Dr Pedro Matias, ITQB, Universidade Nova de Lisboa
The three-dimensional structure of [NiFeSe] hydrogenase from D. vulgaris Hildenborough in its oxidised, “as isolated” state has been determined from a 3-wavelength MAD experiment at the Fe K-edge on Diamond Beamline I04. Crystals were obtained in space group P21 with cell parameters a = 60.6, b = 91.2, c = 66.7 Å, b = 101.7º and one hydrogenase molecule in the asymmetric unit. This work has recently been published [1]. Previously, only the structure of [NiFeSe] hydrogenase from Dm. baculatum [2] in its reduced state was known.
Our results showed that in the oxidised state there is an exogenous sulphur atom present in the active site, which is bound to the Ni atom and to the Se atom of the selenocysteine. This moves the selenocysteine side-chain conformation into a different rotamer than that observed for the reduced structure in Dm. baculatum, shielding the Ni atom from attack by O2 molecules. We speculate that, when the enzyme is activated in production and consumption modes, this sulphur atom can be released as either H2S or HS- and becomes located in a binding site near the active site, which in our structure was found occupied by a Cl- ion. When the enzyme becomes quiescent, the sulphur species can become re-attached to the active site.
These results therefore provide a structural basis for the oxygentolerance of [NiFeSe] hydrogenases.
We also observed the partial oxidation of the proximal Fe4S4 cluster to an apparent stoichiometry of Fe4S3O3. This may be an artifact of the reparation and/or crystallization under aerobic conditions, or it may be an additional protection mechanism of the NiFe active site against inactivation by oxygen.
Work towards the structural determination of this enzyme in the reduced (with H2) and reoxidised (exposure to air after reduction with H2) is in progress. Preliminary data have been obtained, and the current goal is to improve the resolution of the diffraction data.
Presently, hydrogen is a very attractive candidate for storing and transporting energy due to its high energetic content and lack of contribution to the greenhouse effect. Current hydrogen production is mainly from fossil fuels in processes that are not environmentally friendly. Production of hydrogen from renewable sources would be of fundamental importance to its viability as an alternative fuel. As such, the use of microorganisms in biohydrogen production is one of the most interesting options [3].
Biological hydrogen conversion is carried out by extremely efficient enzymes called hydrogenases (Hases). They are the natural molecular reactors that catalyse the production and the oxidation of the hydrogen molecule, which are widely found in microorganisms, particularly in anaerobic bacteria [4].
Both main groups of Hases, the [NiFe] (which includes the [NiFeSe] subgroup) and the [FeFe] Hases, contain a complex metal-based catalytic centre, with two metals (Ni and Fe or two Fe) coordinated by thiolates, CO and CN- ligands [5,6]. Their unique active site architecture is able to carry out the difficult reactions of H2 production/oxidation at ambient conditions, without involving noble metals such as platinum.
Although [NiFeSe] hydrogenases are structurally very similar to the more common [NiFe] hydrogenases, they are more active in hydrogen production and also they are oxygen- tolerant, i.e., do not become inactivated by reaction with small amounts of oxygen. This is reflected by their different EPR spectra, since no Ni-A and Ni-B states are observed for these enzymes, corresponding to a bridging oxygen species between the Ni and Fe atoms in the active site. In addition, FTIR studies show that several structural forms of the active site may be present in the “as-isolated” oxidised form of the enzyme, and that the dominant form after reduction with H2 followed by reoxidation appears to be different from that prevalent in its “as-isolated” oxidised [7].
Typically, [NiFe] hydrogenases and [NiFeSe] hydrogenases are heterodimers, where the smaller chain contains the FexSy clusters (3 Fe4S4 in [NiFeSe] hydrogenases; two Fe4S4 and one Fe3S4 in [NiFe] hydrogenases) responsible for electron transfer and the large subunit contains the NiFe active site. One hydrophobic tunnel leads out from the active site, and divides into several branches as it nears the surface of the molecule.
| Figure 1 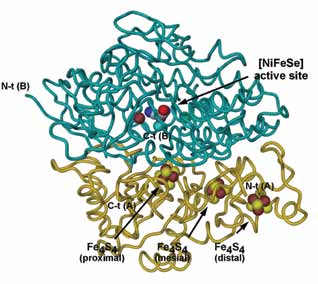 | | [NiFeSe] hydrogenase from D. vulgaris Hildenborough is a bacterial lipoprotein, found in the membrane fractions during purification [8]. It has a lipid group attached to a cysteine residue near its N-terminal. However, over time there is a cleavage of the first 11 N-terminal residues, resulting in a soluble form of the enzyme. Only the soluble form could be so far crystallised. We purified and crystallsed the soluble form of [NiFeSe] hydrogenase from D. vulgaris Hildenborough [9]. Although the enzyme was relatively easy to crystallise, it was more difficult to produce single crystals for diffraction measurements. The crystals obtained were characterised in-house and a dataset to 2.4 Å resolution was measured [9]. A preliminary molecular replacement solution was obtained using the reduced structure of [NiFeSe] hydrogenase from Dm. baculatum (PDB 1CC1) as a search model. However, there were several regions in the structure which were not clear and therefore a 3-wavelength MAD experiment was carried out at Diamond Beamline I04, using the anomalous signal of the expected 13 Fe atoms (corresponding to one Hase heterodimer) present in the structure to derive phase information. Three datasets (peak, inflexion point and high-energy remote) were measured to ca. 2 Å resolution. The crystal suffered some radiation damage during the experiment, but nevertheless all datasets could be used. |
| Figure 1. The structure of D. vulgaris [NiFeSe] Hase. Cα tube diagram representation of the overall structure, with the co-factors represented as spheres; the small subunit (chain A) is coloured gold and the large subunit (chain B) is coloured dark cyan. Atom colours are light blue for carbon, blue for nitrogen, red for oxygen, yellow for sulphur, brown for iron, cyan for nickel and green for Cl-. |
The MAD data set was analyzed with SHELXC, the heavyatom substructure was determined with SHELXD and the phase problem solved with SHELXE. A maximum-likelihood heavy-atom parameter refinement using SHARP was used to obtain improved MAD phases, followed by density modification with SOLOMON and DM. ARP/wARP produced a near complete model, with 690 residues out of the expected 770 built and docked into sequence, with R = 0.180 and Rfree = 0.250. The remaining residues (except for a few disordered N-terminal residues in both subunits) as well as the Fe4S4 and the NiFe co-factors were modelled with COOT.
