Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()
A team of researchers from the Universities of Leeds, Oxford and Imperial College London have captured the 3D atomic models of a single transporter protein in each of its three main structural states, a goal of researchers from around the world for over 25 years.
The discovery offers remarkable insight into the function of one of the body’s most fundamental processes – the movement of essential chemicals into cells of the body - and creates the opportunity to develop brand new drugs.
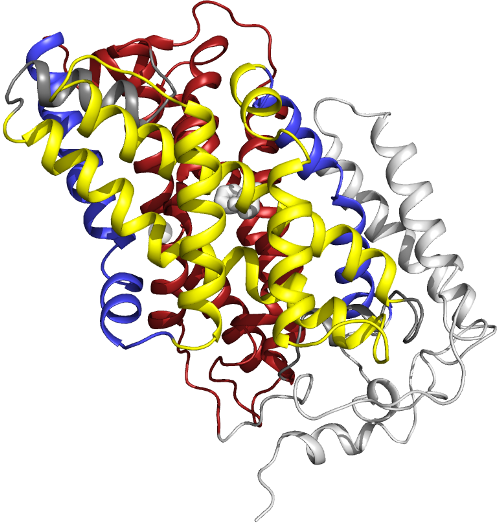 |
| The inward-facing structure of the bacterial Mhp1 transporter protein |
Biologists have surmised that transporter proteins of this type, which sit in the cell membrane, carry molecules through the otherwise impermeable membrane by shifting between at least three distinct structural states, controlled by ion gradients.
In the first state, there is an outward-facing cavity. A compound will enter this cavity and attach to a binding site whereupon the protein will move to a second state with the cargo locked inside. The third state is formed when the protein opens up a cavity on the inward-facing side to release the compound into the cell. The switch between outward and inward-facing sides works rather like a ‘kissing gate’ in which the cavity is either on one side or the other but there is never a direct channel through the protein.
However, until now, scientists had never observed the structural details of these three states in a single protein and theories about how the mechanism worked in detail were based on stitching together their observations from different transporters.
Professor Peter Henderson is from the University of Leeds.
“Previous models gave us a broad understanding of the mechanism involved, but this could never really be usefully applied for drug development. The goal for researchers in this area has always been to observe the entire mechanism in a single protein.”Professor Peter Henderson
The research, published in Science in April 2010, reports the mapping of the inward-facing structure of the bacterial Mhp1 transporter protein, the third structural state that they have determined for this protein. The team has been studying Mhp1 for more than ten years and their observations of the first two structures were published in Science in October 2008.
The protein was produced in Leeds; the structures were determined by X-ray crystallography and analysed at Imperial College and the Imperial College Membrane Protein Laboratory (MPL) located at Diamond Light Source. To further investigate the transitions between the three states of the protein dynamic molecular simulations were carried out at Oxford University.
“This third structure completes the picture and we can now understand Mhp1’s ‘alternating access’ mechanism in great detail. We also unexpectedly found that the structures are similar across many transporter proteins previously thought to be different, so we’re expecting our model to help achieve some rapid progress in the research of colleagues around the world.”Dr Alexander Cameron, Group Leader in the MPL
The detailed knowledge of the mechanism could unlock new drug developments in several ways, says Professor Henderson. “Altering the delivery of compounds into a cell is one potential benefit for treating illness. For example this could be useful in treating conditions where certain chemicals are lacking and need boosting permanently - such as serotonin for those suffering from depression and glucose for those with diabetes.”
The mechanism’s detail is already being used by chemists in the EU-funded European Drug Initiative on Channels and Transporters consortium (EDICT), which Professor Henderson leads. “We’ve found around 20 compounds that match Mhp1’s binding site, and of these, three have been shown to bind. I think we are entering an exciting period of discovery.”
“It’s taken a long time to get to this point – over ten years – but then difficult science takes time. This is the point at which blue skies research evolves into useful applications,” says Professor Henderson. “It’s the best thing I’ve been involved in during my academic career.”
Funded by the Biotechnology and Biological Sciences Research Council (BBSRC), the EU, the Japanese Science and Technology Agency and the Wellcome Trust, the research team comprises: Professor Henderson and colleagues from Leeds, who expressed and purified the Mhp1 protein; Professor So Iwata, Dr Alex Cameron and colleagues from Imperial College London and the MPL who imaged the crystals and combined all the information to propose a mechanism for the alternating access model; and Professor Mark Sansom and Dr Oliver Beckstein from the University of Oxford who authenticated the plausibility of the transitions between the three states.
Professor Henderson says that the next step is to investigate what triggers the protein to change between the states and the team has secured further funding from BBSRC and the EDICT consortium to pursue this.
Clare Elsley, Campuspr Ltd: Tel 44 (0) 113 258 9880, mobile 44 (0) 7767 685168 email clare@campuspr.co.uk
Guy Dixon, University of Leeds press office: Tel 44 (0)113 343 8299, email g.dixon@leeds.ac.uk
Lucy Goodchild, Imperial College London press office: Tel +44 (0)20 7594 6702, email lucy.goodchild@imperial.ac.uk
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.