The structure of an oxidase protein from a key cyanobacteria pathway has been uncovered for the first time, using the new Long-Wavelength Macromolecular Crystallography beamline (I23) at Diamond Light Source. This is the first novel structure to be solved on the I23 beamline using the sulfur single-wavelength anomalous diffraction (S-SAD) phasing method.
The protein, known as ThcOx, is important in the pathway to produce patellamide, which is a cyanobactin (a group of small peptides found in cyanobacteria). Understanding the structure of this protein could help in engineering enzymes with many biotechnology applications, particularly in so called ‘green transformations’ that would otherwise require expensive chemical processes.
However, the selenium variant of the ThcOx protein does not crystallise, making the use of the common selenomethionine substitution to experimentally determine phases unworkable.
Extensive molecular replacement attempts failed to solve the structure, and initial attempts at determining a structure by sulfur phasing using anomalous diffraction data collected on a standard beamline were also unsuccessful. However, S-SAD phasing using I23 meant researchers obtained data at significantly longer wavelengths compared to standard synchrotron beamlines and this did lead to structure determination.
The structure of this protein therefore represented a particularly challenging test for the I23 beamline, and the successful determination of its structure validates its performance.
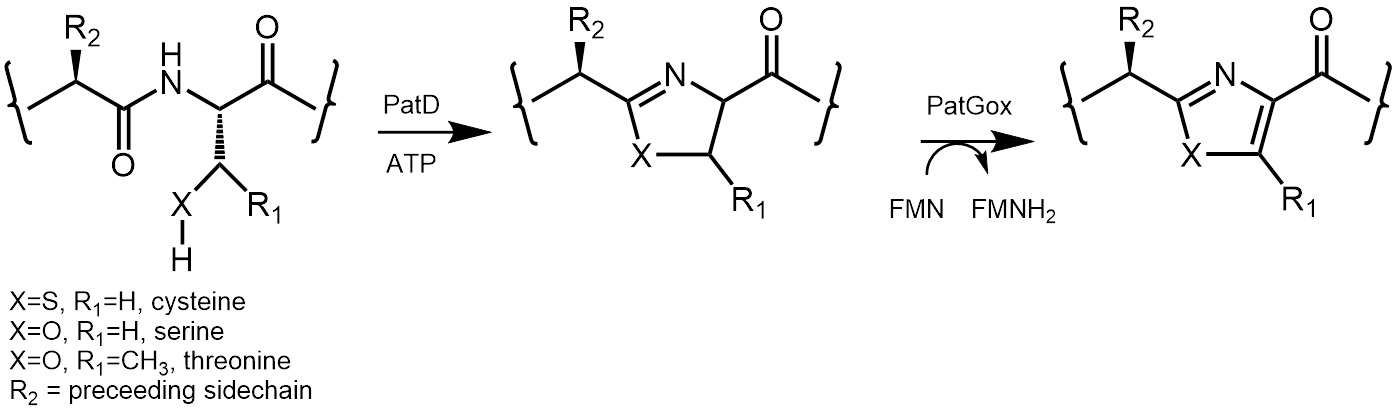
Figure 1: The oxidase enzyme is part of the patellamide biosynthetic pathway. It transforms reactive thiazoline and oxazolines into stable thiazoles and oxazoles. The reaction relies on the co-factor flavin mono nucleotide (FMN).
Proteins, in particular enzymes, are at the heart of biology. Enzymes carry out virtually all the chemistry that humans and all other living things need to survive. Enzymes catalyse conversions in that they make the reaction faster without being changed in the process. Bacteria do not have the luxury of using pre-made chemicals in the way humans do by eating, rather they make molecules from scratch. Some of these molecules have very interesting properties such as reversing drug resistance in cancer. Making such complex molecules using organic chemistry is very hard, expensive and can be polluting. By understanding enzymes , they can be put to use in making new, interesting and useful molecules. Having the three dimensional structure of an enzyme allows scientists to begin to understand how the enzyme works and opens up possible ways to engineer it for further use.