The structure was refined against the 2.04 Å peak dataset with REFMAC to R = 0.144 and and Rfree = 0.201. Electron density was observed for the aliphatic chains of four Sulfobetaine 3-12 detergent molecules (used in purification and crystallization) in two hydrophobic surface pockets and their visible atoms were included in the refinement. Solvent molecules were located with ARP/wARP and by inspection of the 2|Fo|-|Fc| and |Fo|-|Fc| electron density maps with COOT. In the final model, continuous electron density is present for residues 7-283 (C-terminal) in the small subunit and 15-495 (C-terminal) in the large subunit.
In the large subunit, the side-chain of Cys 75 was refined with a doubly oxidised Sg atom (cysteine S-dioxide), the side-chain of SeCys 489 was refined with three-fold disorder (with populations 70%, 15%, 15%), two of those having a bound sulphur atom, and in the proximal subunit Fe4S4 cluster was refined as partly oxidised (40%) to Fe4Se3O3.
 | Figure 2. [S]-SeCys 489 conformers in the active site of D. vulgaris [NiFeSe] Hase. Ca tube diagram representations of the large subunit, with the NiFe(CN)2(CO) cluster and the side-chain atoms of the cysteine and selenocysteine residues that bind it to the protein chain drawn in ball-and-stick. The colouring scheme is the same as in Figure 1. (a) Conformer I, S-SeCys 489; (b) Conformer II, S-SeCys 489; (c) Conformer III, SeCys 489 with H2S/HS- in the “storage” site. |
Our results showed that in the oxidised state there is an exogenous sulfur atom present in the active site, which is bound to the Ni atom and to the Se atom of the selenocysteine. This moves the selenocysteine side-chain conformation into a different rotamer than that observed for the reduced structure in Dm. baculatum, shielding the Ni atom from attack by O2 molecules. These results provide a structural basis for the oxygen-tolerance of [NiFeSe] hydrogenases.
Interestingly, oxidised variants of one of the Ni-bound cysteines and the proximal Fe4S4 cluster were observed. This may be an artifact of the preparation and/or crystallization under aerobic conditions, or it may be an additional protection mechanism of the NiFe active site against inactivation by oxygen.
Finally, the two structural forms of the S-bound SeCys in the active site probably correspond to the different features observed in the FTIR spectra of the “as-isolated” and reoxidised forms of the enzyme.
References
[1] M. C. Marques, R. Coelho, A. L. De Lacey, I. A. C. Pereira, P.M. Matias, J. Mol. Biol. 396: 893-907 (2010). DOI: 10.1016/j. jmb.2009.12.013.
[2] Garcin, E., Vernede, X., Hatchikian, E. C., Volbeda, A., Frey, M. & Fontecilla-Camps, J. C. Structure, 7, 557-566 (1999).
[3] Levin, D. B., Pitt, L. & Love, M. Int. J. Hydrogen Energy, 29, 173-185 (2004).
[4] Vignais, P. M. & Billoud, B. Chem. Rev. 107, 4206-4272. (2007).
[5] Fontecilla-Camps, J. C., Volbeda, A., Cavazza, C. & Nicolet, Y. Chem. Rev. 107, 4273-4303. (2007).
[6] Matias, P. M., Pereira, I. A., Soares, C. M. & Carrondo, M. A. Prog. Biophys. Mol. Biol. 89, 292-329 (2005).
[7] De Lacey, A. L., Gutierrez-Sanchez, C., Fernandez, V. M., Pacheco, I. & Pereira, I. A. J. Biol. Inorg. Chem. 13, 1315-1320 (2008).
[8] Valente, F. M., Pereira, P. M., Venceslau, S. S., Regalla, M., Coelho, A. V. & Pereira, I. A. FEBS Lett. 581, 3341-3344 (2007).
[9] Marques, M., Coelho, R., Pereira, I. A. C. & Matias, P. M. Acta Crystallogr.,Sect F: Biol. Cryst. Commun. 65, 920-922. (2009).
Principal Publications and Authors
M. C. Marques, R. Coelho, A. L. De Lacey, I. A. C. Pereira, P. M. Matias.The three-dimensional structure of [NiFeSe] Hydrogenase from Desulfovibrio vulgaris Hildenborough: a Hydrogenase without a bridging ligand in the active site in its oxidised, “as-isolated” state, J. Mol. Biol. 396: 893-907 (2010). DOI: 10.1016/j.jmb.2009.12.013
Funding Acknowledgement
European Community’s Seventh Framework Programme (FP7/2007-2013) under grant agreement nº 226716. Research grant PTDC/BIA-PRO/70429/2006 funded by Fundação para a Ciência e Tecnologia (FCT, MCES, Portugal) and FEDER program, the research grant CTQ2006-12097 funded by the Ministerio de Ciencia e Innovación (Spain) and a Luso-Spanish Joint Action funded by Conselho de Reitores das Universidades Portuguesas and the Ministerio de Educación y Ciencia.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.