Just as humans have an ancestry, so too do viruses. But whereas we can use fossils to help identify the creatures that roamed the earth before us, viruses are much harder to classify and have left no fossilised remains for us to study. A group of researchers from Diamond Light Source and the Wellcome Trust Centre for Human Genetics are currently working on piecing together a complete history of viruses, mapping out their evolutionary lineage by solving the 3D structures of the organism’s proteins. Research published today (13 July 2011) in Structure fills in an important part of the picture.
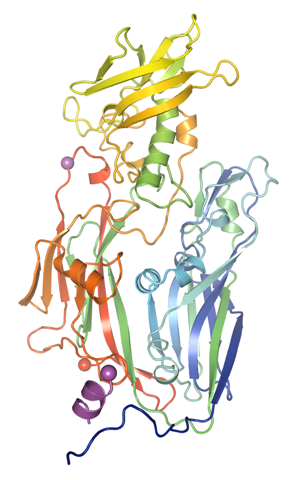 The group have successfully solved the 3D structure of the protein D13 (pictured left) from Vaccinia virus (VACV), a member of the poxvirus family. Known as a scaffolding protein D13 causes the VACV particles, or virons, to take the shape of a sphere as the virus starts to form, before D13 is lost and the viron takes on a more brick-like shape.
The group have successfully solved the 3D structure of the protein D13 (pictured left) from Vaccinia virus (VACV), a member of the poxvirus family. Known as a scaffolding protein D13 causes the VACV particles, or virons, to take the shape of a sphere as the virus starts to form, before D13 is lost and the viron takes on a more brick-like shape.
“This is the first time we have been able to determine the structure of this scaffolding protein,” says Dr Mohammad Bahar of the Structural Biology Division of the Wellcome Trust Centre for Human Genetics. “We found that it has a very distinctive shape, a double jelly-roll fold. This particular feature is also found in large DNA viruses such as human adenovirus, a cause of respiratory and eye infections. By comparing the structural similarities of the different viruses’ scaffolding proteins, we can determine Vaccinia virus’ lineage and place it within our virus ‘family tree’.”
Knowing who or what a particular organism is descended from can tell us a great deal about how it might function. In the case of viruses, this kind of knowledge could help us to find new ways to treat them.
 “If the structural commonalities are known, these links can be used to create therapies that work on a family of viruses, as opposed to just one,” explains Prof David Stuart, Professor of Structural Biology at the Wellcome Trust Centre for Human Genetics and Life Sciences Director at Diamond. “With this approach, it is possible that we could be able to treat patients who are suffering from one of a number of viruses, in the same way that antibiotics are used to treat bacterial infections.”
“If the structural commonalities are known, these links can be used to create therapies that work on a family of viruses, as opposed to just one,” explains Prof David Stuart, Professor of Structural Biology at the Wellcome Trust Centre for Human Genetics and Life Sciences Director at Diamond. “With this approach, it is possible that we could be able to treat patients who are suffering from one of a number of viruses, in the same way that antibiotics are used to treat bacterial infections.”
The image to the right shows Vaccinia Virus within an evolutionary virus tree.
The group used I03, one of Diamond’s Macromolecular Crystallography (MX) beamlines to solve the structure of D13. I03 was one of the first seven beamlines to go into operation at the Diamond synchrotron when it opened in 2007. Since then 210 structures of proteins solved on I03 have been deposited in the Protein Data Bank (PDB) and 157 papers incorporating I03 data have been published in peer reviewed journals.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.