Protein structures are obtained by a variety of methods but in some cases all these approaches can fail. This is because although intensity of X-ray diffraction from a crystal can be measured, the phases cannot be measured, these need to be derived by another approach. Since producing crystals of a protein can take years, failing to solve the structure means being left with a crucial gap in knowledge and a large amount of wasted work. The new beamline I23 at Diamond was designed precisely as a ‘magic bullet’1 for these ‘unsolvable’ protein structures. The premise is simple, since almost all proteins contain sulfur atoms (present in amino acids cysteine and methionine) by collecting diffraction data at a wavelength where one excites electrons in sulfur, one can determine phases and thus the protein structure for almost every protein that crystallises. Phase determination by anomalous scattering can be used for other elements such as selenium that can be introduced by molecular biology into some but certainly not all proteins. Crucially, the closer the wavelength is to the ionisation energy of the atom in question, the larger and more useful the anomalous diffraction effect. Many structures are solved by the native sulfur single anomalous diffraction (S-SAD) technique already but they require high resolution diffraction (not always possible due to crystal quality) and a high percentage of sulfur atoms (rules out many proteins). These limitations are imposed by the wavelength that can be used to collect data (up to 2 Å) on current beamlines. Collecting data at wavelengths that are close to the ionisation of sulfur can overcome these limitations. However, such ‘long’ or ‘low energy’ X-rays create enormous technical challenges in measuring accurate data due to air scattering, absorbance and detector geometry. Inaccuracy would be fatal since one is relying on very small differences. I23 works in vacuum, uses special optics and employs a specially designed curved detector1. This is the first report of its use to solve an ‘unsolvable’ structure.
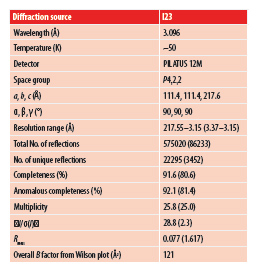
There is great interest in the patellamide biosynthetic pathway2 and all enzyme structures3,4 but that of the oxidase enzyme have been determined (Fig 1). To properly harness the pathway for biotechnology, the oxidase enzyme required to be characterised. Conventional attempts to determine the structure had failed, heavy metal soaking destroyed crystals, a selenium variant did not crystallise and molecular replacement failed. Conventional S-SAD phasing