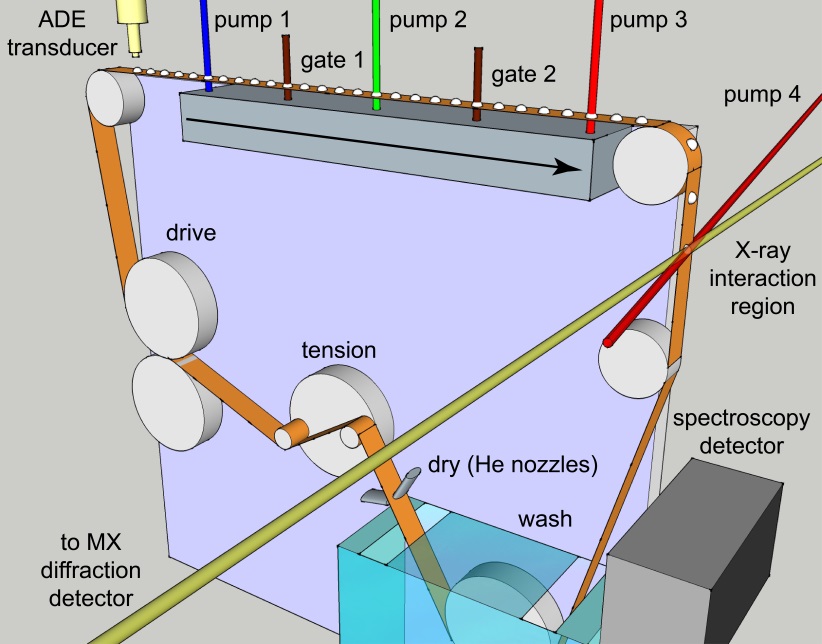
Figure 7: Schematic view of the sample delivery system. Droplets of few nanolitres containing protein crystals are deposited on the Kapton tape via Acoustic Droplet Ejection. The conveyor belt will bring the droplets to the interaction region with the X-ray pulse after exposing them to laser flashes.
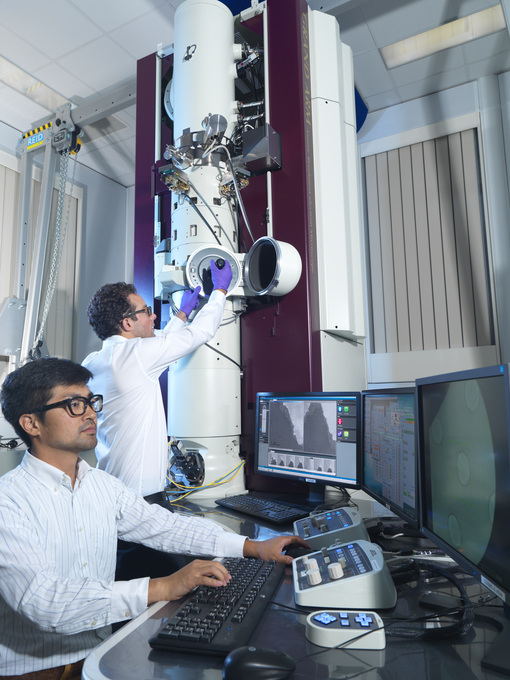
 Figure 1: Acting Principal EM Staff Scientist, Chris Allen, with Akira Yamagishi from JEOL UK (supplier of the microscopes) working with one of the microscopes at ePSIC.
Figure 1: Acting Principal EM Staff Scientist, Chris Allen, with Akira Yamagishi from JEOL UK (supplier of the microscopes) working with one of the microscopes at ePSIC.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.