Two of the Macromolecular Crystallography (MX) beamlines (I02 and I04) at Diamond Light Source were used to initiate the study and solve the crystal structure of a large fragment of WipA. The structure showed that the protein possessed a serine/threonine phosphatase fold that surprisingly targeted tyrosine-phoshorylated peptides. Additionally, it was hypothesised that WipA could transition from a homo-dimeric state to a hetero-dimeric state to interact with a tyrosine-phosphorylated host target. The discoveries will help to clarify the molecular mechanisms underpinning Legionella pathogenicity and could aid the development of inhibitors.
Infected vapour
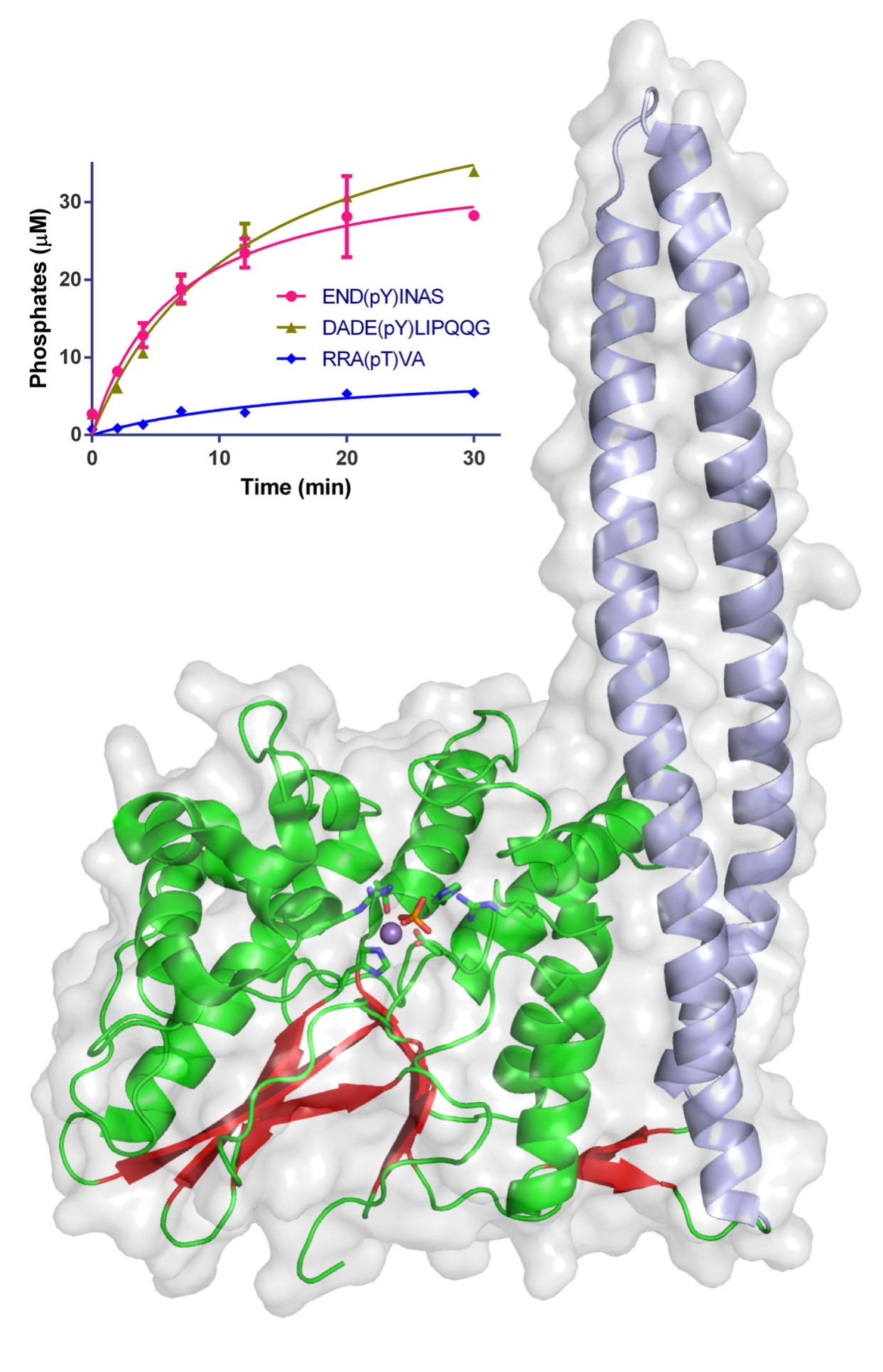
Figure 1: Structure of WipA in cartoon and semi-transparent surface representation. The phosphatase domain is represented in red-green and the attached coiled-coil in light blue. Residues involved in catalysis are represented as sticks around the Mn2+ and the phosphate ions. The surface is shown in light grey. The inset graph displays the time-course desphosphorylation of specific peptides using 20 nM of WipA.
To find out more about the I04 beamline, or to discuss potential applications, please contact Principal Beamline Scientist Prof Dave Hall: david.hall@diamond.ac.uk. The I02 beamline has since closed and been replaced by the fully-automated VMXi beamline, which will welcome users later this year. To find out more about the original I02 beamline or the VMXi beamline, please contact Principal Beamline Scientist Prof Thomas Sorensen: thomas.sorensen@diamond.ac.uk
Pinotsis N and Waksman G. Structure of the WipA protein reveals a novel tyrosine protein phosphatase effector from Legionella pneumophila. Journal of Biological Chemistry (2017) [Epub ahead of print]. DOI: 10.1074/jbc.M117.781948
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.