approaches at Diamond I02 also failed, this is because it needed accurate measurement of a signal (the difference between two measurements) of less than 1 % with poorly diffracting crystals (2.65 Å). In moving to I23 and collecting at a wavelength of 3.1 Å the size of the signal was increased to around 2 %. This would represent a challenge even for normal methods, as it requires very precise measurements. Collecting data at 3.1 Å with such accuracy would be impossible without the design features of I23. Data from many crystals were collected and only one of these was of sufficiently high quality to permit structure determination, reflecting the difficulty of the problem (Fig. 3). The need for so many data sets was a reflection of the variability and low quality of the crystals that were available.
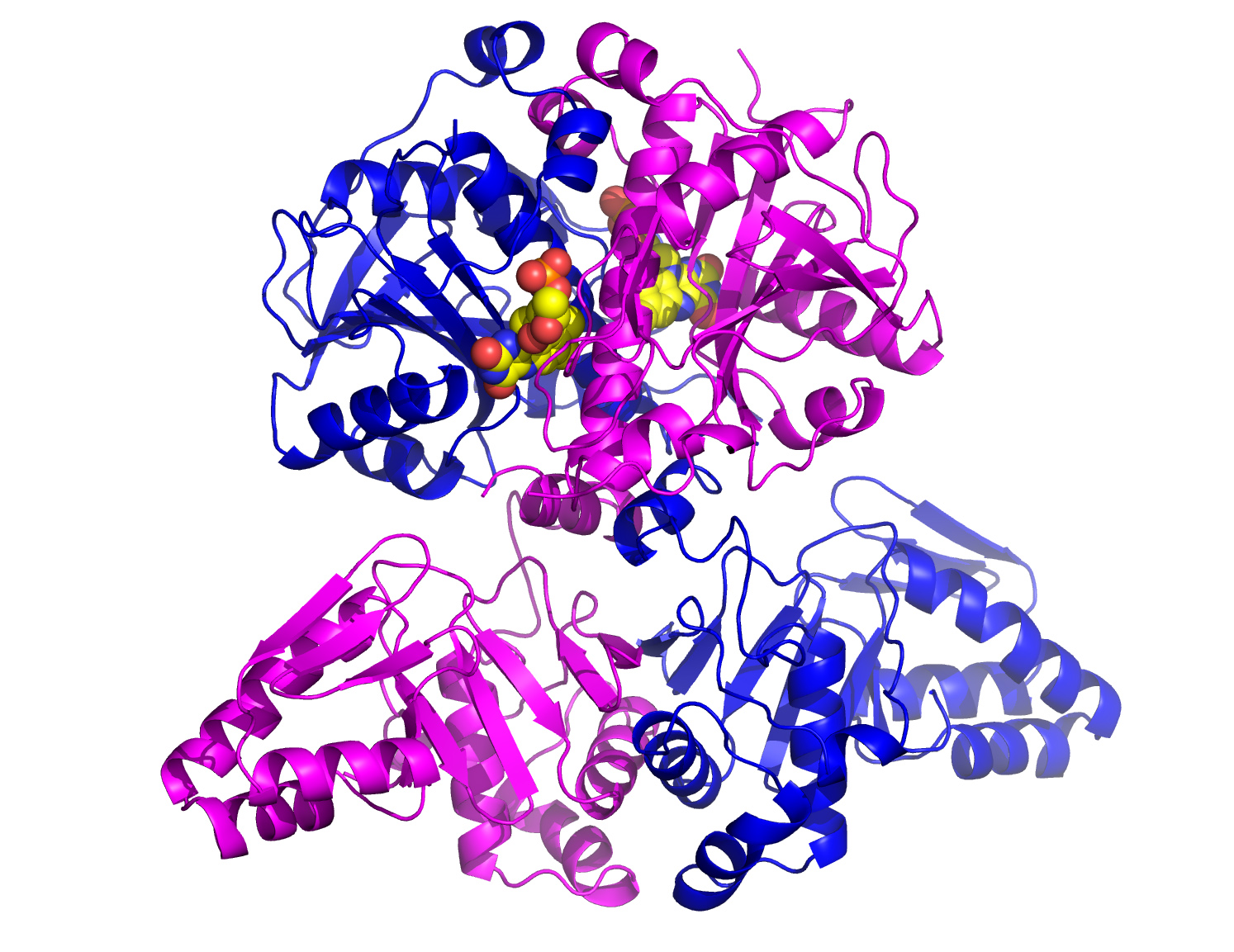
These data allowed the first solution of the structure. The solution was of sufficient quality to allow skilled manual intervention and finally determination of the complete structure (Fig. 2). The structure reveals a dimer, with each monomer consisting of two domains. The C-terminal domain binds the FMN co-factor (Fig. 2) and contains the active site. In fact the N-terminal domain at first appeared entirely novel but can be further subdivided into two peptide clamp domains. This is the first time such an arrangement has been seen and provides valuable clues for engineering studies.
The long wavelength, the in-vacuum sample environment and the large semi-cylindrical PILATUS 12M pixel hybrid detector were all essential in solving this extremely challenging structure. I23 represents a transformational advance in solving structures by native S-SAD.
References:
Funding acknowledgement:
This research was supported by grants from the UK Biotechnology and Biological Research Council (No. BB/K015508/1; JHN and MJ) and the European Research Council (No. 339367; JHN and MJ).
Corresponding author:
Professor James H Naismith, University of St Andrews, [email protected]
Related publication:
Bent AF, Mann G, Houssen WE, Mykhaylyk V, Duman R, Thomas L, Jaspars M, Armin Wagner A, Naismith JH. Structure of the cyanobactin oxidase ThcOx from Cyanothece sp. PCC 7425, the first structure to be solved at Diamond Light Source beamline I23 by means of S-SAD. Acta Crystallographica Section D Structural Biology 72, 1174-1180, doi:10.1107/S2059798316015850 (2016).
Publication keywords:
Cyanobactins; Azoline oxidase; S-SAD; RIPPs; Phasing; Structure; Sulfur; Long -wavelength
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.